Proteomic profile of 4-PBA treated human neuronal cells during ER stress†
Bhavneet
Kaur‡
ab,
Ajay
Bhat‡
ab,
Rahul
Chakraborty
ab,
Khushboo
Adlakha
a,
Shantanu
Sengupta
ab,
SoumyaSinha
Roy
*ab and
Kausik
Chakraborty
 *ab
*ab
aGenomics and Molecular Medicine, CSIR-IGIB, Mathura Road, New Delhi, India. E-mail: kausik@igib.res.in; kausik@igib.in
bAcademy of Scientific & Innovative Research (AcSIR), CSIR-IGIB, Mathura Road Campus, New Delhi, India
First published on 19th January 2018
Abstract
Perturbations affecting the homoeostasis of endoplasmic reticulum (ER) activate an adaptive signaling known as the unfolded protein response or UPR. Many studies have reported the association between neurological disorders and ER stress. Decreasing ER stress may therefore aid in therapeutic control of neuronal diseases. Sodium 4-phenylbutyrate (4-PBA), a small molecule, has been shown to alleviate ER stress and various neurological diseases, but the mechanistic basis of its action is not well understood. Using an iTRAQ based LC-MS technique we have delineated the effect of 4-PBA on the proteome of human neuroblastoma cells (SK-N-SH) during Tunicamycin-induced ER stress. The proteomic profile of 4-PBA-treated cells revealed that 4-PBA does not alter the cellular proteome to adapt towards ER stress. However, it can alleviate both the toxicity and proteomic alterations, induced by an ER stress inducer. Hence, the therapeutic effect of 4-PBA is primarily due to its ability to resolve ER stress rather than its ability to alter the expression of proteins required for maintaining ER proteostasis. Thus, we posit here that 4-PBA acts as an authentic chemical chaperone by aiding protein folding in the ER.
1. Introduction
Cellular stress leads to activation of pathways required for stress tolerance, failure of which can activate death signals.1 Accumulation of misfolded or unfolded proteins in the Endoplasmic Reticulum (ER) leads to ER stress, which is typically resolved by the stress response pathway known as the Unfolded Protein Response (UPR).2,3 The characteristic features of UPR involve the up-regulation of ER chaperones and foldases (folding enzymes), along with inhibition of protein translation and subsequent degradation of misfolded proteins.4 In higher eukaryotes, there are three branches of UPR comprising IRE1 (Inositol-requiring enzyme 1), PERK (PKR like ER resident kinase) and ATF6 (activation transcription factor 6). Activation of Ire1 results in increased expression of molecular chaperones like Hsp90 and GRP78 through splicing and translation of XBP-1. Activated PERK inhibits protein translation to decrease the load of proteins entering the ER for folding. ATF4, a transcription factor translated specifically during PERK activation, leads to expression of CHOP (C/EBP homologous protein), which upregulates pro-apoptotic factors. Activation of ATF6 happens through its release from the ER membrane by a protease-dependent cleavage. The cleaved protein is an active transcription factor that translocates to the nucleus and upregulates various UPR genes including XBP1.5 UPR is activated to resolve ER stress, but a prolonged ER stress that is unresolved by the adaptive changes may lead to activation of apoptotic cell death.6 This feature of chronic ER stress and UPR is associated with various metabolic diseases like cardiovascular disease and diabetes.7–9 It is also associated with Neurodegenerative diseases such as Alzheimer's, Huntington's and Parkinson's disease that are typically caused by aggregation and accumulation of wrongly folded or misfolded proteins.10–14Chaperones play a key role during ER stress in preventing incomplete and unfolded proteins from forming aggregates and assisting in the proper folding of defective proteins in the ER lumen.15,16 There are two major types of chaperones: molecular and chemical chaperones. Molecular chaperones are proteins that help other proteins to acquire proper conformation by interacting with them. Hsp70, a classical example of molecular chaperones, functions by binding to unfolded polypeptides and prevents aggregation. It is known to play a role in reducing the levels of aggregated α-synuclein in both in vivo and in vitro models.17 Chemical chaperones are small molecular weight compounds which assist in folding and stability of proteins. These are two major types: osmotic (polyols, amino acids, amino acid derivatives and methylamines) and hydrophobic compounds (bile salts and sodium 4-phenylbutyrate).18,19 Sodium 4-phenylbutyrate (4-PBA) is a hydrophobic short chain fatty acid. It is an FDA-approved drug for the treatment of urea cycle disorders.20 4-PBA has additionally shown promising results in various diseases such as cancer, spinal muscular atrophy, cystic fibrosis, and neurodegenerative diseases associated with the folding of proteins, such as ALS (amyotrophic lateral sclerosis), Huntington's disease, Alzheimer's disease and Parkinson's disease.21–23 Many reports suggest that 4-PBA can act as a chemical chaperone by preventing aggregation of misfolded proteins.24,25 A competing model proposes that 4-PBA has an HDAC (Histone deacetylase) inhibitor activity, and thereby may regulate the expression of various neuronal genes during neurodegenerative disorders which involve aberrant histone acetylation.23,26,27 However, despite the fact that 4-PBA has therapeutic potential in proteostasis disorders, the validity of the competing proposals has not been authenticated.
In this study using a human neuroblastoma cell line as a model system, we investigated the effect of 4-PBA during Tm (prevents N-linked glycosylation) induced ER stress. With the help of iTRAQ based quantitative proteomics, we identified the pathways affected by 4-PBA. This report highlights the first proteomic profile of 4-PBA-treated neuronal cells during ER stress. We show that the most likely route of 4-PBA action is through the alleviation of ER stress itself and not through the modulation of stress-response genes.
2. Materials and methods
2.1 Cell line and culture
The human neuroblastoma (SK-N-SH) cell line was purchased from ATCC (Rockville, MD, USA). The cells were maintained in Dulbecco's Modified Eagle's medium (catalog #11965092, Gibco) supplemented with 10% FBS (catalog #10082147, Gibco) and 100 units of antimycotic and antibacterial solution. Cells were cultured under standard growth conditions at 37 °C in a humidified atmosphere, with 5% CO2. In all the experiments in this study, three biological replicates were performed.2.2 Cell viability assay
Cell viability in the neuroblastoma cell line was assayed using trypan blue exclusion and lactate dehydrogenase leakage assay (LDH assay). In trypan blue assay, 5 × 104 cells were seeded in 24-well flat bottom culture plates and treated the next day with 3 μg ml−1 concentration of Tm for 15 h in the presence and absence of 1 mM 4-PBA. Following treatment cells were washed with PBS, followed by trypsinization. The trypsinized cells were collected (along with dead cells) and stained with 0.4% trypan blue solution for 2 min. Both live (trypan negative) and dead cell (trypan positive) populations were counted with the help of a hemocytometer and the toxicity was represented as a percentage of dead cells.LDH assay was performed using a cytotoxicity detection kit (cytotoxicity detection kit plus, Roche). In this assay, cells were grown in 96-well flat bottom plates at a seeding density of 1 × 104 cells per well, and upon 70–80% confluency, cells were treated with Tm and 4-PBA. Following treatment, LDH assay was performed according to the manufacturer's protocol. Briefly, 100 μl of freshly prepared reaction mix was added to an equal volume of the supernatant collected from each well for every experimental group and was incubated for 30 min at room temperature. After incubation, absorbance was measured at 490 nm using a microplate reader. Further, relative cytotoxicity was calculated using the following equation: {(Ab_Exp − Ab_Con)/(Ab_Pc − Ab_Con)} × 100, where Ab_Exp = absorbance of the experimental group, Ab_Con = absorbance of the control group, and Ab_Pc = absorbance of the positive control (provided with the kit).
2.3 Western blotting
Harvested cells were lysed in RIPA buffer supplemented with protease and phosphatase inhibitors. An equal concentration of protein (30 μg) was resolved by 8–12% SDS-PAGE and then transferred onto a 0.2 μm PVDF membrane (catalog #ISEQ00010, Millipore). The membranes were blocked by 5% BSA for 2 h and then incubated with the primary antibodies against BIP (catalog #610979, BD Biosciences) and CHOP (catalog #2895s, Cell Signaling Technology) at a 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1000 dilution overnight at 4 °C. The membranes were washed three times, for 15 min each, with TBS containing 0.1% tween and then subjected to horseradish peroxidase (HRP) conjugated secondary antibody at 1
1000 dilution overnight at 4 °C. The membranes were washed three times, for 15 min each, with TBS containing 0.1% tween and then subjected to horseradish peroxidase (HRP) conjugated secondary antibody at 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 3000. Detection was carried out using the supersignal chemiluminescent substrate for HRP (catalog #WBKLS0050, Millipore) and the image was captured using GBox (Chemi X4 Syngene).
3000. Detection was carried out using the supersignal chemiluminescent substrate for HRP (catalog #WBKLS0050, Millipore) and the image was captured using GBox (Chemi X4 Syngene).
2.4 Trypsin digestion and iTRAQ labeling
Total protein for iTRAQ experiment was isolated by cell lysis in RIPA buffer, as mentioned above, for immunoblotting. Trypsin digestion was performed using 60 μg of protein as described previously.28–30 Briefly, total protein from each group was reduced with 25 mM DTT at 56 °C for 30 minutes and then treated with 55 mM IAA at room temperature for 15–20 minutes for blocking cysteine. The samples were then incubated with trypsin in a 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 10 ratio (trypsin to protein) for 16–18 h at 37 °C. The tryptic peptides of each sample were labeled with different 4plex-iTRAQ reagents (114, 115, 116, and 117) as per the manufacturer's instructions (Sciex, Foster City, CA). Briefly, each iTRAQ tag was kept at room temperature for 15–20 minutes and then re-suspended in 70 μl ethanol and mixed properly. Tryptic digested samples were then mixed with the corresponding tag and kept at room temperature for 1 hour. To avoid bias due to the labeling efficiency, we swapped the tags in different experimental groups (ESI,† Fig. S1). In each iTRAQ experiment, all the four individually tagged samples were mixed and pooled into a single centrifuge tube and then vacuum dried at 30 °C.
10 ratio (trypsin to protein) for 16–18 h at 37 °C. The tryptic peptides of each sample were labeled with different 4plex-iTRAQ reagents (114, 115, 116, and 117) as per the manufacturer's instructions (Sciex, Foster City, CA). Briefly, each iTRAQ tag was kept at room temperature for 15–20 minutes and then re-suspended in 70 μl ethanol and mixed properly. Tryptic digested samples were then mixed with the corresponding tag and kept at room temperature for 1 hour. To avoid bias due to the labeling efficiency, we swapped the tags in different experimental groups (ESI,† Fig. S1). In each iTRAQ experiment, all the four individually tagged samples were mixed and pooled into a single centrifuge tube and then vacuum dried at 30 °C.
2.5 Separation of tryptic peptides and LC-MS method
iTRAQ labeled tryptic peptides were separated by cation exchange (SCX) chromatography using an SCX Cartridge (5 micron 300 A bead from Sciex, USA), with a cartridge holder (Sciex, USA) as described previously.29,30 Fractionation of samples was performed by a step gradient of increasing concentration of ammonium formate (35 mM, 50 mM, 75 mM, 100 mM, 125 mM, 150 mM, 250 mM and 350 mM ammonium formate) prepared in 30% v/v ACN and 0.1% formic acid (pH = 2.9). These fractionated peptides were analyzed on a quadrupole-TOF hybrid mass spectrometer (TripleTOF6600, Sciex, USA) coupled to a nano-LC system (Eksigent NanoLC-400). Ten microliters of sample was injected and loaded onto a reverse phase peptide ChromoLC trap (200 μm × 0.5 mm) column and desalted at a flow rate of 2 μl per minute for 45 minutes. After desalting, peptides were separated using a C18 column (75 μm × 15 cm, Eksigent). The samples were run using a gradient method using buffer A (99.9% LC-MS water + 0.1% formic acid) and buffer B (99.9% acetonitrile + 0.1% formic acid). The gradient consists of 95% of buffer A for 2 minutes, and then shifted to 90% of buffer A for 8 minutes, and then decreased to 20% of buffer A in 80 minutes and finally again shifted to 95% of buffer A for 10 minutes at a consistent flow rate of 250 nL min−1. Data were acquired with a NanoSpray source installed in the TripleTOF 6600 System using a nebulizing gas of 25 psi, a curtain gas of 25 psi, an ion spray voltage of 2400 V and a heater interface temperature of 75 °C. Information-dependent acquisition (IDA) mode was setup with a TOF/MS survey scan (400–1600 m/z) with an accumulation time of 250 ms. For fragmentation a maximum of ten precursor ions per cycle were selected with each MS/MS spectrum (100–1800 m/z) accumulated for 70 ms with a total cycle time of approximately 2.05 seconds. Parent ions with a charge state from +2 to +5 and abundance of more than 150 cps were selected for MS/MS fragmentation. Once an ion had been fragmented by MS/MS, its mass and isotopes were excluded for a period of 3 s. High sensitivity mode with ‘adjust collision energy when using iTRAQ reagent’ settings was used to acquire MS/MS spectra.2.6 Database searching and statistical analysis
All the ‘.wiff’ files containing MS and MS/MS spectra generated from Triple TOF 6600 were analyzed using the Protein Pilot v5.0 software (Sciex). The Paragon algorithm was used in a “Thorough ID” search mode against the Uniprot Homo sapiens reference dataset. The following parameters were included in the search: trypsin as the digestion enzyme with two missed cleavages, modifications by IAA as cysteine blocking reagent, iTRAQ 4-plex modification of the N termini of peptides and of the side chains of lysine. An automatic decoy database search was also performed to calculate the false discovery rate (FDR) and 1% global protein level FDR was considered for protein identification. Further, the resulting dataset was normalized using auto-bias correction to remove any experimental bias. Reproducibility of the relative expression of each protein across different biological replicates was assessed by calculating percentage of the coefficient of variation. A cutoff of 1.2 fold was set for identifying proteins differentially expressed by Tm and 4-PBA treatment. Among the Tm induced differentially expressed proteins, to identify the proteins whose expression was restored by 4-PBA, we performed the unpaired Student's t-test, individually for each protein between Tm/control and (Tm + 4-PBA)/control with p < 0.05 set as the criteria for significant change.3. Results
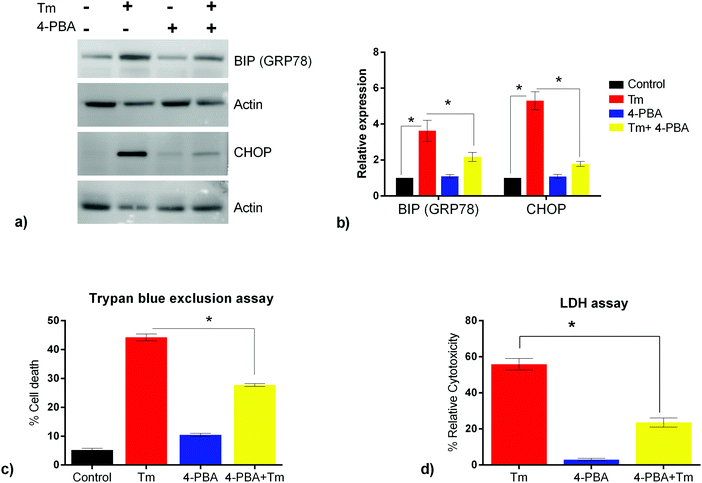
ER stress has been implicated as a contributing factor in neurodegenerative diseases. Previously 4-PBA has been reported to protect against neuronal cell death caused by ER stress31 but the precise mechanism behind 4-PBA mediated protection remains elusive. Thus, in this study, to assess the protective effects of 4-PBA in neuronal cells under ER stress, we treated human neuroblastoma cells (SK-N-SH) with Tm, to induce ER stress. We then analyzed the expression of ER stress markers, Grp78 and CHOP, after treatment with 3 μg ml−1 Tm for 15 h in the presence and absence of 1 mM 4-PBA. We observed a statistically significant increase in the expression of Grp78 and CHOP after Tm treatment in SK-N-SH cells, which reverted on pre-treatment with 4-PBA (Fig. 1a and b). Further, the 4-PBA treatment significantly decreased Tm induced cell death (Fig. 1c and d).To have a comprehensive understanding of the protective effect of 4-PBA during ER stress in neuronal cells, we performed 4-plex iTRAQ-based relative quantitative proteomics experiment under different conditions: control (without any treatment), Tm treated, 4-PBA treated and a combination of 4-PBA and Tm treatment. In the three biological replicates, we identified 3665, 3199, and 3510 proteins respectively, with 1% FDR and at least two unique peptides. Of these, 2023 proteins were identified in all the three experiments (Fig. 2 and ESI,† Table S1). The % CV was calculated across the three biological replicates to check the reproducibility of the experiments. Out of 2023 proteins, 88% of the proteins had a CV of <20% (data not shown). The relative expression of GRP78 (a well-known marker of ER stress) acquired from this experiment was in agreement with the fold change obtained from the immunoblotting technique (ESI,† Fig. S2). This validated the relative quantification acquired through the iTRAQ approach.
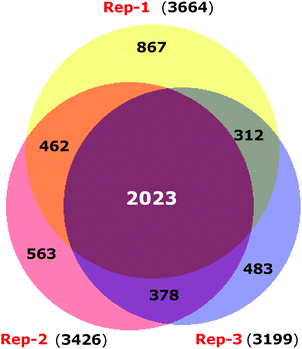 | ||
| Fig. 2 Venn diagram displaying the overlap of proteins identified in three biological replicates of iTRAQ experiments. | ||
We further investigated the differential expression of proteins in Tm-treated cells to identify the cellular response towards an ER stress inducer. Proteins were considered to be upregulated by Tm, if the fold change with respect to the control was ≥1.2 in all biological replicates. Similarly, a fold change of ≤0.8 was used for downregulated proteins. Using these criteria of selection, 101 proteins were found to be differentially expressed of which 84 were upregulated and 17 were downregulated (Table 1). In agreement with a previous report, Tm-treatment led to the overexpression of proteins involved in protein folding and cellular redox homeostasis.32 Several proteins like GRP78, DNAJB11, PDIA4, PDIA3, and ERO1 that are involved in unfolded protein response were upregulated. This further indicates that in our experimental conditions, Tm was able to cause ER stress and induce unfolded protein response. These differentially expressed proteins were classified based on their Gene Ontology (GO) parameters of the biological function and cellular localization using the Database for Annotation, Visualization and Integrated Discovery (DAVID). GO analyses based on cellular localization revealed that the majority of the Tm-induced differentially expressed proteins are localized in the mitochondria, extracellular exosomes and endoplasmic reticulum (Fig. 3 and ESI,† Table S1). Surprisingly 54% of the differentially expressed proteins belonged to extracellular exosomes, highlighting a yet underappreciated connection that demands further investigation to understand the modulation of extracellular communication during ER stress. The biological function analysis further showed that proteins involved in mitochondrial ATP synthesis coupled proton transport, protein folding, response to ER stress, cellular redox homeostasis, and nucleosome assembly were also significantly enriched (Fig. 3).
| Up-regulated | Fold change: Tm/control | |||
|---|---|---|---|---|
| Acession ID | Description | Rep 1 | Rep 2 | Rep 3 |
| P11021 | 78 kDa glucose-regulated protein OS = Homo sapiens GN = HSPA5 PE = 1 SV = 2 | 3.47 | 3.17 | 3.09 |
| P30101 | Protein disulfide-isomerase A3 OS = Homo sapiens GN = PDIA3 PE = 1 SV = 4 | 1.91 | 1.94 | 1.85 |
| P13667 | Protein disulfide-isomerase A4 OS = Homo sapiens GN = PDIA4 PE = 1 SV = 2 | 2.3 | 2.27 | 2.11 |
| P07237 | Protein disulfide-isomerase OS = Homo sapiens GN = P4HB PE = 1 SV = 3 | 1.82 | 1.87 | 1.75 |
| P06576 | ATP synthase subunit beta, mitochondrial OS = Homo sapiens GN = ATP5B PE = 1 SV = 3 | 1.42 | 1.39 | 1.4 |
| P25705 | ATP synthase subunit alpha, mitochondrial OS = Homo sapiens GN = ATP5A1 PE = 1 SV = 1 | 1.37 | 1.32 | 1.36 |
| B4DGP8 | Calnexin OS = Homo sapiens GN = CANX PE = 2 SV = 1 | 1.86 | 1.92 | 1.82 |
| H6VRG3 | Keratin 1 OS = Homo sapiens GN = KRT1 PE = 3 SV = 1 | 1.52 | 6.05 | 1.6 |
| O15240 | Neurosecretory protein VGF OS = Homo sapiens GN = VGF PE = 1 SV = 2 | 1.24 | 1.25 | 1.31 |
| Q6IAW5 | CALU protein OS = Homo sapiens GN = CALU PE = 2 SV = 1 | 1.49 | 1.73 | 1.57 |
| P50454 | Serpin H1 OS = Homo sapiens GN = SERPINH1 PE = 1 SV = 2 | 1.45 | 1.49 | 1.4 |
| K7ELL7 | Glucosidase 2 subunit beta OS = Homo sapiens GN = PRKCSH PE = 4 SV = 1 | 1.54 | 1.48 | 1.51 |
| P08758 | Annexin A5 OS = Homo sapiens GN = ANXA5 PE = 1 SV = 2 | 1.44 | 1.4 | 1.34 |
| P07355 | Annexin A2 OS = Homo sapiens GN = ANXA2 PE = 1 SV = 2 | 1.5 | 1.43 | 1.44 |
| G3XAI2 | Laminin subunit beta-1 OS = Homo sapiens GN = LAMB1 PE = 2 SV = 1 | 1.4 | 1.4 | 1.32 |
| A8K401 | Prohibitin, isoform CRA_a OS = Homo sapiens GN = PHB PE = 2 SV = 1 | 1.5 | 1.37 | 1.45 |
| J3KPX7 | Prohibitin-2 OS = Homo sapiens GN = PHB2 PE = 4 SV = 1 | 1.45 | 1.34 | 1.4 |
| Q5TZZ9 | Annexin OS = Homo sapiens GN = ANXA1 PE = 2 SV = 1 | 1.41 | 1.47 | 1.41 |
| P31930 | Cytochrome b–c 1 complex subunit 1, mitochondrial OS = Homo sapiens GN = UQCRC1 PE = 1 SV = 3 | 1.22 | 1.25 | 1.27 |
| Q9BZQ8 | Protein Niban OS = Homo sapiens GN = FAM129A PE = 1 SV = 1 | 1.22 | 1.32 | 1.28 |
| P05141 | ADP/ATP translocase 2 OS = Homo sapiens GN = SLC25A5 PE = 1 SV = 7 | 1.33 | 1.31 | 1.26 |
| Q96A33 | Coiled-coil domain-containing protein 47 OS = Homo sapiens GN = CCDC47 PE = 1 SV = 1 | 1.34 | 1.22 | 1.2 |
| Q8NCF7 | cDNA FLJ90278 fis, clone NT2RP1000325, highly similar to phosphate carrier protein, mitochondrial precursor OS = Homo sapiens PE = 2 SV = 1 | 1.32 | 1.34 | 1.32 |
| P23284 | Peptidyl-prolyl cis–trans isomerase B OS = Homo sapiens GN = PPIB PE = 1 SV = 2 | 1.97 | 2.22 | 1.92 |
| P35908 | Keratin, type II cytoskeletal 2 epidermal OS = Homo sapiens GN = KRT2 PE = 1 SV = 2 | 1.65 | 1.59 | 1.56 |
| J3KPF3 | 4F2 cell-surface antigen heavy chain OS = Homo sapiens GN = SLC3A2 PE = 4 SV = 1 | 1.45 | 1.46 | 1.43 |
| P13521 | Secretogranin-2 OS = Homo sapiens GN = SCG2 PE = 1 SV = 2 | 1.29 | 1.25 | 1.28 |
| Q5U0D2 | Putative uncharacterized protein DKFZp686P11128 OS = Homo sapiens GN = TAGLN PE = 2 SV = 1 | 1.39 | 1.42 | 1.35 |
| A8K878 | cDNA FLJ77177, highly similar to Homo sapiens arginine-rich, mutated in early stage tumors (ARMET), mRNA OS = Homo sapiens PE = 2 SV = 1 | 2 | 1.85 | 1.87 |
| P30040 | Endoplasmic reticulum resident protein 29 OS = Homo sapiens GN = ERP29 PE = 1 SV = 4 | 1.41 | 1.36 | 1.42 |
| Q9BS26 | Endoplasmic reticulum resident protein 44 OS = Homo sapiens GN = ERP44 PE = 1 SV = 1 | 1.49 | 1.51 | 1.34 |
| Q96HE7 | ERO1-like protein alpha OS = Homo sapiens GN = ERO1L PE = 1 SV = 2 | 1.39 | 1.39 | 1.37 |
| Q15293 | Reticulocalbin-1 OS = Homo sapiens GN = RCN1 PE = 1 SV = 1 | 1.56 | 1.51 | 1.44 |
| Q9BRK5 | 45 kDa calcium-binding protein OS = Homo sapiens GN = SDF4 PE = 1 SV = 1 | 1.31 | 1.21 | 1.31 |
| P16401 | Histone H1.5 OS = Homo sapiens GN = HIST1H1B PE = 1 SV = 3 | 1.77 | 2.18 | 1.83 |
| Q9UBS4 | DnaJ homolog subfamily B member 11 OS = Homo sapiens GN = DNAJB11 PE = 1 SV = 1 | 1.51 | 1.5 | 1.52 |
| Q08ET0 | Cell proliferation-inducing protein 47 OS = Homo sapiens GN = hCG_39985 PE = 2 SV = 1 | 1.29 | 1.48 | 1.21 |
| Q8TAS0 | ATP synthase subunit gamma (fragment) OS = Homo sapiens PE = 2 SV = 1 | 1.27 | 1.36 | 1.39 |
| P48047 | ATP synthase subunit O, mitochondrial OS = Homo sapiens GN = ATP5O PE = 1 SV = 1 | 1.39 | 1.3 | 1.33 |
| Q53GF9 | Full-length cDNA 5-PRIME end of clone CS0DF013YM24 of fetal brain of Homo sapiens (human) variant (fragment) OS = Homo sapiens PE = 2 SV = 1 | 1.58 | 1.53 | 1.43 |
| P80303 | Nucleobindin-2 OS = Homo sapiens GN = NUCB2 PE = 1 SV = 2 | 1.53 | 1.32 | 1.32 |
| Q13162 | Peroxiredoxin-4 OS = Homo sapiens GN = PRDX4 PE = 1 SV = 1 | 1.24 | 1.28 | 1.24 |
| O75947 | ATP synthase subunit d, mitochondrial OS = Homo sapiens GN = ATP5H PE = 1 SV = 3 | 1.29 | 1.31 | 1.31 |
| P20674 | Cytochrome c oxidase subunit 5A, mitochondrial OS = Homo sapiens GN = COX5A PE = 1 SV = 2 | 1.33 | 1.42 | 1.22 |
| Q5T0G8 | Annexin OS = Homo sapiens GN = ANXA11 PE = 2 SV = 1 | 1.4 | 1.36 | 1.48 |
| Q14696 | LDLR chaperone MESD OS = Homo sapiens GN = MESDC2 PE = 1 SV = 2 | 1.22 | 1.63 | 1.36 |
| Q96JZ5 | SM-11044 binding protein, isoform CRA_b OS = Homo sapiens GN = SMBP PE = 2 SV = 1 | 1.42 | 1.21 | 1.27 |
| B4DR61 | Protein transport protein Sec61 subunit alpha isoform 1 OS = Homo sapiens GN = SEC61A1 PE = 2 SV = 1 | 1.75 | 2.04 | 1.71 |
| Q53XJ5 | Peptidyl-prolyl cis–trans isomerase OS = Homo sapiens GN = FKBP2 PE = 2 SV = 1 | 1.39 | 1.44 | 1.37 |
| B7Z5L4 | cDNA FLJ61340, highly similar to Homo sapiens seizure related 6 homolog-like 2 (SEZ6L2), transcript variant 2, mRNA OS = Homo sapiens PE = 2 SV = 1 | 1.62 | 1.72 | 1.76 |
| Q5T8U7 | Surfeit 4 OS = Homo sapiens GN = SURF4 PE = 2 SV = 1 | 1.57 | 1.26 | 1.43 |
| Q8TCT9 | Minor histocompatibility antigen H13 OS = Homo sapiens GN = HM13 PE = 1 SV = 1 | 1.54 | 1.2 | 1.25 |
| P51970 | NADH dehydrogenase [ubiquinone] 1 alpha subcomplex subunit 8 OS = Homo sapiens GN = NDUFA8 PE = 1 SV = 3 | 1.34 | 1.39 | 1.27 |
| Q6FHT8 | RNP24 protein OS = Homo sapiens GN = RNP24 PE = 2 SV = 1 | 1.37 | 1.37 | 1.26 |
| O75367 | Core histone macro-H2A.1 OS = Homo sapiens GN = H2AFY PE = 1 SV = 4 | 1.79 | 1.85 | 1.72 |
| B4DVE1 | cDNA FLJ53478, highly similar to Galectin-3-binding protein OS = Homo sapiens PE = 2 SV = 1 | 1.37 | 1.39 | 1.25 |
| O15173 | Membrane-associated progesterone receptor component 2 OS = Homo sapiens GN = PGRMC2 PE = 1 SV = 1 | 1.32 | 1.34 | 1.29 |
| Q16222 | UDP-N-acetylhexosamine pyrophosphorylase OS = Homo sapiens GN = UAP1 PE = 1 SV = 3 | 1.68 | 1.58 | 1.4 |
| A8KA82 | DnaJ (Hsp40) homolog, subfamily C, member 3 OS = Homo sapiens GN = DNAJC3 PE = 2 SV = 1 | 1.51 | 1.52 | 1.39 |
| Q9BRR6 | ADP-dependent glucokinase OS = Homo sapiens GN = ADPGK PE = 1 SV = 1 | 1.32 | 1.43 | 1.22 |
| P31949 | Protein S100-A11 OS = Homo sapiens GN = S100A11 PE = 1 SV = 2 | 1.5 | 1.44 | 1.47 |
| Q8NI22 | Multiple coagulation factor deficiency protein 2 OS = Homo sapiens GN = MCFD2 PE = 1 SV = 1 | 1.33 | 1.23 | 1.2 |
| P30049 | ATP synthase subunit delta, mitochondrial OS = Homo sapiens GN = ATP5D PE = 1 SV = 2 | 1.3 | 1.61 | 1.62 |
| Q9NZ45 | CDGSH iron–sulfur domain-containing protein 1 OS = Homo sapiens GN = CISD1 PE = 1 SV = 1 | 1.32 | 1.3 | 1.27 |
| E9PN17 | ATP synthase subunit g, mitochondrial OS = Homo sapiens GN = ATP5L PE = 2 SV = 1 | 1.45 | 1.42 | 1.44 |
| Q9Y3A6 | Transmembrane emp24 domain-containing protein 5 OS = Homo sapiens GN = TMED5 PE = 1 SV = 1 | 1.59 | 1.63 | 1.73 |
| P07305 | Histone H1.0 OS = Homo sapiens GN = H1F0 PE = 1 SV = 3 | 2.34 | 1.73 | 1.99 |
| P35610 | Sterol O-acyltransferase 1 OS = Homo sapiens GN = SOAT1 PE = 1 SV = 3 | 1.21 | 1.38 | 1.35 |
| H0Y886 | NADH dehydrogenase [ubiquinone] 1 beta subcomplex subunit 5, mitochondrial (fragment) OS = Homo sapiens GN = NDUFB5 PE = 4 SV = 1 | 1.57 | 1.8 | 1.3 |
| Q9NYB0 | Telomeric repeat-binding factor 2-interacting protein 1 OS = Homo sapiens GN = TERF2IP PE = 1 SV = 1 | 1.27 | 1.66 | 2.73 |
| G3V325 | Pentatricopeptide repeat-containing protein 1, mitochondrial OS = Homo sapiens GN = ATP5J2-PTCD1 PE = 4 SV = 1 | 1.47 | 1.28 | 1.43 |
| A8K0F7 | cDNA FLJ76587, highly similar to Homo sapiens vitamin K epoxide reductase complex, subunit 1-like 1 (VKORC1L1), mRNA OS = Homo sapiens PE = 2 SV = 1 | 1.33 | 1.36 | 1.24 |
| B2R8A2 | cDNA, FLJ93804, highly similar to Homo sapiens gp25L2 protein (HSGP25L2G), mRNA OS = Homo sapiens PE = 2 SV = 1 | 1.41 | 1.53 | 1.45 |
| Q6IBU4 | SDF2 protein OS = Homo sapiens GN = SDF2 PE = 2 SV = 1 | 1.36 | 1.23 | 1.68 |
| Q71UI9 | Histone H2A.V OS = Homo sapiens GN = H2AFV PE = 1 SV = 3 | 2.77 | 2.62 | 2.34 |
| Q6IAM7 | SPC18 protein OS = Homo sapiens GN = SPC18 PE = 2 SV = 1 | 1.69 | 1.21 | 1.35 |
| Q13425 | Beta-2-syntrophin OS = Homo sapiens GN = SNTB2 PE = 1 SV = 1 | 1.37 | 1.49 | 1.21 |
| Q9UDW1 | Cytochrome b–c 1 complex subunit 9 OS = Homo sapiens GN = UQCR10 PE = 1 SV = 3 | 1.58 | 1.41 | 1.58 |
| P05204 | Non-histone chromosomal protein HMG-17 OS = Homo sapiens GN = HMGN2 PE = 1 SV = 3 | 1.76 | 4.21 | 1.29 |
| P68431 | Histone H3.1 OS = Homo sapiens GN = HIST1H3A PE = 1 SV = 2 | 6.1 | 3.9 | 9.3 |
| A8K2Q6 | Peptidyl-prolyl cis–trans isomerase OS = Homo sapiens PE = 2 SV = 1 | 1.57 | 1.37 | 1.37 |
| Q6I9V5 | SLC25A6 protein OS = Homo sapiens GN = SLC25A6 PE = 2 SV = 1 | 1.33 | 1.54 | 1.35 |
| Q5U0C3 | RAP1A, member of RAS oncogene family OS = Homo sapiens PE = 2 SV = 1 | 1.25 | 1.49 | 1.25 |
| Q71DI3 | Histone H3.2 OS = Homo sapiens GN = HIST2H3A PE = 1 SV = 3 | 5.06 | 4.58 | 4.16 |
| Down regulated | Fold change: Tm/control | |||
|---|---|---|---|---|
| Acession ID | Description | Rep 1 | Rep 2 | Rep 3 |
| J3KTA4 | Probable ATP-dependent RNA helicase DDX5 OS = Homo sapiens GN = DDX5 PE = 3 SV = 1 | 0.67 | 0.69 | 0.71 |
| Q14566 | DNA replication licensing factor MCM6 OS = Homo sapiens GN = MCM6 PE = 1 SV = 1 | 0.75 | 0.77 | 0.74 |
| P31689 | DnaJ homolog subfamily A member 1 OS = Homo sapiens GN = DNAJA1 PE = 1 SV = 2 | 0.76 | 0.73 | 0.77 |
| Q9UNF1 | Melanoma-associated antigen D2 OS = Homo sapiens GN = MAGED2 PE = 1 SV = 2 | 0.8 | 0.76 | 0.66 |
| Q96T88 | E3 ubiquitin-protein ligase UHRF1 OS = Homo sapiens GN = UHRF1 PE = 1 SV = 1 | 0.74 | 0.75 | 0.77 |
| Q8IV08 | Phospholipase D3 OS = Homo sapiens GN = PLD3 PE = 1 SV = 1 | 0.69 | 0.67 | 0.74 |
| Q9Y5L4 | Mitochondrial import inner membrane translocase subunit Tim13 OS = Homo sapiens GN = TIMM13 PE = 1 SV = 1 | 0.76 | 0.72 | 0.79 |
| Q7Z7L8 | Uncharacterized protein C11orf96 OS = Homo sapiens GN = C11orf96 PE = 1 SV = 3 | 0.76 | 0.61 | 0.53 |
| Q92558 | Wiskott–Aldrich syndrome protein family member 1 OS = Homo sapiens GN = WASF1 PE = 1 SV = 1 | 0.79 | 0.64 | 0.78 |
| O75243 | R30783_1 OS = Homo sapiens PE = 4 SV = 1 | 0.69 | 0.68 | 0.8 |
| Q7Z7K6 | Centromere protein V OS = Homo sapiens GN = CENPV PE = 1 SV = 1 | 0.8 | 0.44 | 0.28 |
| Q2L6I0 | FB19 protein OS = Homo sapiens GN = PPP1R10 PE = 2 SV = 1 | 0.8 | 0.8 | 0.78 |
| B2R4N3 | cDNA, FLJ92155, highly similar to Homo sapiens ubiquitin-like 5 (UBL5), mRNA OS = Homo sapiens PE = 4 SV = 1 | 0.74 | 0.61 | 0.5 |
| Q9Y6H1 | Coiled-coil-helix–coiled-coil-helix domain-containing protein 2, mitochondrial OS = Homo sapiens GN = CHCHD2 PE = 1 SV = 1 | 0.76 | 0.58 | 0.76 |
| Q59EN5 | Prosaposin variant (fragment) OS = Homo sapiens PE = 2 SV = 1 | 0.75 | 0.51 | 0.5 |
| A9CQZ4 | Dihydropyrimidinase-like 2 long form (fragment) OS = Homo sapiens GN = DPYSL2 PE = 2 SV = 1 | 0.71 | 0.67 | 0.77 |
| Q9UMZ1 | Prothymosin a14 OS = Homo sapiens PE = 1 SV = 1 | 0.72 | 0.77 | 0.48 |
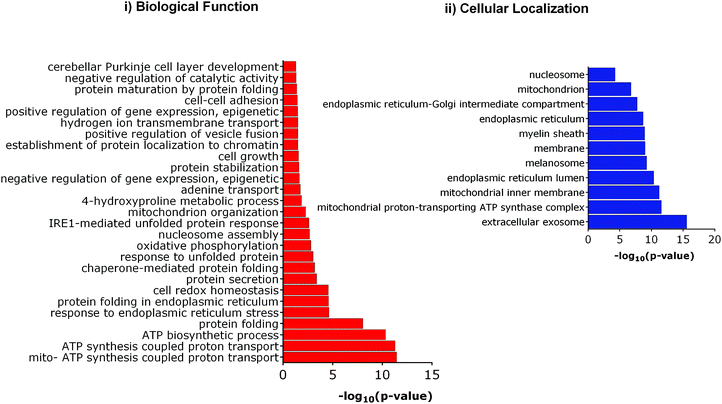 | ||
| Fig. 3 Gene ontology (GO) analysis of Tm induced differentially expressed proteins. (i) Enriched biological functions, (ii) enriched cellular compartment. | ||
4-PBA is known to alleviate ER stress induced by Tm. We thus asked if 4-PBA could upregulate the protein quality control machinery of ER to help survival during ER stress. We then assessed the proteins that are modulated by 4-PBA alone (in the absence of Tm). We found that 4-PBA-treatment did not alter the proteome significantly as only one protein, Tmed5 – Transmembrane emp24 domain-containing protein, was up-regulated and two proteins, Osgep-(Probable tRNA threonylcarbamoyladenosine biosynthesis protein) and Tspo (Putative peripheral benzodiazepine receptor-related protein), were downregulated. These proteins that are differentially expressed upon treatment with 4-PBA are not bonafide members of the protein quality control machinery in the ER. Thus, it is apparent that 4-PBA treatment does not alter the expression of proteins required for maintaining ER homeostasis.
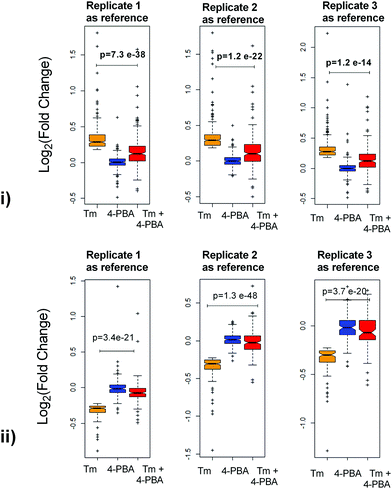
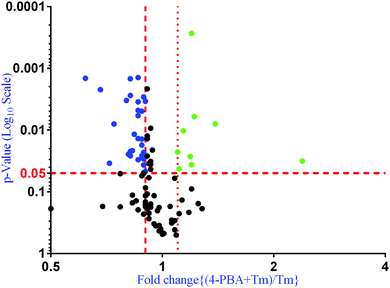
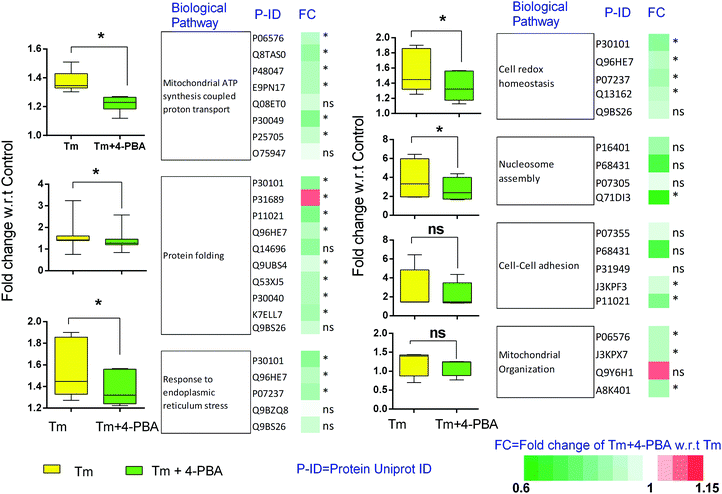
To check if addition of 4-PBA had an effect on the expression of proteins that were modulated by Tm, we performed a Jack-knife resampling analysis30 to investigate the significance of changes induced by 4-PBA during ER stress. For this analysis, proteins that were differentially expressed by Tm treatment in one replicate (fold change ≥1.2 for upregulated proteins, ≤0.8 for downregulated proteins) were taken and the average ratio of (4-PBA + Tm)/control and 4-PBA/control for these proteins in the other two replicates was calculated. This was repeated for all the replicates using the first, second and third replicate as a reference point and a box plot of the fold change was plotted (Fig. 4). In all the three replicates the average ratio of Tm/control was significantly different from (4-PBA + Tm)/control for both up- and downregulated proteins during Tm-treatment. This indicates that on average the expression of proteins induced or repressed by Tm could be reverted to near control levels. Among the Tm-induced differentially expressed proteins, a list of proteins whose expression was significantly reverted by 4-PBA was identified by considering proteins whose fold change for (4-PBA + Tm)/Tm is ≥1.1 or ≤0.9 with p-value ≤0.05 (Table 2 and Fig. 5). Most of the pathways altered by Tm were recovered by 4-PBA (Fig. 6). Both ER quality control and mitochondrial respiration-related proteins were suppressed by 4-PBA during ER stress. Thus the study demonstrates that 4-PBA has a global protective effect on proteomic alterations induced by Tm.
| Acession ID | Description | Fold change (Tm + 4-PBA)/Tm | p-Value | Status in Tm treatment |
|---|---|---|---|---|
| P11021 | 78 kDa glucose-regulated protein OS = Homo sapiens GN = HSPA5 PE = 1 SV = 2 | 0.8 | 3.31 × 10−3 | UP |
| P30101 | Protein disulfide-isomerase A3 OS = Homo sapiens GN = PDIA3 PE = 1 SV = 4 | 0.82 | 2.75 × 10−3 | UP |
| P13667 | Protein disulfide-isomerase A4 OS = Homo sapiens GN = PDIA4 PE = 1 SV = 2 | 0.82 | 1.47 × 10−3 | UP |
| P07237 | Protein disulfide-isomerase OS = Homo sapiens GN = P4HB PE = 1 SV = 3 | 0.86 | 1.41 × 10−3 | UP |
| P06576 | ATP synthase subunit beta, mitochondrial OS = Homo sapiens GN = ATP5B PE = 1 SV = 3 | 0.89 | 2.30 × 10−2 | UP |
| B4DGP8 | Calnexin OS = Homo sapiens GN = CANX PE = 2 SV = 1 | 0.86 | 4.80 × 10−3 | UP |
| Q6IAW5 | CALU protein OS = Homo sapiens GN = CALU PE = 2 SV = 1 | 0.89 | 4.91 × 10−2 | UP |
| K7ELL7 | Glucosidase 2 subunit beta OS = Homo sapiens GN = PRKCSH PE = 4 SV = 1 | 0.86 | 1.37 × 10−2 | UP |
| A8K401 | Prohibitin, isoform CRA_a OS = Homo sapiens GN = PHB PE = 2 SV = 1 | 0.86 | 5.85 × 10−3 | UP |
| J3KPX7 | Prohibitin-2 OS = Homo sapiens GN = PHB2 PE = 4 SV = 1 | 0.89 | 2.94 × 10−2 | UP |
| P05141 | ADP/ATP translocase 2 OS = Homo sapiens GN = SLC25A5 PE = 1 SV = 7 | 0.9 | 3.42 × 10−3 | UP |
| P23284 | Peptidyl-prolyl cis–trans isomerase BOS = Homo sapiens GN = PPIB PE = 1 SV = 2 | 0.82 | 2.61 × 10−2 | UP |
| A8K878 | cDNA FLJ77177, highly similar to Homo sapiens arginine-rich, mutated in early stage tumors (ARMET), mRNA OS = Homo sapiens PE = 2 SV = 1 | 0.82 | 2.22 × 10−2 | UP |
| P30040 | Endoplasmic reticulum resident protein 29 OS = Homo sapiens GN = ERP29 PE = 1 SV = 4 | 0.88 | 4.95 × 10−3 | UP |
| Q96HE7 | ERO1-like protein alpha OS = Homo sapiens GN = ERO1L PE = 1 SV = 2 | 0.89 | 2.91 × 10−3 | UP |
| Q15293 | Reticulocalbin-1 OS = Homo sapiens GN = RCN1 PE = 1 SV = 1 | 0.9 | 4.65 × 10−2 | UP |
| Q8TAS0 | ATP synthase subunit gamma (fragment) OS = Homo sapiens PE = 2 SV = 1 | 0.84 | 1.17 × 10−2 | UP |
| P48047 | ATP synthase subunit O, mitochondrial OS = Homo sapiens GN = ATP5O PE = 1 SV = 1 | 0.88 | 1.75 × 10−2 | UP |
| Q53GF9 | Full-length cDNA 5-PRIME end of clone CS0DF013YM24 of fetal brain of Homo sapiens (human) variant (fragment) OS = Homo sapiens PE = 2 SV = 1 | 0.9 | 2.68 × 10−2 | UP |
| Q13162 | Peroxiredoxin-4 OS = Homo sapiens GN = PRDX4 PE = 1 SV = 1 | 0.9 | 2.68 × 10−2 | UP |
| P20674 | Cytochrome c oxidase subunit 5A, mitochondrial OS = Homo sapiens GN = COX5A PE = 1 SV = 2 | 0.83 | 2.16 × 10−2 | UP |
| B4DR61 | Protein transport protein Sec61 subunit alpha isoform 1 OS = Homo sapiens GN = SEC61A1 PE = 2 SV = 1 | 0.72 | 3.45 × 10−2 | UP |
| Q53XJ5 | Peptidyl-prolyl cis–trans isomerase OS = Homo sapiens GN = FKBP2 PE = 2 SV = 1 | 0.88 | 1.40 × 10−2 | UP |
| P51970 | NADH dehydrogenase [ubiquinone] 1 alpha subcomplex subunit 8 OS = Homo sapiens GN = NDUFA8 PE = 1 SV = 3 | 0.86 | 3.50 × 10−3 | UP |
| O75367 | Core histone macro-H2A.1 OS = Homo sapiens GN = H2AFY PE = 1 SV = 4 | 0.74 | 7.97 × 10−3 | UP |
| O15173 | Membrane-associated progesterone receptor component 2 OS = Homo sapiens GN = PGRM C2 PE = 1 SV = 1 | 0.86 | 3.03 × 10−2 | UP |
| P30049 | ATP synthase subunit delta, mitochondrial OS = Homo sapiens GN = ATP5D PE = 1 SV = 2 | 0.81 | 2.47 × 10−2 | UP |
| E9PN17 | ATP synthase subunit g, mitochondrial OS = Homo sapiens GN = ATP5L PE = 2 SV = 1 | 0.88 | 2.61 × 10−2 | UP |
| B2R8A2 | cDNA, FLJ93804, highly similar to Homo sapiens sgp25L2 protein (HSGP25L2G), mRNA OS = Homo sapiens PE = 2 SV = 1 | 0.89 | 3.67 × 10−2 | UP |
| Q71UI9 | Histone H2A.V OS = Homo sapiens GN = H2AFV PE = 1 SV = 3 | 0.68 | 2.21 × 10−3 | UP |
| Q71DI3 | Histone H3.2 OS = Homo sapiens GN = HIST2H3A PE = 1 SV = 3 | 0.62 | 1.45 × 10−3 | UP |
| J3KTA4 | Probable ATP-dependent RNA helicase DDX5 OS = Homo sapiens GN = DDX5 PE = 3 SV = 1 | 1.14 | 1.03 × 10−2 | Down |
| Q14566 | DNA replication licensing factor MCM6 OS = Homo sapiens GN = MCM6 PE = 1 SV = 1 | 1.1 | 2.26 × 10−2 | Down |
| P31689 | DnaJ homolog subfamily A member 1 OS = Homo sapiens GN = DNAJA1 PE = 1 SV = 2 | 1.11 | 4.24 × 10−2 | Down |
| Q9UNF1 | Melanoma-associated antigen D2 OS = Homo sapiens GN = MAGED2 PE = 1 SV = 2 | 1.19 | 2.67 × 10−2 | Down |
| Q96T88 | E3 ubiquitin-protein ligase UHRF 1 OS = Homo sapiens GN = UHRF1 PE = 1 SV = 1 | 1.2 | 2.71 × 10−4 | Down |
| Q8IV08 | Phospholipase D3 OS = Homo sapiens GN = PLD3 PE = 1 SV = 1 | 1.2 | 3.64 × 10−2 | Down |
| O75243 | R30783_1 OS = Homo sapiens PE = 4 SV = 1 | 1.39 | 7.94 × 10−3 | Down |
| Q7Z7K6 | Centromere protein V OS = Homo sapiens GN = CENPV PE = 1 SV = 1 | 2.39 | 3.17 × 10−2 | Down |
| Q2L6I0 | FB19 protein OS = Homo sapiens GN = PPP1R10 PE = 2 SV = 1 | 1.22 | 6.04 × 10−3 | Down |
4. Discussion
ER stress is associated with various neurodegenerative diseases like Alzheimer's disease, Huntington's disease (HD) and Parkinson's disease (PD).33–35 4-PBA, a small molecule, is known to alleviate ER stress. Several reports have demonstrated the therapeutic potential of 4-PBA in different neurological diseases.21,26,36,37 However, the molecular basis and the breadth of its therapeutic effects are still not clear. In this study using the iTRAQ based global proteomics approach, we intended to identify the global effects of 4-PBA during ER stress in neuronal cells. Using iTRAQ based relative quantitative proteomics, we were able to relatively quantitate the expression of 2023 proteins, 88% of which were under 20% CV across three biological replicates.Tm is a well-known ER stress inducer; it inhibits N-linked glycosylation and leads to accumulation of unfolded glycoproteins in the ER, causing ER stress38,39 The proteomic profile of Tm treated neuronal cells is in agreement with previous reports. We found enhanced accumulation of proteins involved in protein folding and cellular redox homeostasis.32 Various studies have indicated the cross-talk between ER and mitochondria under stress conditions.28,40,41 In the present study, we found that most of the Tm-induced differentially expressed proteins localize in the ER or mitochondria. Enrichment of a large number of mitochondria or ER resident proteins further strengthens the known crosstalk between the two compartments during ER stress.28,40,41 The ER quality control machinery depends upon metabolic energy for proper folding and clearance of misfolded proteins.42,43 Bravo et al. demonstrated that there is an increase in mitochondrial respiration during ER stress. This was found to be an adaptive response.44 When ER stress is not resolved it leads to cell death by inducing mitochondrial dysfunction, and hence, the enrichment of mitochondrial proteins involved in energy synthesis, in our data, underlines the role of mitochondrial respiration in ER stress. However, this does not exclude a more complex and direct cross-talk between ER and mitochondria.
4-PBA reduces Tm-induced cell death and decreases the expression of UPR markers (GRP78 and CHOP). 4-PBA alone does not alter the quality control machinery of ER but in the presence of an ER stress inducer, it restores the altered ER stress induced expression of proteins towards unstressed levels. Most of the pathways affected by Tm were recovered by 4-PBA. 4-PBA suppresses the expression of UPR genes and toxicity induced by Tm and thus directs towards the possibility that 4-PBA works by decreasing ER stress instead of preconditioning the ER to cope better with stress. 4-PBA not only decreases the expression of UPR genes, but also recovers the expression of proteins involved in mitochondrial ATP synthesis. ER stress-induced cell death involves increased mitochondrial respiration, followed by apoptosis.44 Our study reveals that 4-PBA recovers the Tm-induced upregulation of proteins involved in both ER stress and mitochondrial respiration.
4-PBA has an HDAC (Histone deacetylase) inhibitor activity. It thus has the ability to alter the expression of genes which involve aberrant histone acetylation during neurological disorders.23,26,27 In a previous report by Mimori et al.,45 it was shown using structural analogs of 4-PBA that protection was indeed correlated with the in vitro chaperoning activity of the molecule and not HDAC7 binding activity. However, it could not exclude if 4-PBA activated other pathways to protect cells against UPR. Using a global measure of cellular response we support the view proposed by Mimori et al. that 4-PBA indeed does not have a protective effect by upregulating alternate protective pathways to combat ER stress. Its activity most likely is an outcome of its chaperoning activity.
In conclusion, our study demonstrates the first proteomic profile of 4-PBA during Tm treatment in human neuroblastoma cells. This study illustrates that 4-PBA exhibits a global recovery from the proteomic alterations induced by Tm but does not alter the cellular proteome to adapt towards ER stress. This supports the suspected role of 4-PBA as a bonafide chemical chaperone and suggests that 4-PBA may aid in protein folding of ER resident proteins to alleviate ER proteotoxicity.
Abbreviations
| 4-PBA | Sodium 4-phenylbutyrate |
| ALS | Amyotrophic lateral sclerosis |
| ATCC | American Type Culture Collection |
| ATF4 | Activating transcription factor 4 |
| ATF6 | Activating transcription factor 6 |
| ACN | Acetonitrile |
| CHOP | CCAAT-enhancer-binding protein homologous protein |
| % CV | Percentage coefficient of variation |
| DAVID | Database for Annotation, Visualization and Integrated Discovery |
| DMEM | Dulbecco's modified Eagle's medium |
| ER | Endoplasmic reticulum |
| FBS | Fetal bovine serum |
| FDA | Food and Drug Administration |
| FDR | False discovery rate |
| GO | Gene ontology |
| GRP78 | Glucose-regulated protein 78 |
| HDAC | Histone deacetylase |
| HRP | Horseradish peroxidase |
| IAA | Iodoacetamide |
| IDA | Information dependent acquisition |
| IRE1 | Inositol-requiring enzyme 1 |
| iTRAQ | Isobaric tags for relative and absolute quantitation |
| PERK | Protein kinase R (PKR)-like endoplasmic reticulum kinase |
| PVDF | Polyvinylidene fluoride |
| SCX | Strong cation exchange |
| SDS-PAGE | Sodium dodecyl sulfate polyacrylamide gel electrophoresis |
| Tm | Tunicamycin |
| TOF | Time-of-flight |
| UPR | Unfolded protein response |
| XBP1 | X-box binding protein 1 |
Conflicts of interest
There are no conflicts to declare.Acknowledgements
We acknowledge the financial support from the Council of Scientific and Industrial Research (CSIR), India. The study was funded under the project titled ‘CARDIOMED: Centre for Cardiovascular and Metabolic Disease Research (BSC0122)’. R. C. acknowledges the Junior Research Fellowship from UGC.References
- S. Fulda, A. M. Gorman, O. Hori and A. Samali, Int. J. Cell Biol., 2010, 2010, 214074 Search PubMed.
- R. J. Kaufman, D. Scheuner, M. Schroder, X. Shen, K. Lee, C. Y. Liu and S. M. Arnold, Nat. Rev. Mol. Cell Biol., 2002, 3, 411–421 CrossRef CAS PubMed.
- D. Ron and P. Walter, Nat. Rev. Mol. Cell Biol., 2007, 8, 519–529 CrossRef CAS PubMed.
- B. C. Yoo, K. Krapfenbauer, N. Cairns, G. Belay, M. Bajo and G. Lubec, Neurosci. Lett., 2002, 334, 196–200 CrossRef CAS PubMed.
- I. Kim, W. Xu and J. C. Reed, Nat. Rev. Drug Discovery, 2008, 7, 1013–1030 CrossRef CAS PubMed.
- R. V. Rao, H. M. Ellerby and D. E. Bredesen, Cell Death Differ., 2004, 11, 372–380 CrossRef CAS PubMed.
- J. D. Malhotra and R. J. Kaufman, Antioxid. Redox Signaling, 2007, 9, 2277–2293 CrossRef CAS PubMed.
- S. H. Back and R. J. Kaufman, Annu. Rev. Biochem., 2012, 81, 767–793 CrossRef CAS PubMed.
- C. Mozzini, L. Cominacini, U. Garbin and A. M. FrattaPasini, Curr. Atheroscler. Rep., 2017, 19, 33 CrossRef PubMed.
- C. Haass and D. J. Selkoe, Nat. Rev. Mol. Cell Biol., 2007, 8, 101–112 CrossRef CAS PubMed.
- V. N. Uversky, J. Neurochem., 2007, 103, 17–37 CAS.
- A. Bellucci, L. Navarria, M. Zaltieri, E. Falarti, S. Bodei, S. Sigala, L. Battistin, M. Spillantini, C. Missale and P. Spano, J. Neurochem., 2011, 116, 588–605 CrossRef CAS PubMed.
- S. L. Lindquist and J. W. Kelly, Cold Spring Harbor Perspect. Biol., 2011, 3, a004507 Search PubMed.
- H. Y. Zhang, Z. G. Wang, X. H. Lu, X. X. Kong, F. Z. Wu, L. Lin, X. Tan, L. B. Ye and J. Xiao, Mol. Neurobiol., 2015, 51, 1343–1352 CrossRef CAS PubMed.
- F. U. Hartl, Nature, 1996, 381, 571–579 CrossRef CAS PubMed.
- S. W. Fewell, K. J. Travers, J. S. Weissman and J. L. Brodsky, Annu. Rev. Genet., 2001, 35, 149–191 CrossRef CAS PubMed.
- J. Klucken, Y. Shin, E. Masliah, B. T. Hyman and P. J. McLean, J. Biol. Chem., 2004, 279, 25497–25502 CrossRef CAS PubMed.
- M. Gaestel, Molecular chaperones in health and disease, Springer, Berlin, New York, 2006 Search PubMed.
- L. Cortez and V. Sim, Prion, 2014, 8, 197–202 CrossRef CAS.
- N. E. Maestri, S. W. Brusilow, D. B. Clissold and S. S. Bassett, N. Engl. J. Med., 1996, 335, 855–859 CrossRef CAS PubMed.
- G. Gardian, S. E. Browne, D. K. Choi, P. Klivenyi, J. Gregorio, J. K. Kubilus, H. Ryu, B. Langley, R. R. Ratan, R. J. Ferrante and M. F. Beal, J. Biol. Chem., 2005, 280, 556–563 CrossRef CAS PubMed.
- H. Ryu, K. Smith, S. I. Camelo, I. Carreras, J. Lee, A. H. Iglesias, F. Dangond, K. A. Cormier, M. E. Cudkowicz, R. H. Brown, Jr. and R. J. Ferrante, J. Neurochem., 2005, 93, 1087–1098 CrossRef CAS PubMed.
- A. Ricobaraza, M. Cuadrado-Tejedor, A. Perez-Mediavilla, D. Frechilla, J. Del Rio and A. Garcia-Osta, Neuropsychopharmacology, 2009, 34, 1721–1732 CrossRef CAS PubMed.
- R. C. Rubenstein and P. L. Zeitlin, Am. J. Physiol.: Cell Physiol., 2000, 278, C259–267 CrossRef CAS PubMed.
- K. Kubota, Y. Niinuma, M. Kaneko, Y. Okuma, M. Sugai, T. Omura, M. Uesugi, T. Uehara, T. Hosoi and Y. Nomura, J. Neurochem., 2006, 97, 1259–1268 CrossRef CAS PubMed.
- D. M. Chuang, Y. Leng, Z. Marinova, H. J. Kim and C. T. Chiu, Trends Neurosci., 2009, 32, 591–601 CrossRef CAS PubMed.
- N. A. Shein and E. Shohami, Mol. Med., 2011, 17, 448–456 CAS.
- S. Maity, T. Basak, A. Bhat, N. Bhasin, A. Ghosh, K. Chakraborty and S. Sengupta, Proteomics, 2014, 14, 1724–1736 CrossRef CAS PubMed.
- T. Basak, A. Bhat, D. Malakar, M. Pillai and S. Sengupta, Mol. BioSyst., 2015, 11, 2135–2143 RSC.
- S. Maity, A. Rajkumar, L. Matai, A. Bhat, A. Ghosh, G. Agam, S. Kaur, N. R. Bhatt, A. Mukhopadhyay, S. Sengupta and K. Chakraborty, Cell Rep., 2016, 16, 851–865 CrossRef CAS PubMed.
- S. Mimori, Y. Okuma, M. Kaneko, K. Kawada, T. Hosoi, K. Ozawa, Y. Nomura and H. Hamana, Biol. Pharm. Bull., 2012, 35, 84–90 CAS.
- V. H. Bull and B. Thiede, Electrophoresis, 2012, 33, 1814–1823 CrossRef CAS PubMed.
- Y. Jia, T. J. Jucius, S. A. Cook and S. L. Ackerman, J. Neurosci., 2015, 35, 3001–3009 CrossRef PubMed.
- E. R. Perri, C. J. Thomas, S. Parakh, D. M. Spencer and J. D. Atkin, Front. Cell Dev. Biol., 2015, 3, 80 Search PubMed.
- N. T. Sprenkle, S. G. Sims, C. L. Sanchez and G. P. Meares, Mol. Neurodegener., 2017, 12, 42 CrossRef PubMed.
- S. J. Del Signore, D. J. Amante, J. Kim, E. C. Stack, S. Goodrich, K. Cormier, K. Smith, M. E. Cudkowicz and R. J. Ferrante, Amyotrophic Lateral Scler., 2009, 10, 85–94 CrossRef CAS PubMed.
- A. Ricobaraza, M. Cuadrado-Tejedor, S. Marco, I. Perez-Otano and A. Garcia-Osta, Hippocampus, 2012, 22, 1040–1050 CrossRef CAS PubMed.
- F. Del Grosso, M. De Mariano, L. Passoni, R. Luksch, G. P. Tonini and L. Longo, BMC Cancer, 2011, 11, 525 CrossRef CAS PubMed.
- M. F. Beers, M. Zhao, Y. Tomer, S. J. Russo, P. Zhang, L. W. Gonzales, S. H. Guttentag and S. Mulugeta, Am. J. Physiol.: Lung Cell. Mol. Physiol., 2013, 305, L970–980 CrossRef CAS PubMed.
- T. Hayashi and T. P. Su, Cell, 2007, 131, 596–610 CrossRef CAS PubMed.
- D. Senft and Z. A. Ronai, Trends Biochem. Sci., 2015, 40, 141–148 CrossRef CAS PubMed.
- J. Hoseki, R. Ushioda and K. Nagata, J. Biochem., 2010, 147, 19–25 CrossRef CAS PubMed.
- R. Bravo, V. Parra, D. Gatica, A. E. Rodriguez, N. Torrealba, F. Paredes, Z. V. Wang, A. Zorzano, J. A. Hill, E. Jaimovich, A. F. Quest and S. Lavandero, Int. Rev. Cell Mol. Biol., 2013, 301, 215–290 CAS.
- R. Bravo, J. M. Vicencio, V. Parra, R. Troncoso, J. P. Munoz, M. Bui, C. Quiroga, A. E. Rodriguez, H. E. Verdejo, J. Ferreira, M. Iglewski, M. Chiong, T. Simmen, A. Zorzano, J. A. Hill, B. A. Rothermel, G. Szabadkai and S. Lavandero, J. Cell Sci., 2011, 124, 2143–2152 CrossRef CAS PubMed.
- S. Mimori, H. Ohtaka, Y. Koshikawa, K. Kawada, M. Kaneko, Y. Okuma, Y. Nomura, Y. Murakami and H. Hamana, Bioorg. Med. Chem. Lett., 2013, 23, 6015–6018 CrossRef CAS PubMed.
Footnotes |
| † Electronic supplementary information (ESI) available. See DOI: 10.1039/c7mo00114b |
| ‡ Equal contribution. |
| This journal is © The Royal Society of Chemistry 2018 |