Engineering more stable proteins†
Abstract
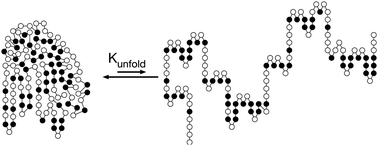
Protein function requires the folded protein form, but this form is unstable mainly because it readily unfolds into a flexible, unstructured form. Protein folding is favored by burying of hydrophobic side chains and hydrogen bonding between the amino acids. Protein unfolding is favored by the increase in conformational freedom of the main chain of amino acids upon unfolding. Protein stability is usually measured by the reversible unfolding of the protein with either heat or chemical additives like urea. Engineering mores stable proteins involves making substitutions that shift the folding–unfolding balance toward the folded form. Stabilizing substitutions can either stabilize the folded conformation or destabilize the unfolded ensemble. This tutorial emphasizes web-based tools to identify substitutions that stabilize proteins. Besides unfolding, other sources of protein instability are chemical modifications like oxidations or cleavage by proteases and aggregation of partly unfolded proteins into insoluble particles.
- This article is part of the themed collection: Protein engineering through chemical, genetic and computational manipulation


 Please wait while we load your content...
Please wait while we load your content...