Effect of trehalose polymer regioisomers on protein stabilization†
Abstract
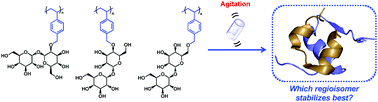
There is considerable interest in the use of proteins as therapeutics and as chemical and biochemical reagents. However, many proteins are unstable and aggregate when exposed to stressors, including increased temperature, pH change, agitation, and desiccation. Polymers with side chain trehalose units were shown to be effective protein stabilizers, preventing aggregation and prolonging activity. Herein, we report the synthesis and characterization of four trehalose regioisomers containing a vinylbenzyl ether moiety at either the 2-O, 3-O, 4-O, or 6-O position. Computational analysis of these regioisomers suggested that they differ in their conformational flexibility, but all retained the native clam shell conformation of trehalose. Polymers were synthesized from the monomers separately via free radical polymerization and one polymer was prepared containing all of the regioisomers. The polymers were tested for their ability to stabilize insulin, and were found to prevent agitation-induced aggregation comparably. The results show that for insulin the effect of trehalose positional modification is minimal and suggest that the clam shell conformation itself may be more important than the polymer backbone attachment site for stabilization of proteins.
- This article is part of the themed collection: Celebrating Excellence in Research: 100 Women of Chemistry


 Please wait while we load your content...
Please wait while we load your content...