Open Access Article
Open Access ArticleProteomic and genetic analysis of the response of S. cerevisiae to soluble copper leads to improvement of the antimicrobial function of cellulosic copper nanoparticles†
Xiaoqing
Rong-Mullins‡
 a,
Matthew J.
Winans‡
a,
Matthew J.
Winans‡
 a,
Justin B.
Lee
a,
Zachery R.
Lonergan§
a,
Justin B.
Lee
a,
Zachery R.
Lonergan§
 a,
Vincent A.
Pilolli¶
a,
Vincent A.
Pilolli¶
 a,
Lyndsey M.
Weatherly
a,
Thomas W.
Carmenzind
b,
Lihua
Jiang
c,
Jonathan R.
Cumming
a,
Lyndsey M.
Weatherly
a,
Thomas W.
Carmenzind
b,
Lihua
Jiang
c,
Jonathan R.
Cumming
 a,
Gloria S.
Oporto
a,
Gloria S.
Oporto
 d and
Jennifer E. G.
Gallagher
d and
Jennifer E. G.
Gallagher
 *a
*a
aDepartment of Biology, West Virginia University, Morgantown, WV, USA. E-mail: jegallagher@mail.wvu.edu
bDepartment of Computer Science, Stanford University, Stanford, CA, USA
cDepartment of Genetics, Stanford University, Stanford University, Stanford, CA, USA
dDivision of Forestry and Natural Resources, West Virginia University, Morgantown, WV, USA
First published on 18th August 2017
Abstract
Copper (Cu) was used in antiquity to prevent waterborne and food diseases because, as a broad-spectrum antimicrobial agent, it generates reactive oxygen species, ROS. New technologies incorporating Cu into low-cost biodegradable nanomaterials built on cellulose, known as cellulosic cupric nanoparticles or c-CuNPs, present novel approaches to deliver Cu in a controlled manner to control microbial growth. We challenged strains of Saccharomyces cerevisiae with soluble Cu and c-CuNPs to evaluate the potential of c-CuNPs as antifungal agents. Cells exposed to c-CuNPs demonstrated greater sensitivity to Cu than cells exposed to soluble Cu, although Cu-resistant strains were more tolerant than Cu-sensitive strains of c-CuNP exposure. At the same level of growth inhibition, 157 μM c-CuNPs led to the same internal Cu levels as did 400 μM CuSO4, offering evidence for alternative mechanisms of toxicity, perhaps through β-arrestin dependent endocytosis, which was supported by flow cytometry and fluorescence microscopy of c-CuNPs distributed both on the cell surface and within the cytoplasm. Genes responsible for genetic variation in response to copper were mapped to the ZRT2 and the CUP1 loci. Through proteomic analyses, we found that the expression of other zinc (Zn) transporters increased in Cu-tolerant yeast compared to Cu-sensitive strains. Further, the addition of Zn at low levels increased the potency of c-CuNPs to inhibit even the most Cu-tolerant yeast. Through unbiased systems biological approaches, we identified Zn as a critical component of the yeast response to Cu and the addition of Zn increased the potency of the c-CuNPs.
Significance to metallomicsSaccharomyces cerevisiae has developed resistance to copper independently multiple times and provides an ideal model to assess the response of fungi to Cu. Previously, we demonstrated that c-CuNPs are an effective antibacterial material. In an effort to understand the mode of action of c-CuNPs, improve potency, ensure broad spectrum activity, and limit the development of antimicrobial resistance, the response of S. cerevisiae to c-CuNPs was characterized using proteomic and genetic approaches. c-CuNPs expose yeast to higher Cu levels and, by comparing the response of copper resistant to copper sensitive yeast, the role of Zn was uncovered. Through these interactions, we suggest that the antimicrobial effectiveness of metallic hybrid nanomaterials will enhance microbial control, which may offer a new and wide range of antimicrobial materials for public health and food safety. |
A. Introduction
Copper (Cu) has been used since ancient times to prevent microbial growth because of its broad-spectrum activity coupled with multiple modes of toxicity.1 Excessive Cu induces damage by catalyzing the generation of reactive oxygen species (ROS).2 ROS generated by extracellular Cu oxidize lipids and proteins of the cell membrane3 and, if cells survive these lesions, then DNA damage can occur from the high levels of internal Cu.4 These mechanisms of toxicity and the response to Cu in microorganisms have been widely studied.5 Importantly, the expression of genes in response to Cu is highly regulated to maintain a fine balance of this essential, yet toxic, trace metal within the cytoplasm.6,7Nanotechnology is a transformative tool that can be used to develop and enhance high-value products from renewable and biocompatible raw materials such as wood.8–12 Cellulose, as the predominant component of woody biomass, can be utilized not only commercially where bio-compatibility and biodegradability are relevant, but also as a support structure for nanoparticle formation in novel applications. The use of the cellulose nanostructure as a template and stabilizer for metallic nanoparticles expands the potential applications of nanomaterials.13 Cellulosic cuprous nanoparticles (c-CuNPs) display effective antimicrobial activity against a non-pathogenic Escherichia coli DH5α.14,15 c-CuNPs contain reduced copper (Cu(I)), which is thought to be slowly released as Cu(I) and then oxidized making Cu(II) available to interact with biological molecules leading to toxicity.
Improving c-CuNPs depends on understanding their range of functions in order to manufacture products with increased antimicrobial effectiveness. We have developed a cost-effective hybrid nanomaterial built on a cellulosic scaffold with Cu(I) chemically linked to cellulose.16 To understand how yeast respond to Cu delivered in c-CuNPs, we carried out toxicological and quantitative proteomic analyses on genetically diverse yeast strains. c-CuNPs delivered more Cu to yeast cells compared to soluble CuSO4, and this may result from the endocytic uptake of c-CuNPs. The levels of hundreds of proteins changed in response to Cu exposure, and yeast strains that do not tolerate Cu fail to increase expression of Zrt1, a high affinity zinc (Zn) transporter. Furthermore, a genome-wide quantitative trait loci (QTL) analysis uncovered a role for Zn transporters in the Cu response. The genetic variation in the low-affinity membrane transporter, Zrt2, directly contributed to variation of Cu tolerance, while yeast lacking Zrt1 or Zrt3 were sensitive to high levels of Cu. Supplementing Zn decreased the growth of all yeast strains tested when also challenged with c-CuNPs. Understanding the interdependence of metal tolerances provides insights into improving the antimicrobial properties of metal-based materials.
B. Materials and methods
Yeast strains and plasmids
The S288c strains used included strains S96 (MATa lys517) and GSY147 (MATa prototroph18). YJM789 (MATalpha lys217,19) was the parent of YJM789K5a (MATa prototroph). YJM789K5a was generated by transforming a PCR product containing LYS2 into YJM789 and then backcrossed to HS959 (MATa lys2) that is otherwise isogenic to YJM789. All S288c strains displayed a similar copper resistance. Both YJM789 strains were copper sensitive. Other strains used were AWRI1631, RM11, and YJM339.20–22 The haploid recombinant segregant collection between YJM789 and S96 was previously generated.17 Knockout strains were previously described.23 Cloning of ZRT2 alleles was carried out by PCR amplified from genomic DNA using primers as follows: 5′ZRT2 GTACCTGCAGGCGTCCTAATGAAAATGGAGG and 3′ZRT2 TTCCCGGGATTACTCTGCCCTTTTC. Endogenous promoter and terminator sequences within 580 nucleotides upstream of the start and 277 nucleotides downstream of the stop codons were included. Genes were cloned into the SbfI and XmaI sites of the pGS36 plasmid with the hygromycin resistance marker.18 Plasmids were kept under selection with hygromycin after LiAc chemical transformations.Media and chemicals
All haploid yeast strains were grown in yeast minimal media (YM), which includes 2% dextrose, 6.7 g L−1 yeast nitrogen base, and 20 g L−1 agar in solid media. In liquid media, the agar was omitted. YM was, when needed, supplemented with lysine, methionine, uracil, leucine, and histidine. Plasmids were maintained by addition of 250 μg mL−1 hygromycin. All antioxidants were added to solid media when it had cooled to 55 °C and used within 24 hours. Chronic exposure was defined as two days of exposure, while acute exposure was defined as two hours of exposure. Viability assays were carried out in liquid culture in biological triplicates. Yeast were grown to log-phase and split into subcultures. Subcultures served as controls, receiving no added Cu; were soluble Cu-treated, receiving various concentrations of CuSO4·5H2O (Ricco Chemicals, CAS 7732-18-5); or received c-CuNPs. Carboxymethyl cellulose (CMC) copper nanoparticles (c-CuNPs) were created and physically characterized as previously reported,16 where Cu is reduced and bound onto free hydroxyls of the cellulose scaffold. 10 μg mL−1 of c-CuNP corresponded to 157 μM of soluble Cu determined by atomic absorption spectrometry (AAS). Yeast medium is acidic (pH 5), which would cause dissociation and oxidization of Cu from the c-CuNPs. Indeed, 27% more free copper was found in c-CuNPs suspended in YM than in water alone.Atomic absorption spectrometry
Strains YJM789K5a, GSY147, RM11, and YJM339 were grown in YM media in triplicate to the log phase. The control samples (YM) were collected. The remaining cultures were treated with 400 μM CuSO4 or 157 μM Cu as c-CuNPs (10 μg mL−1 of CMC) for 2 hours. For cell collection, an equal number of cells, as determined by optical density at 600 nm, was obtained (equivalent to 4 ODu). These samples were centrifuged and washed twice with distilled water. One ODu was separated for protein quantification. Samples were split, frozen in liquid nitrogen and stored at −80 °C. Soluble protein concentration was determined via Bradford assay. For AAS, the cell pellets were treated with 600 μL of concentrated HNO3 and 200 μL of 30% H2O2 for digestion. These solutions were transferred to glass tubes and placed in a boiling water bath for 2 hours until clear. A 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 10 dilution with distilled water was analyzed for Cu at 324.8 nm. Copper concentrations were given in mg L−1, adjusted for dilution, and normalized to soluble protein concentration for each sample. AAS was performed across four days, and a total average for the four repeated samples was determined for each day. From these values, the overall average was determined. To calculate the correction factor for each day's analysis, the overall average was divided by the daily average. This correction factor was then applied to all the values for that particular day, prior to dilution adjustments.
10 dilution with distilled water was analyzed for Cu at 324.8 nm. Copper concentrations were given in mg L−1, adjusted for dilution, and normalized to soluble protein concentration for each sample. AAS was performed across four days, and a total average for the four repeated samples was determined for each day. From these values, the overall average was determined. To calculate the correction factor for each day's analysis, the overall average was divided by the daily average. This correction factor was then applied to all the values for that particular day, prior to dilution adjustments.
Staining of yeast cells and cellulose copper nanoparticles for flow cytometry and microscopy
Cells of each strain were cultured from frozen stock in YM media to the log phase and harvested by centrifugation. The cells were then transferred to 10 mM 4-(2-hydroxyethyl)-1-piperazineethanesulfonic acid (HEPES) buffer (pH of 7.4) containing 5% Rhodamine B (RhodB) was added to final concentration of 100 nM, and the cells were incubated at room temperature for 20 minutes in the dark. c-CuNPs were added to phosphate buffered saline (PBS) to a concentration of 400 μM with Fluorescein isothiocyanate (FITC) in dimethyl sulfoxide (DMSO) to a final concentration of 2.568 μM. The FITC-dyed PBS nanoparticle solution was incubated in the dark for 8 hours at 4 °C. Because excess FITC could not be removed from the solution, 5.136 mM in DMSO was added to PBS to a final concentration of 2.568 μM and incubated in the dark for 8 hours at 4 °C to serve as a control.DMSO was purchased from Biotium (Hayward, CA). A four laser LSR/Fortessa (BD Biosciences, San Jose CA) with BD FACS Diva software (BD Biosciences) located in the WVU Flow Cytometry & Single Cell Core Facility was used to collect flow cytometric data. Cells of each strain were transferred from HEPES buffer, treated, and transferred to PBS. To determine the extent of association of c-CuNPs with the yeast cell wall, 1 mL of PBS containing 1 million dyed cells was added to 1 mL of 400 μm dyed c-CuNP PBS solution. To control for FITC labeling of cells, 1 mL of PBS containing 1 million dyed cells was added to 1 mL of 2.568 μM FITC in PBS. Solutions were allowed to incubate at 30 °C for 2 hours. Samples in PBS were analyzed on an LSR Fortessa (BD Biosciences San Jose CA) using BD FACS Diva version 8.0 software (Fig. S1, ESI†). A total of 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 events were collected/sampled and the median fluorescence intensity for each sample was calculated. Also, in order to ensure that the FITC signal only came from c-CuNPs that were inside the yeast, trypan blue was added to the samples prior to analysis on the flow cytometer.
000 events were collected/sampled and the median fluorescence intensity for each sample was calculated. Also, in order to ensure that the FITC signal only came from c-CuNPs that were inside the yeast, trypan blue was added to the samples prior to analysis on the flow cytometer.
After treatment, samples were harvested by centrifugation and re-suspended in 1 mL PBS containing 4% paraformaldehyde and incubated for 30 minutes at 30 °C. Cells were washed twice with PBS and re-suspended in 500 μL of 80% glycerol before sonicating at 10% amplitude for six seconds with one second pulses for 5 intervals of 1 second bursts on a Branson model SSE-1 Sonifier. Concanavalin A (Sigma – L7647) coated microscope slides were utilized to mount 10 μL of the samples and sealed with clear fingernail polish. Fluorescent yeasts were viewed using a laser scanning confocal microscope (Olympus FV1000) equipped with red/green HeNe and argon lasers. Images were acquired using a 60× oil immersion magnification optical objective. Distances between confocal planes were optimized for the objective using Fluoview software (FV10-ASW Version 04.00.02.09). Pixel resolution was adjusted for best resolution capable of performing a z-stack scan. Images were only modified for contrast enhancement.
Viability assay
After two hours with 400 μM of copper or 157 μM c-CuNPs, the cultures were washed twice with water, and the OD600 was measured. 0.1 OD unit (ODu) of each sample was diluted 100-fold and 100 μL was plated on YPD. The number of untreated colonies that grew was considered 100% viability, and the viability of treated cells was taken as a percentage of untreated cells.Proteomics
To measure the global proteomic changes of S288c and YJM789 in response to Cu, cells were treated for 90 minutes in YM with or without 800 μM CuSO4. Peptides were prepared using a modified FASP method.24 Treated cells were harvested by briefly spinning down 5 ODu of cells, washing once with water and then freezing cell pellets in liquid nitrogen in 2 mL screw cap tubes. Cells were lysed in 4% SDS, 100 mM DTT, and 100 mM Tris pH 8.5 in a ratio of 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 10 v/v cell pellet to buffer. 0.5 mm zirconium beads were added and the cells were lysed in a Fastprep for 3 × 20 seconds at speed 5 with a 5 minute rest at room temperature. Using a 24-gauge needle, the bottom of the tube was punctured and placed into a 2 mL low protein binding locking cap tube. All liquid and cellular debris were then spun at 1000 rpm into a 2 mL tube. The tubes were incubated at 95 °C for 4 minutes. Cellular debris was pelleted for 10 minutes at 14
10 v/v cell pellet to buffer. 0.5 mm zirconium beads were added and the cells were lysed in a Fastprep for 3 × 20 seconds at speed 5 with a 5 minute rest at room temperature. Using a 24-gauge needle, the bottom of the tube was punctured and placed into a 2 mL low protein binding locking cap tube. All liquid and cellular debris were then spun at 1000 rpm into a 2 mL tube. The tubes were incubated at 95 °C for 4 minutes. Cellular debris was pelleted for 10 minutes at 14![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 rpm at room temperature. The supernatant was transferred to new tubes. Protein concentration was calculated using Bradford reagent on a 1
000 rpm at room temperature. The supernatant was transferred to new tubes. Protein concentration was calculated using Bradford reagent on a 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 100 diluted sample. 200 μg of protein was diluted with 8 M urea/100 mM Tris pH 8.5 to a final SDS concentration of 1% and transferred to a Microcon YM-30 column. The Microcon columns were spun at 10
100 diluted sample. 200 μg of protein was diluted with 8 M urea/100 mM Tris pH 8.5 to a final SDS concentration of 1% and transferred to a Microcon YM-30 column. The Microcon columns were spun at 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 rpm. Proteins were washed five times with 8 M urea/100 mM Tris pH 8.5 and resuspended in 100 μL of 0.05 M idoacetamide in 8 M urea/0.1 M Tris HCl 8.5. The samples were alkylated for 45 minutes in the dark. Liquid was spun through the Microcon. The buffer was exchanged for 200 mM TEAB by washing the proteins five times. Protein concentrations were retested using a BCA kit (Thermo Fisher Scientific) on 1
000 rpm. Proteins were washed five times with 8 M urea/100 mM Tris pH 8.5 and resuspended in 100 μL of 0.05 M idoacetamide in 8 M urea/0.1 M Tris HCl 8.5. The samples were alkylated for 45 minutes in the dark. Liquid was spun through the Microcon. The buffer was exchanged for 200 mM TEAB by washing the proteins five times. Protein concentrations were retested using a BCA kit (Thermo Fisher Scientific) on 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 100 diluted samples. Trypsin was freshly dissolved in 200 mM TEAB and incubated at 30 °C for 20 minutes. Trypsin was incubated with proteins on the Microcon wrapped in parafilm overnight at 37 °C at 1
100 diluted samples. Trypsin was freshly dissolved in 200 mM TEAB and incubated at 30 °C for 20 minutes. Trypsin was incubated with proteins on the Microcon wrapped in parafilm overnight at 37 °C at 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 20 dilution. Digested peptides were spun through the Microcon and collected into fresh tubes. The concentration of peptides was approximated, and 5 μg of peptides were labeled with Thermo Fisher Tandem Mass Tags 6plex in a 10
20 dilution. Digested peptides were spun through the Microcon and collected into fresh tubes. The concentration of peptides was approximated, and 5 μg of peptides were labeled with Thermo Fisher Tandem Mass Tags 6plex in a 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 4.1 ratio according to the manufacturer's instructions for 1.5 hours at room temperature. The reaction was quenched with 5% hydroxylamine diluted in 200 mM TEAB. Equal amounts of labeled peptides were mixed and dried.
4.1 ratio according to the manufacturer's instructions for 1.5 hours at room temperature. The reaction was quenched with 5% hydroxylamine diluted in 200 mM TEAB. Equal amounts of labeled peptides were mixed and dried.
Peptides were resuspended in 100 mM ammonia formate at pH 10 and separated by reverse phase chromatography at high pH in the first dimension, followed by an orthogonal separation at low pH in the second dimension. In the first dimension the mobile phases were buffer A: 20 mM ammonium formate at pH 10 and buffer B: acetonitrile. Peptides were separated on a Xbridge 300 μm × 5 cm C18 5.0 μm column (Waters) using a 15 discontinuous step gradient at 2 μL min−1. Peptides eluted off from the first dimension were trapped by a Symmetry 180 μm × 2 cm C18 5.0 μm trap column (Waters). In the second dimension, peptides were loaded into an in-house packed 75 μm ID/15 μm tip ID × 20 cm C18-AQ 3.0 μm resin column with buffer A (0.1% formic acid in water). Peptides were separated with a linear gradient from 5% to 30% buffer B (0.1% formic acid in acetonitrile) at a flow rate of 300 nL min−1 over 180 minutes. Each sample separation was repeated three times.
Peptides were identified by using a linear trap quadrupole (LTQ)-Orbitrap Velos instrument (Thermo Fisher Scientific). The mass spectrometer was run in a data dependent mode. One survey scan acquired in the Orbitrap mass analyzer with resolution 60![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 was followed by MS/MS of the 10 most intense peaks with charge state ≥2 and above an intensity threshold of 5000. Peptides were fragmented in the high collisional cell (HCD) with normalized collision energy of 40% and activation time of 0.1 s. The MS/MS scan was acquired in the Orbitrap at resolution of 7500.
000 was followed by MS/MS of the 10 most intense peaks with charge state ≥2 and above an intensity threshold of 5000. Peptides were fragmented in the high collisional cell (HCD) with normalized collision energy of 40% and activation time of 0.1 s. The MS/MS scan was acquired in the Orbitrap at resolution of 7500.
Peptides were identified and quantitated using Proteome Discoverer 1.4.1.14 (Table S1, ESI†). Spectra were searched against a protein database consisting of all the S288c (http://downloads.yeastgenome.org/sequence/S288C_reference/genome_releases/S288C_reference_genome_R64-1-1_20110203.tgz) and YJM789 (http://downloads.yeastgenome.org/sequence/strains/YJM789/YJM789_Stanford_2007_AAFW02000000) proteins available in the Saccharomyces Genome Database (http://www.yeastgenome.org). Common contaminants such as human keratin and trypsin were also included in the database to prevent peptides of those contaminants from being falsely identified as yeast peptides (Table S2, ESI†). TMT6plex was used for quantification labeling according to the manufacturer's instructions and the six isobaric labels of 126–131 Da were assigned to samples as follows: 126 – S288c, YM, replicate 1; 127 – S288c, YM, replicate 2; 128 – S288c, Cu treated, replicate 1; 129 – S288c, Cu treated, replicate 1; 130 – YJM789, YM; 131 – YJM789, Cu treated.
The default method used by Proteome Discoverer to quantify protein fold change from peptide intensities is to calculate the median intensity ratio between two isobaric labels for all the peptide spectrum matches (PSMs) associated with the protein in question. This method is problematic when a peptide sequence is identified in multiple PSMs where ratios have large variation. For example, a peptide can have different phosphorylation states between two treatments, and consequently show very different ratios for the phosphorylated PSMs and the unphosphorylated PSMs. This type of variation can skew quantification results when there are only a few PSMs usable for quantification; therefore, we used different methods than the Proteome Discoverer default for protein quantification. For each peptide sequence identified by Proteome Discoverer, the reporter ion intensities of all PSMs associated with this peptide were summed into the total intensity for this peptide in each sample. For each protein, only its unique (not belonging to any other protein) peptides of high to middle confidence (false discovery rate 0.01–0.05) were considered usable for quantification. Four comparisons, each between two experimental scenarios, were performed by calculating intensity ratios of reporter ions: (1) S288c, Cu vs. YM – (128 + 129)/(126 + 127); (2) YJM789, Cu vs. YM – 131/130; (3) YJM789 vs. S288c, YM – 130 × 2/(126 + 127); (4) YJM789 vs. S288c, Cu – 131 × 2/(128 + 129). For the comparisons between strains, (3) and (4), only peptides showing no sequence polymorphisms between the two strains were considered usable for quantification. Four methods were used to calculate protein ratios for each comparison: (1) ‘means then ratio’ – the mean of all the usable peptides belonging to this protein was calculated for each experimental scenario, then the ratio between the two scenarios was calculated from the two means; (2) ‘medians then ratio’ – the median of usable peptides was calculated for each scenario, then the ratio was calculated from the two medians; (3) ‘ratios then mean’ – the ratio between the two scenarios was calculated for each usable peptide, then the mean of all the ratios was calculated; (4) ‘ratios then median’ – the ratio was calculated for each usable peptide, then the median of all the ratios was calculated. For each of the four methods and each comparison of two scenarios, the resulting ratios for all the proteins were divided by the median of the ratios for normalization. This normalization method assumes that the majority of reported proteins do not change much in each comparison, which was the case in our experiment (see Results). For each comparison, only the proteins showing ratios >2 or <0.5 by all the four methods were included in Table S1 (ESI†). The ratios calculated using method (1) ‘means then ratio’ were transformed into logarithms of 2 (two).
63![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 563 PSMs were reported with high to middle confidence (false discovery rate 0.01–0.05), representing 27
563 PSMs were reported with high to middle confidence (false discovery rate 0.01–0.05), representing 27![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 794 unique peptide sequences. Peptides were labeled with isobaric mass tags for quantification with Tandem Mass Tag (TMT). The TMT labeling rates of these PSMs were 92.7–97.3%. 4431 yeast proteins were reported containing at least one peptide with middle to high confidence (each case of two homologous proteins from S288c and YJM789 were counted as one protein), among which 2991 were quantified with at least two unique peptides and considered for differential abundance quantification. 112 proteins showed greater than two-fold change (absolute value of log
794 unique peptide sequences. Peptides were labeled with isobaric mass tags for quantification with Tandem Mass Tag (TMT). The TMT labeling rates of these PSMs were 92.7–97.3%. 4431 yeast proteins were reported containing at least one peptide with middle to high confidence (each case of two homologous proteins from S288c and YJM789 were counted as one protein), among which 2991 were quantified with at least two unique peptides and considered for differential abundance quantification. 112 proteins showed greater than two-fold change (absolute value of log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2 (fold change) or |log
2 (fold change) or |log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2fc| >1) between strains and/or copper treatments (Table S1, ESI†) by all the four calculation methods for fold change.
2fc| >1) between strains and/or copper treatments (Table S1, ESI†) by all the four calculation methods for fold change.
Quantitative trait loci studies on copper response
Strains from the recombinant haploid segregant collection17 from crossing S288c (S96) and YJM789 were grown to saturation in YM supplemented with lysine. The cells were then diluted to 0.2 OD600 in 200 μL in YM or YM with 50 μM CuSO4 in triplicate in 96 well plates. The plates were sealed and readings taken every hour in a TECAN M200, an automatic plate reader with no shaking to mimic growth on solid media. The culture density at 12 hours (late-log phase) and at 18 hours (endpoint or stationary phase) was used to map quantitative trait loci (QTL). All readings in Cu were subtracted from the same strain grown in YM for QTL values (Table S3, ESI†). For each locus, we calculated a growth score and a significance score. To calculate the growth score, all strains that inherited the segregant YJM789 (1) allele were assigned to group 1, and all strains that inherited the segregant S96 (2) allele were assigned to group 2. Average growth was calculated for strains in group 1 and was subtracted from the average growth for strains in group 2. The calculations resulted in the growth score. A positive growth score indicates that the segregant 1 allele leads to high growth and Cu resistance while a negative growth score indicates the segregant 2 allele confers Cu sensitivity. The significance score was also calculated, which is the probability of these growth scores occurring if the alleles were assigned randomly. To calculate the significance score, each yeast strain was randomly assigned to group 1 or group 2 regardless of which allele was inherited. The growth score was calculated given these 2 randomly assigned groups and repeated 100![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 times using a Monte Carlo simulation. The resulting distribution over growth scores was approximately normal. This normal distribution was used to calculate the significance score. We then studied the loci that have large magnitude growth scores and high significance scores for 12 hour growth (Table S4) and 18 hour growth (Table S5) (ESI†).
000 times using a Monte Carlo simulation. The resulting distribution over growth scores was approximately normal. This normal distribution was used to calculate the significance score. We then studied the loci that have large magnitude growth scores and high significance scores for 12 hour growth (Table S4) and 18 hour growth (Table S5) (ESI†).
C. Results
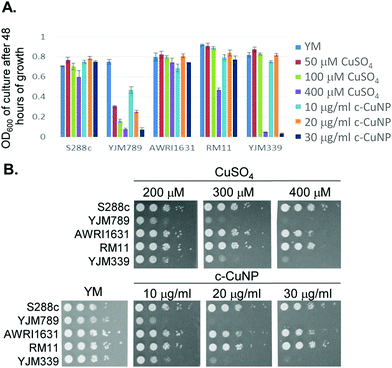
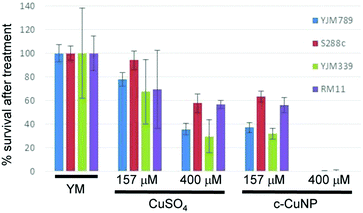
Clinical isolates are more sensitive to copper than agricultural isolates
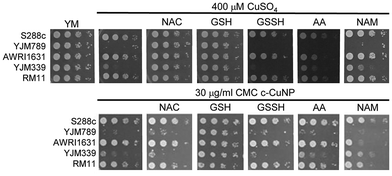
S. cerevisiae is among the oldest domesticated species and has evolved high tolerances to copper independently multiple times both in the wild and in lab-evolution experiments.5,25–29 To assess the antimicrobial potency of copper incorporated onto cellulosic nanoparticles, genetically diverse yeasts were incubated in liquid culture with different concentrations of c-CuNPs.16 The agricultural isolates AWRI1631,20 RM11,21,30,31 and S288c32,33 were resistant to Cu whereas YJM33919 and YJM789,19,34 clinical isolates of yeast, had lower Cu tolerances (Fig. 1A). The c-CuNPs were diluted to a level that would inhibit growth of YJM789 to a similar degree to 400 μM of soluble Cu when measured for growth in a plate reader without shaking, mimicking their growth on solid media. Both 30 μg mL−1 (by CMC concentration) of c-CuNPs and 400 μM of soluble copper decreased the growth of YJM789 by 90% compared to YM media. Similarly, both the soluble Cu and c-CuNPs embedded into solid agar media inhibited the growth of yeast, with the agricultural isolates being more resistant than the clinical isolates (Fig. 1B). However, growth patterns on agar suggested that c-CuNPs may be more toxic than soluble Cu, with growth at 10 μg mL−1 c-CuNPs inhibiting growth to the same extent as 200 μM Cu (Fig. 1B).Because copper can catalyze the generation of reactive oxygen species,2 we tested the ability of antioxidants to quench free radicals induced by soluble copper and c-CuNPs. Both glutathione (GSH) and N-acetylcysteine (NAC), a glutathione precursor, rescued growth inhibition by soluble copper in clinical strains YJM789 and YJM339 (Fig. 2). No growth rescue was detected with oxidized glutathione (GSSH) or ascorbic acid (another antioxidant) in the presence of Cu for YJM789, but there was a slight improvement for YJM339. When cells were treated with nicotinamide (NAM) and ascorbic acid (AA), the Cu tolerance of YJM339, but not YJM789, increased slightly, suggesting that, although YJM789 and YJM339 are Cu sensitive, there are other genetic differences that affect Cu response. However, NAC could not rescue growth inhibition caused by c-CuNPs. While NAC can function as an antioxidant when it is imported into cells, NAC is the precursor to glutathione, which is more potent at reducing ROS.35 Therefore, while the pattern of c-CuNP sensitivity approximately mirrored soluble Cu sensitivity, the difference in NAC rescue suggests differences in Cu delivery, and likely, cellular internalization, between cells exposed to soluble Cu and c-CuNPs.
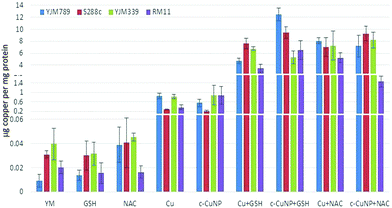
To determine how much Cu is imported into yeast, internal copper levels were measured by atomic absorption spectrometry (AAS). When grown in YM, YJM789 has three-fold less Cu than YJM339 or S288c. Furthermore, YJM789 accumulates as much Cu as RM11, a Cu-resistant strain (Fig. 3). Thus, basal untreated internal Cu levels do not correspond to Cu sensitivity. Treatment with GSH or NAC did not significantly change internal copper levels except for YJM789, in which cellular Cu increased with NAC to levels comparable to S288c and YJM339 (Fig. 3). Treatment with 400 μM Cu increased internal Cu concentrations in all strains, although to varying extents, by between 9- and 90-folds (Fig. 3). YJM789 and YJM339 had ∼0.8 μg of internal Cu per mg of protein, whereas Cu-resistant strains S288c and RM11 had levels 66% to 53% lower, respectively. To establish similar internal levels of copper from c-CuNPs to the levels from 400 μM soluble copper, the amount of c-CuNPs was decreased to 157 μM of Cu bound to CMC (= 10 μg mL−1 CMC). The acute exposure of yeast to 157 μM Cu as c-CuNPs led to higher Cu accumulation than levels measured when exposed to soluble Cu (Fig. 3): the internal Cu concentrations of yeast treated with 157 μM Cu as c-CuNPs were similar to those of yeast treated with 2.5 times higher levels of CuSO4. Co-treatment of yeast with glutathione and soluble Cu or c-CuNPs increased Cu uptake by 4- to 10-fold compared to copper treatments alone. The internal levels of Cu for GSH- and NAC-treated yeast, in general, were increased and similar irrespective of the copper source with two exceptions. YJM798 accumulated Cu from c-CuNPs when supplemented with GSH and RM11 exposed to c-CuNPs and NAC had lower internal Cu levels.
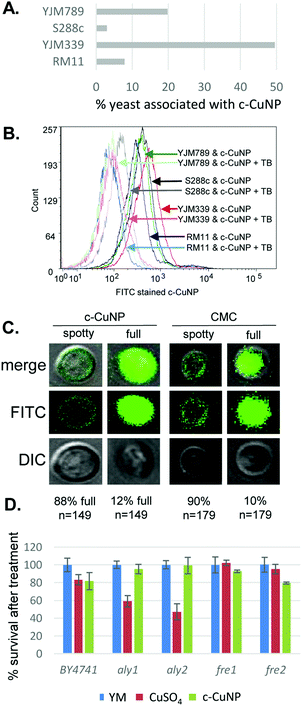
Because the c-CuNPs increased the efficiency of Cu delivery and lacked the NAC rescue to c-CuNP toxicity, we turned our focus to additional paths of Cu delivery by c-CuNPs. c-CuNPs are 10–20 nm particles with a unique three-dimensional spherical structure built on a cellulose scaffold.16 To assess the interaction of c-CuNPs with the outer surface of yeast cells, yeast cells were measured by flow cytometry in the presence of c-CuNPs. The yeasts were stained with Rhodamine B, which crosses cell membranes and is sequestered by mitochondria without inducing cell lysis, while c-CuNPs were stained with FITC. Because c-CuNPs labeled with FITC are too small to be directly detected by flow cytometry, FITC labeling can only be measured when c-CuNPs are associated with yeast cells (Fig. S1, ESI†). The fluorescence intensities of S288c, YJM789, YJM339, and RM11 cells incubated for two hours with 157 μM of FITC pre-stained c-CuNPs increased compared to yeast cells alone (Fig. 4A), suggesting the binding or uptake of c-CuNPs by yeast cells. In addition, a greater proportion of cells of YLM789 and YJM339 accumulated FITC-labeled c-CuNPs (Fig. 4A) and they accumulated these to a greater extent (Fig. 4B) than the S288c or RM11 lines.
To evaluate the physical location of c-CuNPs on yeast cells, cells exposed to FITC-labeled c-CuNPs were further treated with trypan blue, which quenches FITC fluorescence. Trypan blue does not enter living cells and can only quench FITC if the dye was physically located outside the yeast. This addition decreased the fluorescence intensity of FITC-stained c-CuNPs associated with yeast (Fig. 4B). To confirm that c-CuNPs were both outside and inside, the yeasts were again treated with c-CuNPs and CMC (cellulose backbone of the nanoparticles). Further examination revealed two populations of cells: most of the FITC stained c-CuNPs appeared as speckled spots on the outside of the yeast, while about 10% of the cells exhibited fluorescence inside the cells as seen through z-stack using confocal microscopy (Fig. 4C). The extracellular and intracellular fluorescence was not dependent on the presence of Cu on CMC (Fig. 4C).
To determine whether c-CuNPs were actively internalized by endocytosis, the viability of mutants in aly1 and aly2, arrestins that act as adapters for clatherin-mediated endocytosis,36 were evaluated. Also tested were mutants of fre1 and fre2,37,38 which reduce oxidized Cu for transport by Ctr1. These knockout strains were constructed in the BY4741 strain background that is related to S288c.23 Testing the viability of yeast under acute exposure can reveal smaller differences in sensitivity and determine if changes in viability to c-CuNP liquid culture is different from on plates. Cells were grown under similar conditions as when measuring internal copper levels (Fig. 3). When endocytosis is affected by deletion of Aly1 and Aly2, cellular viability was restored in the presence of c-CuNPs but not soluble copper (Fig. 4D). Loss of Fre1 and Fre2 increased cellular viability in the presence of soluble copper but not c-CuNPs (Fig. 4D).
Previous assessment of growth inhibition with c-CuNPs was carried out under chronic 2 day exposure measuring changes in growth in solid media (e.g., Fig. 1 and 2). Physical interactions between the yeast and c-CuNPs were measured under acute exposure in liquid media (e.g., Fig. 3 and 4). Viability assays were undertaken to evaluate the impact of soluble Cu and c-CuNPs on cell survival under acute exposure at comparable total Cu concentrations. Overall, the greater toxicity of c-CuNPs is evident across all isolates, with near complete loss of viability following exposure to 400 μM Cu as c-CuNPs (Fig. 5). Differential resistance of the isolates was also confirmed, with S288c and RM11 exhibiting greater viability than YJM789 or YJM339 regardless of the Cu source (Fig. 5).
Genetic and proteomic responses to copper in genetically diverse yeast
The four diverse strains of yeast used in this study have underlying genetic differences that contribute to variation in Cu response. S288c and YJM789 exhibited the most extreme differences in their abilities to tolerate high doses of Cu and moderate doses of c-CuNPs (Fig. 1 and 2). To evaluate potential molecular underpinnings of these responses to Cu, proteome- and genome-wide measurements were carried out. While many mRNAs are known to change in response to copper,7 we found only 22 proteins that showed at least 2-fold changes in S288c in response to 800 μM copper by quantitative mass spectrometry (Table 1 and Table S1, ESI†). Many other proteins showed more than 2-fold changes between strains but not between copper treatments (Fig. S2, ESI†). 19 out of the 22 copper-responsive proteins showed very little change (less than 1.2-fold change) in Cu-sensitive YJM789 cells when treated with 800 μM CuSO4. Among the 19 proteins, Cup1, Fit2, Fet3, and Zrt1 were upregulated, all of which are involved in metal transport (Table 1). Cup1 is a protein that binds Cu intracellularly and mediates tolerance to high Cu concentrations.39 The S288c genome has two CUP1 alleles annotated with two synonymous SNPs and identical protein sequences (Cup1-1 and Cup1-2), and they cannot be differentiated in proteomics.27 Cup1YJM789 exhibited a higher abundance in YM relative to Cup1S288c. Following Cu exposure, Cup1YJM789 demonstrated lower abundance relative to Cup1S288 due to the upregulation of Cup1S288c combined with little change in Cup1YJM789 abundance (Table 1). Fit2 and Fet340,41 are involved in iron (Fe) transport, and Zrt1 is involved in Zn transport.42 Fit2 and Zrt1 exhibited similar protein abundance between S288c and YJM789 in YM. After Cu exposure, Fit2 and Zrt1 showed higher abundance in S288c than in YJM789 because they were upregulated in S288c and changed little in YJM789 (Table 1). As for Fet3, S288c exhibited higher abundance than YJM789 in YM, and the upregulation by S288c in response to Cu exposure made strain-specific differences even greater (Table 1).![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2-fold ratio) between S288c and YJM789 with and without 800 μM CuSO4
2-fold ratio) between S288c and YJM789 with and without 800 μM CuSO4
| Order in dendro-gram | Systematic name | Protein name | S288cCu/S288cYM | YJM789Cu/YJM789YM | YJM789YM/S288cYM | YJM789Cu/S288cCu |
|---|---|---|---|---|---|---|
| 81 | YHR053C/YHR055C | Cup1 | 3.477 | 0.196 | 2.585 | −0.686 |
| 82 | YOR382W | Fit2 | 2.448 | −0.117 | 0.167 | −2.387 |
| 84 | YMR058W | Fet3 | 1.180 | −0.105 | −1.429 | −2.630 |
| 93 | YGL255W | Zrt1 | 1.141 | 0.123 | −0.268 | −0.848 |
| 63 | YDR424C | Dyn2 | −1.325 | −0.013 | 0.207 | 1.527 |
| 64 | YAL007C | Erp2 | −1.119 | 0.228 | 0.386 | 1.743 |
| 53 | YHR051W | Cox6 | −1.097 | −0.038 | 1.094 | 2.162 |
| 60 | YOL077W-A | Atp19 | −1.327 | −0.195 | 1.525 | 2.667 |
| 68 | YER057C | Hmf1 | −1.528 | 0.033 | 0.596 | 2.166 |
| 65 | YIL051C | Mmf1 | −1.328 | −0.043 | 0.445 | 1.739 |
| 58 | YBR230C | Om14 | −1.020 | −0.051 | 0.618 | 1.597 |
| 71 | YOR130C | Ort1 | −1.236 | −0.050 | 0.010 | 1.205 |
| 52 | YNL227C | Jjj1 | −1.050 | 0.015 | 0.796 | 1.869 |
| 72 | YDL081C | Rpp1a | −2.002 | −0.009 | −0.373 | 1.629 |
| 70 | YOR236W | Dfr1 | −1.134 | 0.574 | −0.346 | 1.429 |
| 80 | YNL190W | Ynl190w | 1.474 | −0.756 | 3.492 | 1.271 |
| 85 | YPR124W | Ctr1 | −1.966 | −0.494 | −1.380 | 0.101 |
| 35 | YAL046C | Aim1 | −0.168 | −1.334 | 1.136 | −0.021 |
| 96 | YMR173W | Ddr48 | 0.653 | 0.006 | −0.564 | −1.202 |
To further search for genetic loci contributing to Cu tolerance, we performed a genome-wide linkage study using a hybrid segregant collection17,43,44 to find association of genetic variation with differences in Cu tolerance between S288c and YJM789. 125 segregants from an S288c/YJM789 cross were grown at sub-lethal concentrations of copper (50 μM) in liquid culture (Fig. S3A–C, ESI†). In contrast to previous experiments that used an acute exposure and high Cu concentration, we used a long exposure and lower dose of Cu to model the adaptive response to low level copper exposure. S288c grew better in low doses of Cu than in media without additional Cu (Fig. S3C, ESI†) as opposed to YJM789, which was strongly inhibited by Cu (Fig. S3B, ESI†). In the presence of bathocuproinedisulfonic acid (BCS), a Cu chelator, S288c growth decreased (Fig. S3D, ESI†). The differences in growth of haploid segregants between YM and Cu treatment at 12 hours (mid-log phase) and 18 hours (endpoint) were used as quantitative trait loci (QTL). Using a threshold of LOD (logarithm of odds) >5, three regions of the genome were identified at the mid-log phase (Table S4 and Fig. S4A, ESI†) and one region was identified at the 18 hour endpoint linked to Cu resistance (Table S5 and Fig. S4B, ESI†). The genetic variations within these regions are most likely associated with differences in copper tolerance between S288c and YJM789.
One region of the genome that was identified in the 12 hour, but not 18 hour, QTL mapping was on chromosome VIII between coordinates 190![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 664 and 223
664 and 223![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 431, which contains twenty-two genes (Fig. S4C, ESI†). In the center of this region is the CUP1 locus, and its contribution to Cu tolerance has been well studied. The common laboratory strain S288c and many other Cu-tolerant strains contain independent amplifications of the CUP1 locus.27,39 Other studies have also found that laboratory-evolved strains amplify this region of the genome.45 The Saccharomyces Genome Database reference strain, S288c, has two alleles of CUP1 with YHR054c, a gene of unknown function, between them. This region is actually amplified seven times in S288c with fourteen copies of the CUP1 gene; YJM789 contains one copy of CUP1 in each repetitive unit that is amplified seven times.27
431, which contains twenty-two genes (Fig. S4C, ESI†). In the center of this region is the CUP1 locus, and its contribution to Cu tolerance has been well studied. The common laboratory strain S288c and many other Cu-tolerant strains contain independent amplifications of the CUP1 locus.27,39 Other studies have also found that laboratory-evolved strains amplify this region of the genome.45 The Saccharomyces Genome Database reference strain, S288c, has two alleles of CUP1 with YHR054c, a gene of unknown function, between them. This region is actually amplified seven times in S288c with fourteen copies of the CUP1 gene; YJM789 contains one copy of CUP1 in each repetitive unit that is amplified seven times.27
To pinpoint the specific genes that were contributing to Cu tolerance, we focused on the genes that had amino acid polymorphisms or copy number variation. The region on chromosome XI between 74![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 608 and 92
608 and 92![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 505 (Fig. S4D, ESI†) contained ten genes, among which MNN4, PEX1, MIA40, MST1, and DHP2 encoded amino acid polymorphisms between S288c and YJM789. Mia40 works with other proteins for the maturation of the mitochondrial fraction of Sod1.46 Sod1 is a major contributor to converting superoxides to hydrogen peroxide, which is further catalyzed into water and hydrogen47 and is important for tolerance to metals, including copper.48 In our proteomic analysis, Sod1 levels were not significantly changed between strains or in response to Cu.
505 (Fig. S4D, ESI†) contained ten genes, among which MNN4, PEX1, MIA40, MST1, and DHP2 encoded amino acid polymorphisms between S288c and YJM789. Mia40 works with other proteins for the maturation of the mitochondrial fraction of Sod1.46 Sod1 is a major contributor to converting superoxides to hydrogen peroxide, which is further catalyzed into water and hydrogen47 and is important for tolerance to metals, including copper.48 In our proteomic analysis, Sod1 levels were not significantly changed between strains or in response to Cu.
Another region that was identified in the endpoint linkage analysis was on chromosome II between coordinates 466![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 289 and 482
289 and 482![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 822 (Fig. S4B, ESI†). Within this interval is the LYS2 gene, which is mutated in YJM789 as an auxotrophic marker. As this peak was not seen at the 12 hour time point (Fig. S4A, ESI†), this may be interpreted as Lys2 becoming required as lysine present in the media was depleted as cells grew to saturation. Therefore, LYS2 may be selected by growth conditions instead of by Cu treatment.
822 (Fig. S4B, ESI†). Within this interval is the LYS2 gene, which is mutated in YJM789 as an auxotrophic marker. As this peak was not seen at the 12 hour time point (Fig. S4A, ESI†), this may be interpreted as Lys2 becoming required as lysine present in the media was depleted as cells grew to saturation. Therefore, LYS2 may be selected by growth conditions instead of by Cu treatment.
In both 12 and 18 hour association analyses, one strong peak was shared on chromosome XII between 393![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 415 and 414
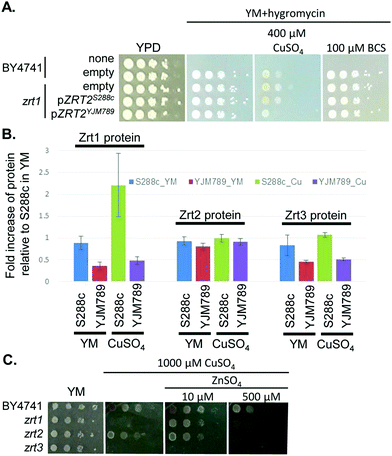
415 and 414![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 994 (S6 and Fig. S4A, B, ESI†), which contained eleven genes (Fig. S4E, ESI†). One of the genes known to be relevant to Cu tolerance was ACE2, which encodes the activating transcription factor of CUP1 expression.49 However, ACE2 showed no significant change in mRNA abundance in YM,50 no amino acid polymorphisms between S288c and YJM789, and minimal change in protein levels in response to Cu exposure. Near ACE2 was another gene of interest, ZRT2, which encodes a low affinity Zn transporter with five amino acid polymorphisms between S288c and YJM789.33 Like Ace2, Zrt2 exhibited less than 2-fold changes in protein abundance between strains or treatments. However, we cannot rule out changes in post-translational modifications of Zrt2 in response to Cu or the impact of the polymorphisms on transporter function. Therefore, we cloned ZRT2 from each strain, including the endogenous promoters and terminators, into plasmids. When the plasmids were expressed in BY4741 zrt2 deleted yeast, only yeast expressing Zrt2YJM789 grew more slowly on media containing 400 μM Cu and showed no change in growth on BCS (Fig. 6A). The polymorphisms within the Zrt2 protein apparently contribute to the Cu-sensitive phenotype of YJM789.
994 (S6 and Fig. S4A, B, ESI†), which contained eleven genes (Fig. S4E, ESI†). One of the genes known to be relevant to Cu tolerance was ACE2, which encodes the activating transcription factor of CUP1 expression.49 However, ACE2 showed no significant change in mRNA abundance in YM,50 no amino acid polymorphisms between S288c and YJM789, and minimal change in protein levels in response to Cu exposure. Near ACE2 was another gene of interest, ZRT2, which encodes a low affinity Zn transporter with five amino acid polymorphisms between S288c and YJM789.33 Like Ace2, Zrt2 exhibited less than 2-fold changes in protein abundance between strains or treatments. However, we cannot rule out changes in post-translational modifications of Zrt2 in response to Cu or the impact of the polymorphisms on transporter function. Therefore, we cloned ZRT2 from each strain, including the endogenous promoters and terminators, into plasmids. When the plasmids were expressed in BY4741 zrt2 deleted yeast, only yeast expressing Zrt2YJM789 grew more slowly on media containing 400 μM Cu and showed no change in growth on BCS (Fig. 6A). The polymorphisms within the Zrt2 protein apparently contribute to the Cu-sensitive phenotype of YJM789.
In the proteomic analysis, one other metal-responsive protein, Zrt1, was upregulated in S288c, but not in YJM789, when treated with Cu (Table 1). Zrt1 is a high affinity zinc transporter that is induced in Zn-depleted cells.42 While Zrt1S288c was upregulated approximately 2-fold by Cu treatment, other Zn transporters, including Zrt2 and Zrt3, an intracellular Zn transporter, did not exhibit significant changes in response to Cu in either strain (Fig. 6B). YJM789 displayed slightly lower Zrt3 expression compared to S288c regardless of Cu. Due to this association between Zn transporters and resistance to Cu, we tested whether increased levels of exogenous Zn could alter the Cu response of BY4741 and the Zn transporter mutant yeast lines. The zrt1 and zrt3 knockout lines were sensitive to 1000 μM Cu, whereas the zrt2 line was not (Fig. 6C). The addition of 10 μM Zn improved the growth of zrt1, but not zrt3, mutants, and the zrt2 line was still resistant to Cu and unaffected by supplemental Zn (Fig. 6C). These growth promotion effects were lost at 500 μM Zn in the presence of 1000 μM of Cu (Fig. 6C).
To determine if the relationship between Cu and Zn was a more general yeast response, additional yeast strains were tested. When S288c, YJM339, and RM11 were grown in 400 μM Cu in either the soluble form or from c-CuNPs, adding 10 μM Zn resulted in modest growth increases, which were lost or became negative at and above 25 μM Zn (Fig. 7). In contrast, YJM789 did not respond to supplemental Zn (Fig. 7). These results indicate that Zn, at low concentrations, plays a role in modulating Cu toxicity and this benefit may result from different Zn uptake capacities among yeast strains.
D. Discussion
We have characterized the interaction of the genetically diverse model eukaryotic organism, S. cerevisiae, with soluble Cu and c-CuNPs towards improving future applications of hybrid nanomaterials for microbial control. Copper has been used extensively throughout history as an antimicrobial agent and microorganisms exposed to Cu gain the opportunity to develop resistance mechanisms to this metal. Metabolic mechanisms that reduce internal metal accumulation or free metal speciation, which will reduce the production of toxic ROS in the cytoplasm, are important in conferring heavy metal resistance. Overcoming existing microbial resistance pathways and altering the balance of metals in cells may enhance the efficacy of existing metal-based antimicrobial materials and aid in the development of novel antimicrobial agents.Cellulosic copper nanoparticles exhibit enhanced toxicity
In the current study, the antimicrobial potency of Cu was increased by the delivery of Cu in the form of c-CuNPs. Exposure of 157 μM copper in NPs was as effective as 400 μM copper sulfate in yeast growth inhibition, and the increase in toxicity resulted from enhanced delivery of Cu to the cytoplasm (Fig. 3 and 5). Flow cytometry evidence and c-CuNP fluorescence suggested that CuNPs bound in large part to the cell walls of S. cerevisiae and such binding may presage the transport of Cu into the cell (Fig. 4B and C). Variation in binding of yeast cells and c-CuNPs mirrored resistance, with low binding strains exhibiting greater resistance to Cu (Fig. 4A). This differential binding may enhance exposure in Cu sensitive strains. Based on the size of c-CuNPs (10–20 nm),16 it is also possible that c-CuNPs were absorbed into S. cerevisiae by endocytosis and endocytotic activity varied between the strains. An endocytosis vesicle is 50–80 nm in S. cerevisiae51 and some cells in the current study exhibited internal fluorescence (Fig. 4C). Although most of the yeasts with incubated c-CuNPs appeared speckled with c-CuNP on the cell surface, approximately 10% of the yeast took up c-CuNPs as represented by intracellular FITC staining. We were unable to test if these two subpopulations had differences in survival, but 13% of the laboratory yeast had internal FITC stained c-CuNPs which was not statistically different from 18% of the yeast that fail to form colonies after c-CuNP treatment (Fig. 4C). Arrestin mutants in the laboratory strain (S288c background) displayed rescued cellular viability in the presence of c-CuNPs suggesting that endocytosis may be one mechanism of action of c-CuNPs, which would effectively deliver Cu to the cytoplasm.Differential expression of metal transporters is linked to toxicity
Copper uptake and subsequent toxicity, regardless of the mechanism of enhanced delivery of copper via CuNPs, will most likely be governed by the reduction and transport of metals across the plasma membrane and their cellular compartmentation in different strains. In S288c, exposure to Cu down-regulated the Cu transporter, Ctr1,52 while expression of the Cu and Fe oxidizer, Fet3,53,54 the plasma membrane bound facilitator of iron transport, Fit3, and the high affinity Zn transporter, Zrt1, increased. These patterns suggest that Cu exposure perturbs trace metal homeostasis and the more resistant S288c strain responds to these perturbations.40,55 YJM789 did not show significant changes in these proteins and we propose that the lack of regulation of Cu uptake underlies the sensitivity of YJM789 to Cu.The expression levels of Crt1 tracked cell Cu concentrations in YJM789 and S288c. Expression is lower in YJM789 than S288c under control conditions, yet reverse under exposure to Cu (Table 1), leading to YJM789 exhibiting low basal Cu levels in YM media, but increased Cu under exposure to Cu (Fig. 3). S288c regulation of expression more tightly regulates cell Cu concentrations under these conditions. Thus, S288c apparently has more responsive metabolic regulation of Cu uptake. Changes in other metal transport genes may reflect mass balance displacement of Fe and Zn by Cu from transporters or secondary stresses requiring additional Fe or Zn to overcome toxicity. Fet3 is a ferroxidase required for the reduction of Fe(II) and subsequent transport across the plasma membrane, and its upregulation may reflect increased demand for Fe due to inhibition of Fe(II) transport by Cu or increased demand for redox metabolism within the cell. Similarly, Fit2 is an iron and iron-siderophore-binding protein expressed on the cell surface40 that facilitates Fe uptake and may be directly regulating Cu response by adjusting internal iron levels for Sod2, a mitochondrial superoxide dismutase.56
The up-regulation of Zrt1, the high affinity Zn transporter, also points to Cu-induced Zn limitation in S. cerevisiae. High affinity transporters are typically activated by mineral deficiency, and the low km for Zrt1 in yeast (∼10 nM) suggests that it would be effective in overcoming deficiency.57 The fact that supplemental Zn rescued zrt1 knockouts (Fig. 6) and, to a lesser degree, native strains under high Cu exposure (Fig. 7) suggests that excess Cu ions at the plasma membrane surface may be impeding the binding of Zn to Zn transporters. It is interesting that the more Cu resistant strain S288c upregulates Zrt1 whereas the sensitive YJM789 does not, suggesting that metal homeostasis responses may be limited in cases where excessive Cu accumulation does not lead to loss of cellular signaling systems or excessive ROS damage as would occur in strains that cannot limit Cu accumulation (Fig. 3).
Intracellular copper binding reduces copper toxicity
Transporters, intracellular sequestration, and chaperonins/chelation maintain metal concentrations within strict bounds and tightly regulate the balance and potential toxicity of metals within the cytoplasm. In the current study, S288c up-regulated expression of Cup1, the main metallothionein in S. cerevisiae, in response to Cu. YJM789 did not show significant changes in levels of this metal-binding protein in the presence of Cu and we propose that the lack of response to increased cytoplasmic Cu increased YJM789 sensitivity to this metal. Unregulated free Cu concentrations in the cytoplasm will increase the production of ROS and lead to broad disruptions in cell homeostasis.In support of this, providing certain antioxidants increased the copper levels of all treated strains (Fig. 2). Glutathione can bind copper transported into cells by Ctr1 before delivery to copper chaperones.58 More importantly, glutathione and NAC may serve to prime phytochelatin and metallothionein production within S. cerevisiae.59,60 These peptides/proteins function to regulate internal free metal concentrations and maintain cytoplasmic metal homeostasis. The large increases in Cu accumulation from both soluble Cu and CuNPs may reflect enhanced production of these metal-binding systems, the chelation of Cu intercellularly, and internal storage of nontoxic metal complexes.
Addition of zinc increases copper toxicity
Although the toxicity of Cu was alleviated in some strains by the addition of low concentrations of Zn as discussed above, the antimicrobial activity of Cu was enhanced in the current study by the application of higher Zn concentrations (Fig. 7). As with Cu, Zn plays critical roles as a catalytic and structural cofactor in a wide array of proteins, and maintenance of free Zn2+ concentrations within the cytoplasm is important in maintaining metal homeostasis.61 Zinc is transported in the cell by the membrane-bound transporters Zrt1 and Zrt2 and is sequestered in the vacuole by Zrt3.62 Given that exposure of yeast to Cu induces the expression of Zrt1 in Cu-tolerant strains (Table 1), the addition of Zn into the environment may lead to the over-accumulation of Zn and subsequent toxicity to S. cerevisiae.When in excess, Zn will induce many of the same lesions in S. cerevisiae as does Cu, including oxidative damage and negative interactions with Fe transport. Thus, the enhanced microbial activity of Cu and Zn combined may be due to the additional sympatric lesions induced by Zn. However, there is evidence that the stresses induced by Cu and Zn are distinctly different. In a unique multi-metal study of S. cerevisiae response to stress,63,64 significant differences were found between the response of yeast to Cu and Zn. While there was some overlap in the common stress responses, such as metal transport and oxidative stress, significant groups of differentially expressed genes did not overlap and only two of the 10 most metal-responsive protein–protein and protein–DNA interacting networks were shared.64 Thus, there may be limited overlap in the cell's metabolic rescue capacity when, under Cu exposure, a second divergent metal is introduced. The addition of Zn to the production of Cu-based nanomaterials would improve the antimicrobial properties of these novel antimicrobial systems and may represent a new approach for public health and food safety.
Author contributions
XRM carried out the proteomics analysis and LJ developed the proteomics workflow. MJW carried out the microscopy, characterized Zrt2, and performed flow cytometry with JBL. JBL and VAP carried out viability assays. LMW and JRC measured internal copper concentrations. TWC analyzed QTL data. ZRL characterized Cup1 levels. ZRL, JRC, and JEGG wrote the manuscript. JEGG conceived and directed the experiments.Conflicts of interest
There are no conflicts to declare.Acknowledgements
We thank Mike Snyder for the use of the Orbitrap used in these studies. Randall Mann generously shared his common protein contaminant list. Angela Lee generously gave us the yeast knockout collection. Jenay Grant assisted in confirming the genomic amplification of the CUP1 locus. Flow cytometry was carried out on the WVU Flow Cytometry & Single Cell Core Fortessa. Microscopy was carried out with the aid of Phil Chapman. This research was funded by the National Science Foundation (grant #1144676), the West Virginia Clinical and Translational Sciences Institute (grant #GM104942), and the National Institutes of Health (grant #OD016165).Notes and references
- H. H. A. Dollwet and J. R. J. Sorenson, Trace elements in Medicine, 1985, vol. 2, pp. 80–87 Search PubMed.
- R. V Lloyd, P. M. Hanna and R. P. Mason, Free Radical Biol. Med., 1997, 22, 885–888 CrossRef.
- S. V Avery, N. G. Howlett and S. Radice, Appl. Environ. Microbiol., 1996, 62, 3960–3966 Search PubMed.
- D. Quaranta, T. Krans, C. E. Santo, C. G. Elowsky, D. W. Domaille, C. J. Chang and G. Grass, Appl. Environ. Microbiol., 2011, 77, 416–426 CrossRef CAS PubMed.
- M. I. Samanovic, C. Ding, D. J. Thiele and K. H. Darwin, Cell Host Microbe, 2012, 11, 106–115 CAS.
- T. J. Lyons, A. P. Gasch, L. A. Gaither, D. Botstein, P. O. Brown and D. J. Eide, Proc. Natl. Acad. Sci. U. S. A., 2000, 97, 7957–7962 CrossRef CAS.
- C. Gross, M. Kelleher, V. R. Iyer, P. O. Brown and D. R. Winge, J. Biol. Chem., 2000, 275, 32310–32316 CrossRef CAS PubMed.
- T. Wegner and P. Jones, Wood Fiber Sci., 2005, 37, 549–551 CAS.
- T. H. Wegner and P. H. Jones, Cellulose, 2006, 13, 115–118 CrossRef CAS.
- T. H. Wegner and E. P. Jones, Nanosci. Technol. Renewable Biomater., 2009, 1, 1–41 Search PubMed.
- J. McCrank, Nanotechnology applications in the forest sector, Natural Resources Canada, 2009 Search PubMed.
- R. J. Moon, A. Martini, J. Nairn, J. Simonsen and J. Youngblood, Chem. Soc. Rev., 2011, 40, 3941–3994 RSC.
- J. Cai, S. Kimura, M. Wada and S. Kuga, Biomacromolecules, 2008, 10, 87–94 CrossRef PubMed.
- T. Zhong, G. S. Oporto, J. Jaczynski and C. Jiang, BioMed Res. Int., 2015, 456834 Search PubMed.
- C. Jiang, G. S. Oporto, T. Zhong and J. Jaczynski, Cellulose, 2015, 1–10 Search PubMed.
- T. Zhong, G. S. Oporto, J. Jaczynski, A. T. Tesfai and J. Armstrong, Wood Fiber Sci., 2013, 45, 1–8 Search PubMed.
- L. M. Steinmetz, H. Sinha, D. R. Richards, J. I. Spiegelman, P. J. Oefner, J. H. McCusker and R. W. Davis, Nature, 2002, 416, 326–330 CrossRef CAS PubMed.
- J. W. Wenger, K. Schwartz and G. Sherlock, PLoS Genet., 2010, 6, e1000942 Search PubMed.
- J. H. McCusker, K. V Clemons, D. A. Stevens and R. W. Davis, Genetics, 1994, 136, 1261–1269 CAS.
- A. R. Borneman, A. H. Forgan, I. S. Pretorius and P. J. Chambers, FEMS Yeast Res., 2008, 8, 1185–1195 CrossRef CAS PubMed.
- R. K. Mortimer, P. Romano, G. Suzzi and M. Polsinelli, Yeast, 1994, 10, 1543–1552 CrossRef CAS PubMed.
- M. J. McCullough, K. V Clemons, C. Farina, J. H. McCusker and D. A. Stevens, J. Clin. Microbiol., 1998, 36, 557–562 CAS.
- A. H. Tong, M. Evangelista, A. B. Parsons, H. Xu, G. D. Bader, N. Page, M. Robinson, S. Raghibizadeh, C. W. Hogue, H. Bussey, B. Andrews, M. Tyers and C. Boone, Science, 2001, 294, 2364–2368 CrossRef CAS PubMed.
- J. R. Wisniewski, A. Zougman and M. Mann, J. Proteome Res., 2009, 8, 5674–5678 CrossRef CAS PubMed.
- A. C. Gerstein, J. Ono, D. S. Lo, M. L. Campbell, A. Kuzmin and S. P. Otto, Genetics, 2015, 199, 555–571 CrossRef PubMed.
- A. Hodgins-Davis, A. B. Adomas, J. Warringer and J. P. Townsend, Genome Biol. Evol., 2012, 4, 1061–1079 CrossRef PubMed.
- Y. Zhao, P. K. Strope, S. G. Kozmin, J. H. McCusker, F. S. Dietrich, R. J. Kokoska and T. D. Petes, G3, 2014, 2259–2269 CrossRef PubMed.
- J. C. Fay, H. L. McCullough, P. D. Sniegowski and M. B. Eisen, Genome Biol., 2004, 5, R26 CrossRef PubMed.
- D. J. Kvitek, J. L. Will and A. P. Gasch, PLoS Genet., 2008, 4, e1000223 Search PubMed.
- T. Torok, R. K. Mortimer, P. Romano, G. Suzzi and M. Polsinelli, J. Ind. Microbiol., 1996, 17, 303–313 CrossRef CAS.
- R. B. Brem, G. Yvert, R. Clinton and L. Kruglyak, Science, 2002, 296, 752–755 CrossRef CAS PubMed.
- R. K. Mortimer and J. R. Johnston, Genetics, 1986, 113, 35–43 CAS.
- J. M. Cherry, E. L. Hong, C. Amundsen, R. Balakrishnan, G. Binkley, E. T. Chan, K. R. Christie, M. C. Costanzo, S. S. Dwight, S. R. Engel, D. G. Fisk, J. E. Hirschman, B. C. Hitz, K. Karra, C. J. Krieger, S. R. Miyasato, R. S. Nash, J. Park, M. S. Skrzypek, M. Simison, S. Weng and E. D. Wong, Nucleic Acids Res., 2012, 40, D700–D705 CrossRef CAS PubMed.
- O. W. Tawfik, C. J. Papasian, A. Y. Dixon and L. M. Potter, J. Clin. Microbiol., 1989, 27, 1689–1691 CAS.
- A. M. Sadowska, Ther. Adv. Respir. Dis., 2012, 6, 127–135 CrossRef CAS PubMed.
- C. H. Lin, J. A. MacGurn, T. Chu, C. J. Stefan and S. D. Emr, Cell, 2008, 135, 714–725 CrossRef CAS PubMed.
- E. Georgatsou, L. A. Mavrogiannis, G. S. Fragiadakis and D. Alexandraki, J. Biol. Chem., 1997, 272, 13786–13792 CrossRef CAS PubMed.
- Y. Yamaguchi-Iwai, M. Serpe, D. Haile, W. Yang, D. J. Kosman, R. D. Klausner and A. Dancis, J. Biol. Chem., 1997, 272, 17711–17718 CrossRef CAS PubMed.
- M. Karin, R. Najarian, A. Haslinger, P. Valenzuela, J. Welch and S. Fogel, Proc. Natl. Acad. Sci. U. S. A., 1984, 81, 337–341 CrossRef CAS.
- O. Protchenko, T. Ferea, J. Rashford, J. Tiedeman, P. O. Brown, D. Botstein and C. C. Philpott, J. Biol. Chem., 2001, 276, 49244–49250 CrossRef CAS PubMed.
- D. M. De Silva, C. C. Askwith, D. Eide, J. Kaplan, D. M. De Silva, C. C. Askwith, D. Eide and J. Kaplan, J. Biol. Chem., 1995, 270, 1098–1101 CrossRef CAS PubMed.
- H. Zhao and D. Eide, Proc. Natl. Acad. Sci. U. S. A., 1996, 93, 2454–2458 CrossRef CAS.
- J. E. G. Gallagher, W. Zheng, X. Rong, N. Miranda, Z. Lin, B. Dunn, H. Zhao and M. P. Snyder, Genes Dev., 2014, 28, 409–421 CrossRef CAS PubMed.
- E. Mancera, R. Bourgon, A. Brozzi, W. Huber and L. M. Steinmetz, Nature, 2008, 454, 479–485 CrossRef CAS PubMed.
- G. M. Adamo, M. Lotti, M. J. Tamas and S. Brocca, Microbiology, 2012, 158, 2325–2335 CrossRef CAS PubMed.
- A. Varabyova, U. Topf, P. Kwiatkowska, L. Wrobel, M. Kaus-Drobek and A. Chacinska, FEBS J., 2013, 280, 4943–4959 CrossRef CAS PubMed.
- K. T. Tamai, E. B. Gralla, L. M. Ellerby, J. S. Valentine and D. J. Thiele, Proc. Natl. Acad. Sci. U. S. A., 1993, 90, 8013–8017 CrossRef CAS.
- M. M. Pena, K. A. Koch and D. J. Thiele, Mol. Cell. Biol., 1998, 18, 2514–2523 CrossRef CAS PubMed.
- G. Butler and D. J. Thiele, Mol. Cell. Biol., 1991, 11, 476–485 CrossRef CAS PubMed.
- W. Zheng, H. Zhao, E. Mancera, L. M. Steinmetz and M. Snyder, Nature, 2010, 464, 1187–1191 CrossRef CAS PubMed.
- C. Buser and D. G. Drubin, Microsc. Microanal., 2013, 19, 381–392 CrossRef CAS PubMed.
- S. A. Knight, S. Labbe, L. F. Kwon, D. J. Kosman and D. J. Thiele, Genes Dev., 1996, 10, 1917–1929 CrossRef CAS PubMed.
- D. de Silva, S. Davis-Kaplan, J. Fergestad and J. Kaplan, J. Biol. Chem., 1997, 272, 14208–14213 CrossRef CAS PubMed.
- C. Askwith, D. Eide, A. Van Ho, P. S. Bernard, L. Li, S. Davis-Kaplan, D. M. Sipe and J. Kaplan, Cell, 1994, 76, 403–410 CrossRef CAS PubMed.
- J. A. Stadler and R. J. Schweyen, J. Biol. Chem., 2002, 277, 39649–39654 CrossRef CAS PubMed.
- A. Naranuntarat, L. T. Jensen, S. Pazicni, J. E. Penner-Hahn and V. C. Culotta, J. Biol. Chem., 2009, 284, 22633–22640 CrossRef CAS PubMed.
- D. J. Eide, Biochim. Biophys. Acta, Mol. Cell Res., 2006, 1763, 711–722 CrossRef CAS PubMed.
- E. B. Maryon, S. A. Molloy and J. H. Kaplan, Am. J. Physiol., 2013, 304, C768–C779 CrossRef CAS PubMed.
- R. Kneer, T. M. Kutchan, A. Hochberger and M. H. Zenk, Arch. Microbiol., 1992, 157, 305–310 CrossRef CAS PubMed.
- J. Wünschmann, A. Beck, L. Meyer, T. Letzel, E. Grill and K. J. Lendzian, FEBS Lett., 2007, 581, 1681–1687 CrossRef PubMed.
- C. W. MacDiarmid, L. A. Gaither and D. Eide, EMBO J., 2000, 19, 2845–2855 CrossRef CAS PubMed.
- F. Winston, C. Dollard and S. L. Ricupero-Hovasse, Yeast, 1995, 11, 53–55 CrossRef CAS PubMed.
- X. Liu, J. Jia, R. Popat, C. A. Ortori, J. Li, S. P. Diggle, K. Gao and M. Cámara, BMC Microbiol., 2011, 11, 26 CrossRef CAS PubMed.
- Y. H. Jin, P. E. Dunlap, S. J. McBride, H. Al-Refai, P. R. Bushel and J. H. Freedman, PLoS Genet., 2008, 4, e1000053 Search PubMed.
Footnotes |
| † Electronic supplementary information (ESI) available. See DOI: 10.1039/c7mt00147a |
| ‡ These authors contributed equally. |
| § Current address: Department of Pathology, Microbiology, and Immunology, Vanderbilt University, Nashville, TN, USA. |
| ¶ Current address: Department of Biological Sciences and Chemical Engineering, Youngstown State University, OH, USA. |
| This journal is © The Royal Society of Chemistry 2017 |