Open Access Article
Open Access ArticleAn advanced conjugation strategy for the preparation of quantum dot-antibody immunoprobes
Veronika
Dvorakova
a,
Michaela
Cadkova
a,
Vladimira
Datinska
b,
Karel
Kleparnik
b,
Frantisek
Foret
b,
Zuzana
Bilkova
a and
Lucie
Korecka
 *a
*a
aDepartment of Biological and Biochemical Sciences, Faculty of Chemical Technology, University of Pardubice, Studentska 573, 532 10, Pardubice, Czech Republic. E-mail: lucie.korecka@upce.cz; Tel: +420 466 037 711
bInstitute of Analytical Chemistry of the CAS, v.v.i., The Czech Academy of Sciences, Veveri 967/97, 602 00, Brno, Czech Republic
First published on 22nd March 2017
Abstract
An advanced site-specific conjugation strategy for the preparation of quantum dot-based antibody probes applicable in various immunoassays from fluorescence to electrochemical biosensors is described. The combination of antigen (protein ApoE) modified magnetic particles providing protection to antibody binding sites, simple carbodiimide chemistry and carboxylate quantum dots (QDs) made of CdSe/ZnS is used for efficient labelling of anti-ApoE antibodies representing a model system. Polyacrylamide gel electrophoresis, fluorescence spectra measurements, capillary electrophoresis-laser induced fluorescence (CE-LIF) and square wave anodic stripping voltammetry (SWASV) were used for experimental verification of labelling efficiency. A simple change in antibody type makes this approach versatile and exploitable in a wide range of applications.
1. Introduction
Immunoglobulins and alternative antigen binding formats have the key position in a wide range of conventionally designed immunoassays fully utilizing the natural characteristics of antibodies to specifically target molecules or antigens.1–4 The conjugation of specific antibodies with signal generating molecules including conventional radioactive isotopes, organic dyes, reporter enzymes, biotin or new generation probes such as quantum dots, carbon- and graphene-based nanomaterials, dendrimers or fullerenes enables the detection of bioactive compounds of various physico-chemical nature occurring over a wide concentration range.5,6Recently semiconductor QDs have been considered as a new class of luminescent probes well suited for biological research and clinical medicine.7,8 Luminescent nanocrystal QDs represent spherical particles with diameters in the range of 1–15 nm.9 Their typical core–shell structure and material composition reflect their impressive opto-chemical properties for instance size-tunable emission, excellent signal brightness, and nearly no-photobleaching8 whereas charged ligands on the surface provide water solubility and biocompatibility.10 The ability to link QDs with various bioactive molecules without losing the aforementioned characteristics provides wide-field applications. Quantum dot-based immunochromatographic assays11,12 or QD-FRET biosensing devices13 are only two examples of numerous methods using QDs as a signal generating probe. The improved sensitivity, selectivity and multiplexity of QD-based methods enabled us to meet the high demands of the requirements for diagnostic methods for detecting tumours and other clinically important biomolecules.8,14
The unique properties of QDs such as their ability to generate fluorescence, chemiluminescence, conductivity or electrochemical signals enable us to select an appropriate way of detection based on the characteristics of the biomarker to be detected, its expected concentration, and taking into account also the complexity of the biological material and laboratory instrumentation.15,16
The surface properties of QDs play an important role in the conjugation process through which QDs can be attached to biomolecules.17,18 The structure of core–shell QDs with a “coating” made from polymer molecules provides enough active functional groups for covalent binding of various bioactive molecules. Generally, there is an effort to apply a user-friendly and low-cost conjugation strategy suited for routine production of quantum dot-based probes. In particular, traditional bioconjugation techniques including random, site-selective, covalent or non-covalent strategies are widely used in the case of core–shell QDs.8,19
Carbodiimide mediated coupling represents one of the top covalent strategies used for crosslinking between amine and carboxylic functional groups. Although this approach is quick and very effective due to the high reactivity of crosslinking reagents, the most common strategy is the combination of EDC with sulfo-NHS. The limited possibility to control the orientation of biomolecules to be conjugated, influencing the total activity of the modified carrier, is a drawback needed to be taken into account in the case of use. Conjugates prepared via carbodiimide chemistry are suitable for e.g. cellular imaging and tracking, for cancer cell targeting.20–22
Thiol chemistry is another frequently used conjugation strategy for the preparation of QD-nanoprobes for example for immunolabeling of cells or tissue specimens. Maleimide coupling is an example of the site-specific method strictly dependent on pH aimed at the conjugation of primary amines with thiols. However, the modification of proteins before conjugation is required in many cases.23,24
The carboxyl conjugation technique fully exploits the presence of carbohydrate groups located on the Fc fragment of immunoglobulins, and the reactive aldehyde groups react with the hydrazide modified counterpart. Conjugates prepared via the carboxyl crosslinking procedure are biocompatible and fit very well for sensing and bioimaging.25
His-tag site-specific coupling via polyhistidine residues is a great option for recombinant proteins to be conjugated with QDs. In this case Ni-NTA represents a biofunctional adaptor covalently linked to the QD whereas histidine-tagged antibodies bind the nickel ion by the chelation process.8 Low production cost is the main advantage. These conjugates have been already used for in vivo imaging applications.8,25,26
The coupling strategy based on the biospecific pair streptavidin-biotin belongs to the simplest and widely used non-covalent techniques. This technique provides quick and relatively stable binding among all cooperative partners. Despite the size of this conjugate, which could be limiting in some applications, usability in cell biology has already been mentioned.27,28
Besides the possibility to prepare a “lab-made” quantum dot-based conjugate by one of the methods mentioned above, there is also a commercially available labeling kit (SiteClick™ Antibody Labeling Kit) enabling specific conjugation of QDs with IgG molecules via modified carbohydrate domains. However, this antibody labeling system brings a few disadvantages such as the limited amount of QDs per 1 mol of IgG molecules and contamination of the final conjugation product by free non-conjugated QDs. Final purification steps are desired however with significant losses.
Our work is focused on the development of an advanced conjugation technique suitable for routine preparation of QD conjugates of various specificities. To prepare quantum dot-based probes of high quality is a necessary prerequisite for their integration into sensitive and robust bioassays.
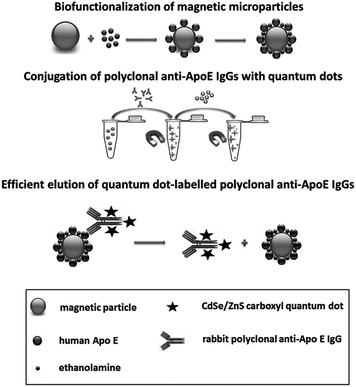
As many frequently used conjugation techniques, our protocol benefits from a well-known carbodiimide-based chemistry. Our original procedure includes (i) the biofunctionalization of the solid phase based on magnetic microparticles, (ii) capturing of the molecules to be conjugated with QDs by the molecules of the biospecific partner located on the magnetic carrier and covalent binding of QDs with biomolecules to be labeled, and (iii) the elution step providing a ready-to-use fraction of biomolecules labeled with QDs, i.e., probe molecules.
The main goal was to prepare specific probes for bioassay combined with highly sensitive electrochemical detection. Apolipoprotein E (ApoE) and rabbit polyclonal anti-ApoE IgGs were used as a model of the biospecific pair. Our intention was to develop such a labeling strategy to obtain a fraction of QD probes without the need to purify the product before use. Also the specific IgG molecules conjugated with QDs have to keep the characteristics such as high specificity and affinity for biomarkers to be detected. During the whole labeling procedure their binding activity has to remain unaffected. Their functionality was proven by a set of control methods based on luminescence spectra measurements, and electrophoretic and electrochemical analyses.
2. Materials and methods
2.1 Reagents and chemicals
Qdot® 565 ITK™ carboxyl quantum dots (CdSe/ZnS carboxyl QDs) were purchased from Thermo Fisher Scientific (Waltham, MA, USA). Magnetic particles SiMAG-carboxyl were from Chemicell GmbH (1 μm in size; Berlin, Germany). Recombinant human apolipoprotein E (ApoE) was obtained from BioVision (CA, USA) and rabbit polyclonal anti-apolipoprotein E IgG (polyclonal anti-ApoE IgGs) antibodies were the product of Moravian Biotechnology (Brno, CZ). Precision Plus Protein™ Standards Unstained were supplied by Bio-Rad (Bio-Rad, Hercules, CA, USA) and acrylamide, N,N,N′,N′-tetramethylethylenediamine (TEMED), ethylenediaminetetraacetic acid (EDTA), Tris(hydroxymethyl)aminomethane and N,N′-(1,2-dihydroxyethylene)bisacrylamide were supplied by Sigma-Aldrich (St. Louis, MO, USA). All other chemicals were of reagent grade and supplied by Lach-Ner (Neratovice, Czech Republic) or Penta (Chrudim, Czech Republic).2.2 Instrumentation
A spectrofluorimeter Jasco FP-8500, SAF-851 was purchased from JASCO Inc. (Easton, MD, USA), electrophoresis equipment and a ChemiDoc™ XRS+ Imaging System with Image Lab™ Software were from Bio-Rad (Hercules, California, USA).A diode laser, RLTMLL-405 Series was supplied by Roithner Lasertechnik GmbH (Vienna, Austria) and it was connected with a 405/10 nm laser clean-up filter MaxDiode™ provided by Semrock (Rochester, USA). A microscope (objective: 40 × 0.65 NA) was obtained from Oriel. The long pass edge filter EdgeBasic™ and a 607/70 nm band-pass filter BrightLine® were products of Semrock (Rochester, USA). A photomultiplier tube R647 was obtained from Hamamatsu (Iwata City, Japan). Silica capillaries (75 μm i.d., 375 μm o.d.) were purchased from Polymicro Technologies (Phoenix, AZ, USA) and CSW 1.6 software was guaranteed from Data Apex Dobrichovice (Dobrichovice, Czech Republic). A potentiostat/galvanostat PalmSens2 with PSTrace software was obtained from PalmSens, the Netherlands, and screen-printed carbon electrodes modified with mercury films (MeSPCE) were obtained from ItalSens, Italy.
2.3 Preparation of quantum dot-based anti-ApoE IgG conjugates
The particles with bound immunocomplexes ApoE–antiApoE IgG (0.5 mg) were mixed with 0.5 mg of EDC and 0.083 mg of sulfo-NHS (both in 500 μL of 0.1 M phosphate buffer). After 10 min incubation under gentle mixing at room temperature, 8 μL Qdot® 565 ITK™ carboxyl quantum dots (250 μM stock solution) mixed with 0.1 M phosphate buffer (500 μL) were added to the magnetic microparticles and incubated under gentle mixing overnight at 4 °C. After incubation, the particles were washed three times with 0.1 M phosphate and 0.1 M phosphate buffer with 0.2 M NaCl. The washing fractions were retained for subsequent analyses.
2.4 Luminescence spectra measurements
The luminescence signals of pure QDs, and fractions related to the labeling protocol such as pure QDs, unbound QDs, washing fractions and poly anti-ApoE IgGQDs conjugates were measured by using a spectrofluorimeter Jasco FP-8500 enabling one-drop measurement unit analysis at 25 °C. Luminescence emission spectra were recorded from 400 to 700 nm with excitation at 340 nm.2.5 SDS-PAGE and native TBE-PAGE
Denaturing SDS-PAGE in a standard arrangement (10% Tris–glycine separating gel, 5% Tris–glycine focusing gel)29 was performed mostly to control the purity and homogeneity of prepared conjugates. Whereas slightly modified native TBE-PAGE (15% Tris–borate separating gel, 5% Tris–borate focusing gel, both with addition of EDTA)30 was performed in order to monitor the quality and quantity of QDs (Fig. 1). Electrophoretic separation was carried out under these conditions: 0.1 M TBE buffer pH 8.3 running buffer, 180 V, 30 mA per one gel, approximately 50 minutes. TBE gels with samples of QDs were evaluated by UV illumination using Molecular Imager® ChemiDoc™ XRS+ with Image Lab™ software from Bio-Rad (Bio-Rad, Hercules, California, USA). Software Image J was used for QD quantification.2.6 CE-LIF analysis of poly anti-ApoE IgGQDs conjugates
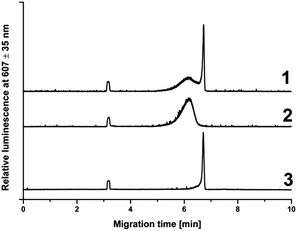
QDs and anti-ApoE IgGQDs conjugates were analyzed by separation in a laboratory-built CE-LIF system. Prior to each analysis, a 40 cm long (30 cm effective length) bare fused silica capillary (75 μm i.d., 375 μm o.d.) was rinsed successively with 0.1 M NaOH, water and finally with the running buffer (50 mM TRIS/TAPS, pH 8.6). The CE separation was driven by a Spellman CZE 1000R high power supply at a voltage of 10 kV. The samples were introduced electrokinetically at 10 kV for 5 s. At the positive voltage, the dominant mobility of electro-osmotic flow (EOF) carried all negatively charged analytes to the detector. A diode laser, RLTMLL-405 Series with a 405/10 nm laser clean-up filter was used as the excitation source at 405 nm. The fluorescence emission was collected by using a microscope objective (40 × 0.65 NA). A system of two filters including a 405 nm long pass edge filter and a 607/70 nm band-pass filter was used to block the 405 nm laser line and allow the maximum intensity emission wavelength of QDs to enter the photomultiplier tube type R647 window. The detector signal was recorded at a sampling rate of 25 Hz with a data acquisition and evaluation system CSW 1.6. The value of EOF mobility was determined using coumarin (1,2-benzopyrone) as the neutral marker in all pH of background electrolytes used.2.7 Electrochemical evaluation of anti-ApoE IgGQDs conjugates
The conjugation efficiency was confirmed by square wave anodic stripping voltammetry (SWASV). All tested samples were mixed with 0.1 M HCl in a ratio of 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
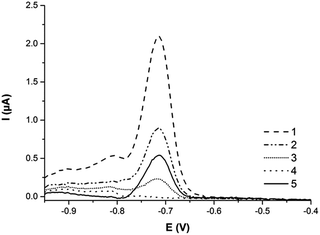
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 40 μL (v/v) in the final volume of 50 μL and incubated for 3 minutes. Before sample application, the surface of MeSPCE was electrochemically pretreated at a potential of −1 V for 300 s followed by subsequent measurement of the blank (three times). In all cases, 0.1 M HCl was used. In each measurement, the surface of the screen-printed electrode was washed three times with 0.1 M HCl. Detection conditions for sample measurement were as follows: condition potential 0 V, condition time 0 s, deposition potential −1 V, dep. time 120 s, potential range from −0.95 to −0.15 V, frequency 20 Hz and amplitude 0.0285 V. The current response at potential (−0.72 V) characteristic of Cd(II) ions released after acidic dissolution of CdSe/ZnS QDs was monitored.
40 μL (v/v) in the final volume of 50 μL and incubated for 3 minutes. Before sample application, the surface of MeSPCE was electrochemically pretreated at a potential of −1 V for 300 s followed by subsequent measurement of the blank (three times). In all cases, 0.1 M HCl was used. In each measurement, the surface of the screen-printed electrode was washed three times with 0.1 M HCl. Detection conditions for sample measurement were as follows: condition potential 0 V, condition time 0 s, deposition potential −1 V, dep. time 120 s, potential range from −0.95 to −0.15 V, frequency 20 Hz and amplitude 0.0285 V. The current response at potential (−0.72 V) characteristic of Cd(II) ions released after acidic dissolution of CdSe/ZnS QDs was monitored.
3. Results and discussion
Our work was aimed at the development of a versatile and highly efficient labeling protocol in order to prepare specific immunoprobes based on the conjugation of IgG molecules with Qdot® 565 ITK™ carboxyl quantum dots as an essential element of bioassays with electrochemical sensing.Our strategy for IgG molecule labelling is based on the assumption which has been many times experimentally confirmed. This method is called hydrogen/deuterium (H/D) exchange for monitoring of biospecific interactions by mass spectrometry.32,33 This method is used to localize the site of a molecule interacting with a ligand. To briefly summarize, the polypeptide chain forming the binding site of the IgG molecule is protected by ligand molecules (antigen) and H/D exchange does not occur. So our strategy was to protect the binding sites of specific antibodies by interactions with antigen molecules and perform the labeling with QDs only on sterically accessible parts of polypeptide chains.
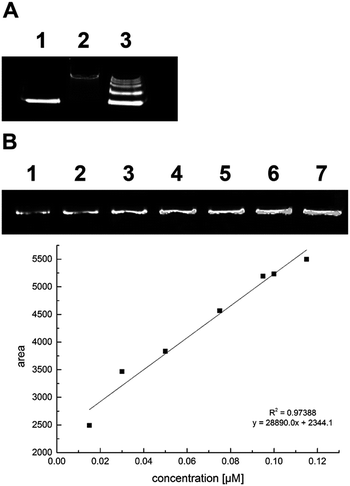
According to our experience, the quality and stability of QDs used for conjugation were decisive for the final outcome of the whole conjugation procedure. Native TBE-PAGE was modified for separation and quality characterization of commercial QDs (Fig. 2A). TBE-PAA gel electrophoresis was also suited for QD quantification (Fig. 2B).
With the use of Qdot® 565 ITK™ carboxyl QDs we have successfully designed and validated the three stages of the IgG labeling approach which is schematically presented in Fig. 1. The biofunctionalization of SiMAG-carboxyl magnetic microparticles with antigen ApoE molecules was the first step, where the particle surface was fully covered with antigens corresponding to the specificity of antibodies to be labeled. The suitable amount of antigen was calculated with respect to the antigen molecular weight and the size of magnetic particles utilized for the reaction. In that case, we applied the following equation: S = 6C/ρSΔd where S is the amount of representative protein (with respect to the molecular weight) required to achieve surface saturation (mg protein per g of microspheres), ρS is the density of the solid sphere (g cm−3), d represents the mean particle diameter (μm) and C is the capacity of the microsphere surface for a given protein (mg of protein per m2 of sphere surface).31 For antigen binding, we applied the carbodiimide-mediated crosslinking reaction where EDC was combined with sulfo-NHS. We tested one- and two-step immobilization protocols with overnight incubation in both cases, whereas the two-step protocol with 30 min-long pre-activation and application of 7.5 mg of EDC and 1.25 mg of sulfo-NHS has been considered as more beneficial, because of better immobilization efficiency. To avoid nonspecific binding and residual reactivity, an additional blocking step was added. Ethanolamine as an intact molecule of low molecular weight was used to block remaining activated carboxyl groups. Different time intervals (30–180 min, 24 hours) and concentrations (0.01–1 M) were tested. The 60 min blocking time with 0.1 M ethanolamine was assumed as optimal.
Thus the ApoE biofunctionalized magnetic carrier was incubated with specific IgG molecules to create stable immunocomplexes with site-specific oriented IgG molecules. In the case of antigens with molecular weights in the range of 25–40 kDa the double molar amount of antibodies added into the binding mixture is the most efficient (data not shown). Also different times (30, 60, 90, 120 minutes and overnight incubation) required for the formation of specific immunocomplexes have been tested. In the case of rabbit polyclonal anti-ApoE IgG, the time of immunocomplex formation 90 min was evaluated as the sufficient time for maximal coverage. To label the specific IgG molecules fixed in immunocomplexes on magnetic particles, EDC, sulfo-NHS and an appropriate amount of carboxyl QDs was added. Incubation was performed under gentle mixing (see 2.3 b). Finally, the magnetic beads were repeatedly washed to remove all reagents and free QDs.
The third phase of the protocol followed immediately. The molecules of poly anti-ApoE IgGQDs conjugates had to be released from the complexes with ApoE. Elution represents the disruption of many non-covalent bonds between the specific binding sites of IgG molecules and antigenic determinants of complex antigen molecules. We tested various elution reagents from the acidic and basic pH range (Table 1), and the time and number of elution steps were also optimized. Elution reagents were applied in three independent 15 minute-long intervals. The most efficient elution reagent has been determined based on spectrophotometric measurements of immunocomplexes before and after treatment with the elution reagent. The standard ELISA arrangement in a microtiter plate, with enzyme horseradish peroxidase as the antibody label, was used. To prove that the binding activity of antibodies is not affected by the used elution reagent, eluted antibodies were repeatedly used for immunocomplex formation (data not shown). Based on our results, the acidic medium namely 0.05% TFA with 0.5% SDS (pH 2.5) provided the strongest elution efficiency. Considering the very low pH the released conjugate was directly transferred to 0.1 M phosphate buffer (pH 7.3).
| Elution reagents | Efficiency [%] |
|---|---|
| Acidic media | |
| 0.1 M acetic acid with 20 mM CaCl2 (pH 4) | 41 |
| 0.1 M acetic acid with 0.2 M MgCl2 (pH 4) | 36 |
| 1 M arginine–hydrochloride (pH 4) | 37 |
| 0.5 M citrate (pH 4.3) | 9 |
| 0.1 M glycine with 0.2 M NaCl (pH 2.6) | 65 |
| 0.05% TFA with 0.5% SDS (pH 2.5) | 76 |
| 0.05% TFA with 1% SDS (pH 2.5) | 73 |
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) |
|
| Alkaline media | |
| 0.1 M citrate with 0.1 M NaOH (pH 8) | 73 |
| 0.1 M citrate with 0.1 M NaOH and 0.2 M MgCl2 (pH 7) | 16 |
| 0.1 M glycine with 0.1 M NaOH and 0.2 M MgCl2 (pH 8) | 12 |
| 1 M MgCl2 (pH 7.5) | 14 |
| 0.1 M NH4OH (pH 11) | 25 |
| 0.1 M NH4OH with 0.2 M MgCl2 (pH 11) | 12 |
| 0.1 M triethanolamine (pH 9) | 7 |
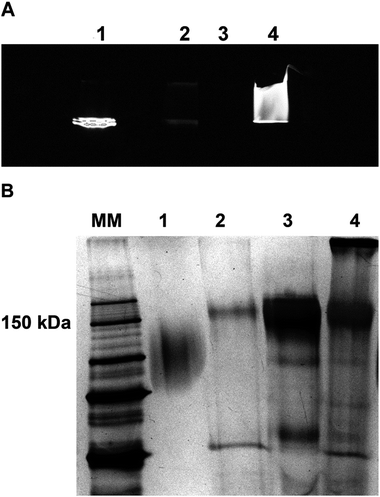
Fluorescence spectra measurements (Fig. 4A) and polyacrylamide gel analyses (Fig. 3 and 4B) supplemented by CE-LIF (Fig. 5) were chosen as fundamental methods for evaluating all the reactive compounds, bioactive molecules entering the reaction and also all the fractions obtained during the three steps of the labeling protocol. Square wave anodic stripping voltammetry (Fig. 6) was considered as the main confirmatory technique for the assessment of the functionality of the designed labeling protocol.
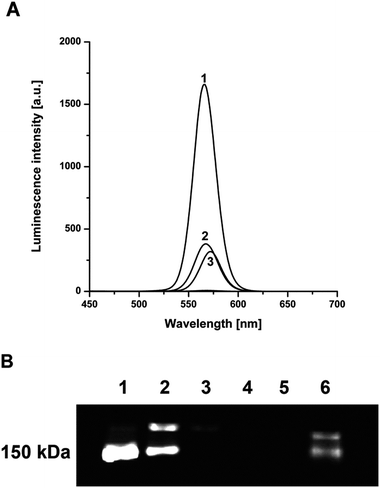
The luminescence emission spectra are shown in Fig. 4A. The pure QDs, unbound QDs and poly anti-ApoE IgGQDs conjugates provided detectable signals in contrast to washing fractions, which did not contain any traces of QDs. The samples were also loaded into the slab gel to perform TBE-PAGE accompanied by UV detection (Fig. 4B). The results of the luminescence spectra and TBE-PAGE demonstrated the functionality of the designed labeling procedure.
The quality and purity of poly anti-ApoE IgGQDs conjugates were confirmed by the CE-LIF assay. Due to the different electrophoretic mobilities of pure QDs and conjugates this technique allowed us to confirm the high efficiency of the labeling procedure and prove the absence of free QDs in the conjugate fraction (Fig. 5).
Square wave anodic stripping voltammetry was another confirmation technique for labeling efficiency assessment. The current response at a potential of −0.72 V, characteristic of CdSe/ZnS quantum dots, was obtained by the use of a disposable miniaturized screen-printed three-electrode sensor with a mercury film modified carbon working electrode (MeSPCE) (Fig. 6). The efficiency of the conjugation of quantum dots with antibodies was calculated as the ratio of electrochemical current responses of quantum dots of all fractions connected with the performed labeling protocol. The difference of the electrochemical current responses of the original sample (the original fraction of quantum dots before conjugation), the fraction of conjugates (antibodies conjugated with QDs) and the sum of current responses of binding (fraction after conjugation – the rest of the unbound QDs) and washing fractions was calculated with a resulting conjugation efficiency of 30%.
The quality of the poly anti-ApoE IgGQDs conjugate in terms of its ability to target the antigen specifically as well as further optimization of reaction conditions was verified.34
4. Conclusions
Here we have designed and validated a three-stage conjugation procedure to prepare QD-based immunoprobes for highly sensitive biosensors combined with electrochemical detection. This protocol fully exploited the benefits provided by superparamagnetic microparticles, i.e., easy, rapid and quantitative separation of the liquid phase with reagents from the solid phase carrying the molecules of antigens, immunocomplexes or bioconjugates. The conjugation strategy was based on traditional carbodiimide chemistry but combined with site-specific immobilization of IgG molecules on the surface of antigen biofunctionalized magnetic microparticles. The binding sites of all IgG molecules to be labeled are protected by antigen molecules just in immunocomplexes, and IgG affinity and specificity remained unaffected during all steps of the covalent coupling procedure. Another advantage of this labeling approach is the purity of the final product – the poly anti-ApoE IgGQDs conjugate. Since no contamination of the conjugate by free QDs was observed, no additional separation step was needed. This protocol for the efficient conjugation of IgG molecules with QDs is suitable wherever there are high demands on quality and purity of signal generating probes.Acknowledgements
At the Department of Biological and Biochemical Sciences, Faculty of Chemical Technology, University of Pardubice the work was supported by the Czech Science Foundation, project no. 15-16549S and project SGS-2016-004 of the Faculty of Chemical Technology (University of Pardubice, Czech Republic). At the Institute of Analytical Chemistry ASCR, v.v.i., the research was supported by the Czech Science Foundation, project no. 17-01995s.References
- I. Vainshtein and R. Lee, Bioanalysis, 2014, 6, 1939–1951 CrossRef PubMed.
- T. Fodey, P. Leonard, J. O'Mahony, R. O'Kennedy and M. Danaher, TrAC, Trends Anal. Chem., 2011, 30, 254–269 CrossRef CAS.
- F. Okda, S. Lawson, X. Liu, A. Singrey, T. Clement, K. Hain, J. Nelson, J. Christopher-Jennings and E. A. Nelson, BMC Vet. Res., 2016, 12, 95 CrossRef PubMed.
- C. Spiess, Q. Zhai and P. J. Carter, Mol. Immunol., 2015, 67, 95–106 CrossRef CAS PubMed.
- T. Wróbel, G. Mazur, J. Diana, M. Biedro, R. Badowski and K. Kuliczkowski, J. Oncol., 2005, 55, 316–319 Search PubMed.
- M. Arruebo, M. Valladares and A. Gonzále-Fernandéz, J. Nanomater., 2009, 2009, 1–24 CrossRef.
- A. M. Smith, S. Dave, S. Nie, L. True and X. Gao, Expert Rev. Mol. Diagn., 2006, 6, 231–244 CrossRef CAS PubMed.
- Y. Xing, Q. Chaudry, C. Shen, K. Y. Kong, H. E. Zhau, L. W. Chung, J. A. Petros, R. M. O'Regan, M. V. Yezhelyev, J. W. Simons, M. D. Wang and S. Nie, Nat. Protoc., 2007, 2, 1152–1165 CrossRef CAS PubMed.
- O. I. Mićić, H. M. Cheong, H. Fu, A. Zunger, J. R. Sprague, A. Mascarenhas and A. J. Nozik, J. Phys. Chem., 1997, 101, 4904–4912 CrossRef.
- J. J. Beato-López, C. Fernández-Ponce, E. Blanco, C. Barrera-Solano, M. Ramírez-del-Solar, M. Domínguez, F. García-Cozar and R. Litrán, Nanomater. Nanotechnol., 2012, 2, 1–9 CrossRef.
- L. Trapiella-Alfonso, J. M. Costa-Ferna, R. Pereiro and A. Sanz-Medel, Talanta, 2013, 106, 243–428 CrossRef CAS PubMed.
- A. N. Berlina, N. A. Taranova, A. V. Zherdev, M. N. Sankov, I. V. Andreev, A. I. Martynov and B. B. Zantiev, PLoS One, 2013, 8, 1–8 Search PubMed.
- D. Geißler and N. Hildebrandt, Anal. Bioanal. Chem., 2016, 408, 4475–4483 CrossRef PubMed.
- U. Resch-Geger, M. Grabolle, S. Cavaliere-Jaricot and R. Nitschke, Nat. Methods, 2008, 5, 763–775 CrossRef PubMed.
- C. Frigerio, D. S. Ribeiro, S. S. Rodrigues, V. L. Abreu, J. A. Barbosa, J. A. Prior, K. L. Marques and J. L. Santos, Anal. Chim. Acta, 2012, 735, 9–22 CrossRef CAS PubMed.
- D. Vasudevan, R. R. Gaddam, A. Trinchi and I. Cole, J. Alloys Compd., 2015, 636, 395–404 CrossRef CAS.
- H. Yukawa, R. Tsukamoto, A. Kano, Y. Okamoto, M. Tokeshi, T. Ishikawa, M. Mizuno and Y. Baba, J. Cell Sci. Ther., 2013, 4, 1–7 Search PubMed.
- K. Zhu, R. Dietrich, A. Didier, D. Doyscher and E. Märtlbauer, Toxins, 2014, 6, 1325–1348 CrossRef CAS PubMed.
- K. T. Yong, Y. Wang, I. Roy, H. Rui, M. T. Swihart, W. C. Law, S. K. Kwak, L. Ye, J. Liu, S. D. Mahajan and J. L. Reynolds, Theranostics, 2012, 2, 681–694 CrossRef CAS PubMed.
- T. A. Zdobnova, O. A. Stremovskiy, E. N. Lebedenko and S. M. Deyev, PLoS One, 2012, 7, 2–9 Search PubMed.
- P. Suriamoorthy, X. Zhang, G. Hao, A. G. Joly, S. Singh, M. Hossu, X. Sun and W. Chen, Cancer Nanotechnol., 2010, 1, 19–28 CrossRef CAS PubMed.
- D. A. East, D. P. Mulvihill, M. Todd and I. J. Bruce, Langmuir, 2011, 27, 13888–13896 CrossRef CAS PubMed.
- S. Avvakumova, M. Colombo, P. Tortora and D. Prosperi, Trends Biotechnol., 2014, 32, 11–20 CrossRef CAS PubMed.
- R. Bilan, K. braznik, P. Chames, D. Batyl, I. Nabiev and A. Sukhanova, Phys. Procedia, 2015, 73, 228–234 CrossRef CAS.
- Y. Zhang and T. H. Wang, Theranostics, 2012, 2, 631–654 CrossRef CAS PubMed.
- C. Schieber, A. Bestetti, J. P. Lim, A. D. Ryan, T. Nguyen, R. Eldridge, A. R. White, P. A. Gleeson, P. S. Donnelly, S. J. Williams and P. Mulvaney, Angew. Chem., Int. Ed., 2012, 51, 10523–10527 CrossRef CAS PubMed.
- A. Valizadeh, H. Mikaeili, M. Samiei, S. M. Farkhani, N. Zarghami, M. Kouhi, A. Akbarzadev and S. Davaran, Nanoscale Res. Lett., 2012, 7, 19–21 CrossRef PubMed.
- J. L. Liu, S. A. Walper, K. B. Turner, A. Brozozog, I. L. Medintz, K. Susumu, E. Oh, D. Zabetakis, E. R. Goldman and G. P. Anderson, Biotechnology Reports, 2016, 10, 56–65 CrossRef.
- U. K. Laemmli, Nature, 1970, 227, 680–685 CrossRef CAS PubMed.
- L. Holubova, L. Korecka, S. Podzimek, V. Moravcova, J. Rotkova, T. Ehlova, V. Velebny and Z. Bilkova, Carbohydr. Polym., 2014, 112, 271–276 CrossRef CAS PubMed.
- L. A. Cantarero, J. E. Butler and J. W. Osborne, Anal. Biochem., 1980, 105, 375–382 CrossRef CAS PubMed.
- A. J. Percy, M. Rey, K. M. Burns and D. C. Schriemer, Anal. Chim. Acta, 2012, 721, 7–21 CrossRef CAS PubMed.
- A. C. Hogenboom, A. R. de Boer, R. J. E. Derks and H. Irth, Anal. Chem., 2001, 73(16), 3816–3823 CrossRef CAS PubMed.
- M. Cadkova, V. Dvorakova, R. Metelka, Z. Bilkova and L. Korecka, Monatsh. Chem., 2016, 147, 69–73 CrossRef CAS.
| This journal is © The Royal Society of Chemistry 2017 |