Degradation of disperse blue 2BLN by oleaginous C. sorokiniana XJK
Li Xie,
Lin Zhou,
Tingting Liu and
Xiaolin Xu*
Key Laboratory for Green Process of Chemical Engineering of Xinjiang Bingtuan, School of Chemistry and Chemical Engineering, Shihezi University, Shihezi, 832003, People's Republic of China. E-mail: xuxl@shzu.edu.cn; Fax: +86-0993-2057270; Tel: +86-0993-2055015
First published on 25th October 2016
Abstract
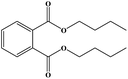
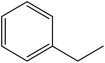
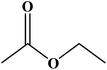
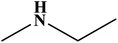
In this study, an oil-producing freshwater microalgae Chlorella sorokiniana XJK was identified and used for the degradation of disperse blue 2BLN. According to the decolorization rate, the optimal conditions were found to be as follows: initial dye conc. = 60 mg L−1, initial biomass density = 8 × 106 cell per mL, initial pH = 4, light intensity 470 μmol (m2 s)−1 and CO2/air (v/v) ratio = 1%. The decolorization rate reached up to 83% in 6 days cultivation with Chlorella sorokiniana XJK, which also obtained 570 mg L−1 of biomass and 43% of lipid content under the optimum conditions. Moreover, the enzyme activities (laccase and manganese peroxidase) in XJK were determined to be 3.89 U mL−1 and 4.86 U mL−1, respectively, which played a significant role in dye decolorization. The degradation products of the dye were analyzed by UV-vis, FTIR and GC-MS methods. This revealed that the major chromophore group was broken down and disperse blue 2BLN was converted into several low-molecular compounds including dibutyl phthalate, ethylbenzene and ethyl acetate during the microalgae-based treatment. This indicated that XJK shows a potential for dye degradation and providing materials for biofuel.
1. Introduction
Nowadays, with the rapid development of economy and industry, synthetic dyes are commonly and widely applied in energy sources, agriculture and all types of industries. According to their chemical structure,1 anthraquinone dyes are the critical groups after azo dyes to have attracted increasing attention to their toxicological and environmental effects with their increasing applications. It has been reported that some losses of the dyes are incurred during the dyeing process and a large quantity of partial dyes are emitted from the textile industry each year.2 These effluents are usually discharged in high concentrations and the toxicity of multiple pollutants, such as the complex aromatic structure of dyes, pigments, heavy metals, biocides, surfactants, multifarious organic and inorganic matter can give rise to complexation phenomena.3,4 Moreover, the contaminants may cause carcinogenic and teratogenic effects on human health, such as dermatitis, ulceration of skin and cancer and pollute water bodies too.5 Hence, there is a significant need to treat dye wastewater prior to discharge. Thus far, physical, chemical and biological processes are available for dyestuff treatment. Nevertheless, the development of physicochemical methods6 has been blocked due to the high-cost, time and material consumption, lack of management and disturbance by other wastewater components on the settlement of dye wastewater. There is a considerable need to put forward biological processes to dispose wastewater, particularly in light of their economic efficiency, eco-friendliness and extensive adaptability.7Various microorganisms in all biological processes such as fungi, bacteria and algae display efficiency in the degradation of synthetic dyestuff.8 Different types of bacteria such as Escherichia coli9 and Enterobacter sp. F NCIM 5545 (ref. 10) are employed for different dyes and dye wastewater. Next to bacteria, the role of fungi, such as Tinctoporellus sp. CBMAI 1061, Ganoderma sp. En3 and Irpex lacteus F17,11–13 in dye removal have been comprehensively studied. Certain microalgae including Gonium sp.8 and Chlorella pyrenoidosa14 have also been effectively applied in the disposal of dye-containing wastewater and water purification. Among microorganisms, microalga have potential applications in treating wastewater because of their adaptability in hostile conditions.7 According to Kim,15 algae-based piggery effluent treatment efficiently removed inorganic nutrients and accumulated algal biomass using an ozonation pretreatment with a dose of 1.1 mg O3 per mg C or light transmittance magnification in liquor. In view of the recycling utilization of the resource, algae-based treatment has been widely used on wastewater, which could make full use of the wastewater as a growth medium and promote carbon dioxide fixation. Besides the treatment of wastewater with significantly low cost, a great amount of microalgae can serve as crude materials of renewable fuels such as bio-oil.16
At present, some synthetic dyes, involving anthraquinone groups, have been processed by fungi and bacteria. Liu et al.1 found that alizarin red (anthraquinone) was destroyed and transformed into small molecules using chloroperoxidase obtained from Caldariomyces fumago. The biodecolorization and biodegradation of the dye simultaneously occurred during the treatment process with mixed fungi.17
In this study, the decolorization efficiency of one oleaginous Chlorella sorokiniana XJK was investigated with dye (disperse blue 2BLN) wastewater in an air-lift column photobioreactor. Various parameters including the dye concentration, biomass density, initial pH value, light intensity, and CO2 concentration were assayed, and the decolorization rate served as an index to achieve the optimal conditions. Considering the role of the enzyme on dye removal, we attempted to assay the ligninolity enzyme activities of Chlorella sorokiniana XJK during the algal growth. Furthermore, another important aspect of this work was to confirm the degradation mechanism of anthraquinone dye using UV-vis, FTIR and GC-MS analysis.
2. Materials and methods
2.1 Dye
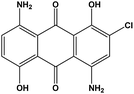
Disperse blue 2BLN (1,5-diamino-2-chloro-4,8-dihydroxy-9,10-anthracenedione) was purchased from Zhejiang Runtu Co. Ltd in a chemically pure form. It is one of the anthraquinone dyes and belongs to the group of the most durable dyes. It is widely utilized in dyeing polyester fibre and its blended fabrics. The chemical structure of disperse blue 2BLN used for decolorization research is shown in Fig. 1.2.2 Identification of microalga
One freshwater oleaginous microalga XJK was isolated from Keketuohai, XinJiang Province, China. The microalga was preliminary identified through the observation of its morphological characteristics, including cell size, shape, and chloroplast core protein number. Nucleic acid extraction of the algal cells was performed using the NuClean PlantGen DNA kit (Beijing Com Win Biotech Co., Ltd., China) according to the manufacturer's instructions. The internal transcribed spacer (ITS) was amplified by polymerase chain reaction (PCR) using the universal primer ITS-F (5′-GGAAGTAA AAGTCGTAACAAGG-3′) and ITS-R (5′-TCCTCCGCTTATTGATATGC-3′) according to the procedure of Katia et al.18 The amplified sequence was sent for analysis. The sequence obtained for the ITS gene was aligned to the published sequences obtained from GenBank using Clustal X 1.83. A neighbor-joining tree was constructed from the data by the bootstrap method (1000 replicates) using Mega 5.0 software.XJK was then conserved in Key Laboratory for Green Process of Chemical Engineering of Xinjiang Bingtuan, Shihezi University.
2.3 Culture medium and conditions
The basal culture medium BG-11 was modified as follows: NaNO3 – 0.83 g L−1, K2HPO4·3H2O – 0.09 g L−1, MgSO4·7H2O – 0.12 g L−1, CaCl2 – 27.2 mg L−1, citric-acid – 6.6 mg L−1, Fe(NH4)3(C6H5O7)2 – 6.0 mg L−1, NaCO3 – 0.02 g L−1, Na2EDTA·2H2O – 1.1 mg L−1, 0.2 mol L−1 and trace metal mix (H3BO3 – 2.86 g L−1, MnCl2·4H2O – 1.86 g L−1, ZnSO4·7H2O – 0.22 g L−1, NaMoO4·2H2O – 0.39 g L−1, CuSO4·5H2O – 0.08 g L−1 and Co(NO3)2·6H2O – 0.05 g L−1).The dye medium was prepared by adding disperse blue 2BLN at different concentrations into the modified medium BG11.
Seed cultured photoautotrophically in a 2 L Erlenmeyer flask containing 1.3 L medium was placed in an illumination chamber under 830 μmol (m2 s)−1 with a 12/0 h light/dark photoperiod in a constant temperature (25 °C) for 5 days. Aeration and mixing were achieved by sparging air through a 0.2 μm gas filter (Millipore) at a flow rate of 0.4 m3 h−1.
Photobioreactor and photoautotrophic cultivation conditions: A 1 L air-lift column photobioreactor (diameter: 5.00 cm; height: 55.00 cm) containing 700 mL working volume was used for the photoautotrophic cultivation. The inoculum biomass density was 9.9 × 106 cell mL−1. The culture conditions were maintained at room temperature (25 °C) under a white fluorescent light with 12/0 day/dark photoperiod for 6 days. The light intensity was approximately 830 μmol (m2 s)−1 at the surface of the photobioreactor for two sides illumination. Aeration and mixing were achieved by sparging air with CO2 through a 0.2 μm gas filter (Millipore) at a flow rate of 0.4 m3 h−1.
2.4 Optimization on decolorizing ability
According to biomass, lipid production, and the best dye decolorization rate of microalga XJK, the initial concentration of disperse blue 2BLN, incubating biomass density, initial pH, light intensity and CO2/air (v/v) ratio were optimized in sequence. All the experiments were conducted in triplicate.The optimum initial concentration of disperse blue 2BLN was firstly selected from 60–110 mg L−1 at intervals of 10 mg L−1. Next, an inoculum biomass density ranging from 3 × 106 to 1.5 × 107 cell per mL on the decolorization efficiency was examined, based on the selected dye concentration.
The dye medium with the dye was then prepared at pH 2, 4, 6, 8, 10 and 11, adjusted using 1 mol L−1 NaOH and HCl solution and measured with a pH meter (PHS-3C, INESA). The effect of the initial pH value on the decolorization efficiency was examined using the optimized initial dye concentration and biomass density.
Subsequently, the effect of light intensity on the decolorization efficiency was investigated, where the dye concentration (60 mg L−1), biomass density (8 × 106 cell mL−1) and initial pH value at 4 were kept constant. Different light densities were chosen: 830 μmol (m2 s)−1 (designated as high light HL), 470 μmol (m2 s)−1 (designated as low light LL), natural light (designated as NL), 0 μmol (m2 s)−1 (designated as photophobism PP), 830 μmol (m2 s)−1 for the first three days and 470 μmol (m2 s)−1 for the last three days (designated as decreasing photoperiod DP), 470 μmol (m2 s)−1 for the first three days and 830 μmol (m2 s)−1 for the last three days (designated as increasing photoperiod IP).
Finally, six levels of CO2/air (v/v) ratios (0%, 1%, 3%, 5%, 7%, and 9%) were used to determine the best concentration for maximum decolorization, where the dye concentration (60 mg L−1), biomass density (8 × 106 cell mL−1), initial pH (4) and 470 μmol (m2 s)−1 of light intensity were kept constant.
2.5 Determination of ligninolytic enzyme
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 rpm for 5 min. The supernatant fractions were used to determine the extracellular enzyme. Cells were frozen and ground with liquid nitrogen and reconstituted in buffer. The intracellular enzyme was then obtained by centrifugation at 10
000 rpm for 5 min. The supernatant fractions were used to determine the extracellular enzyme. Cells were frozen and ground with liquid nitrogen and reconstituted in buffer. The intracellular enzyme was then obtained by centrifugation at 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 rpm for 5 min, after removing the accumulated sedimentation. The enzymatic activities were determined spectrophotometrically. In all cases, the reaction mixture, in a final volume of 4 mL, contained 400 μL of sample.
000 rpm for 5 min, after removing the accumulated sedimentation. The enzymatic activities were determined spectrophotometrically. In all cases, the reaction mixture, in a final volume of 4 mL, contained 400 μL of sample.![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 M cm−1).
000 M cm−1).![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 590 M cm−1) at 25 °C.
590 M cm−1) at 25 °C.2.6 Analysis
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 rpm for 5 min to remove the cells. Absorbance of supernatant was first scanned by UV-vis from 400 to 800 nm using the culture medium as the control. The decolorization rate at the maximum absorption wavelength of 560 nm was defined as the changes of initial absorbance (A0) and absorbance of t moment (At) calculated as follows:
000 rpm for 5 min to remove the cells. Absorbance of supernatant was first scanned by UV-vis from 400 to 800 nm using the culture medium as the control. The decolorization rate at the maximum absorption wavelength of 560 nm was defined as the changes of initial absorbance (A0) and absorbance of t moment (At) calculated as follows:
 | (1) |
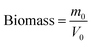 | (2) |
The total lipids were extracted following the protocol of Bligh and Dyer.22 Briefly, 10 mg of the lyophilized algal powder and distilled water of 1 mL served as layering and were added to 6 mL of a chloroform![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) methanol (2
methanol (2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1, v/v) solution. The mixture was shaken at 40 °C and 200 rpm for 4 hours. The solution was centrifuged at 10
1, v/v) solution. The mixture was shaken at 40 °C and 200 rpm for 4 hours. The solution was centrifuged at 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 rpm for 5 min. The chloroform layer was evaporated using a rotary evaporator (model RE-201D, Shanghai Dongxi) at 40 °C for 10 min. Herein, m1 was defined as the weight of extracted liquid and the evaporation flask, and m2 was defined as the weight of the evaporation flask dried in an oven at 105 °C for 2 hours.
000 rpm for 5 min. The chloroform layer was evaporated using a rotary evaporator (model RE-201D, Shanghai Dongxi) at 40 °C for 10 min. Herein, m1 was defined as the weight of extracted liquid and the evaporation flask, and m2 was defined as the weight of the evaporation flask dried in an oven at 105 °C for 2 hours.
The lipids content in % and lipids yield in mg (L d)−1 were determined according to the difference value of m1 and m2. The calculations were as follows:
 | (3) |
 | (4) |
Infrared spectra of the preprocessing liquid samples were analyzed using a Fourier transform infrared spectrometer (FTIR) (Magna-IR 750, Thermo Nicolet). KBr was used as the background; liquid samples were pointed at potassium bromide tableting to obtain their infrared spectra. The resolution and scan number were set at 8.0 cm−1 and 16, respectively. The scan range of the wavenumber was set at the mid IR region (4000–400 cm−1).
The degradation products of disperse blue 2BLN were also analyzed by GC-MS, which comprised of a GC system (model 7890A, Agilent) equipped with a HP-5MS capillary chromatographic column (30 × 0.32 × 0.25) and coupled to a MS system (model 5975C, Agilent). Helium served as the carrier gas at a flow rate of 0.8 mL min−1 and the injector temperature was 250 °C in splitless mode. The extracted samples were measured under the following temperature gradient:23 the initial column temperature was kept constant at 40 °C for 10 min, increased to 100 °C at a rate of 12 °C min−1, then ramped at 5 °C min−1 to 200 °C, further raised at 20 °C min−1 to 270 °C and held constant for 5 min, then with increase to 300 °C at 10 °C min−1 remained constant for 5 min. The degradation products in different periods were analyzed by comparing the GC-MS spectra patterns with those of standard mass spectra in the National Institute of Standards and Technology (NIST) library.
3. Results and discussion
3.1 Microalgae identification
According to its morphological characteristics, including cell size, shape, and chloroplast core protein number, the screened oleaginous microalga was named as Chlorella sp.-XJK. A micrograph of the microalga XJK is shown in Fig. 2a. Besides the characterization of lipid production (20%), it was found that XJK could also grow in the dye medium with a lipid production of 24% (Fig. 2b). When compared to the basal modified medium BG11, the accumulation of biomass evidently decreased in the dye medium because the presence of chlorides, complex aromatic structures, heavy metals, and synthetic dyes limit the light penetration and inhibit the growth of the microalgae.5 However, the production of lipids was slightly increased in XJK with the existence of the dye. Gopalakrishnan et al. found that lipids act as a secondary metabolite, maintaining specific cell signaling pathways and membrane functions while responding to changes in the environment.7 That is, the screened microalga XJK can be used to remove the synthetic dye from water and accumulate lipids for biofuels at the same time.Further identification of XJK was carried out by PCR amplification and subsequent DNA sequencing. The length of the ITS sequence was obtained as 751 bp (Fig. 2c). The ITS sequence of XJK showed 99% sequence homology with Chlorella sorokiniana RP JX014220.1 (Fig. 2d). It was thus named Chlorella sorokiniana XJK (C. sorokiniana XJK). Related sequences were downloaded and used to build a phylogenetic tree using Mega 5.0 software.
3.2 Optimization on dye decolorization
Environmental variables can remarkably affect the dye removal process by microalga. A series of experiments were carried out and the optimum conditions of disperse blue 2BLN removal were acquired according to the decolorization rate.According to the degradation rate of dye using C. sorokiniana XJK, the optimum conditions were described as follows: dye conc. = 60 mg L−1, biomass density = 8 × 106 cell per mL, pH = 4, light intensity 470 μmol (m2 s)−1 and CO2/air (v/v) ratio = 1%, which gave efficient decolorization and the desired decolorization rate was 83%.
3.3 Analysis
![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O and C
O and C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C.37 Therefore, the large chromophore structure of disperse blue 2BLN was degraded into smaller molecules.
C.37 Therefore, the large chromophore structure of disperse blue 2BLN was degraded into smaller molecules.| Time (d) | Laccase (U mL−1) | MnP (U mL−1) | LiP (U mL−1) | |||
|---|---|---|---|---|---|---|
| Intracellular | Extracellular | Intracellular | Extracellular | Intracellular | Extracellular | |
| 2 | 0.05 | 0.22 | 0.87 | 2.40 | 3.53 | 3.53 |
| 3 | 0.08 | 0.48 | 1.53 | 3.27 | 3.25 | 2.17 |
| 4 | 0.19 | 1.99 | 3.92 | 4.86 | 2.99 | 1.09 |
| 5 | 0.21 | 3.89 | 3.27 | 3.49 | 1.09 | 0.27 |
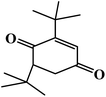
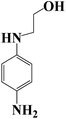
The degradation pathway was proposed starting from the products numbered 2 and 3, which may be isomers. The m/z value of 191 belonged to 2,6-di-tert-butyl-p-benzoquinone (number 4) generated by an open-loop, with the molecular formula being C17H24O3 (Fig. 5a); the m/z value of 149 may extrapolate to 2-(2-aminoanilino) ethanol (number 5) (Fig. 5b). The m/z value of 88 and 59 belong to small molecular products. In addition, these products could further be degraded to give products such as CO2 and H2O (ref. 42) in other ways. However, numerous peaks were indistinguishable and these peaks may come from the secretion of Chlorella sorokiniana XJK by the degrading enzyme.43
 | ||
| Fig. 5 The GC-MS spectra of the identified compounds in the disperse blue 2BLN solution of the algae-based treatment. | ||
Disperse blue 2BLN was confirmed to be broken down by UV-vis, FTIR, GC-MS and enzymatic analysis. Experimental evidence for dye degradation: the characteristic absorption peak of dye had a dramatic decrease (UV-vis), the chromophore C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O was fractured (FTIR) and some new peaks were found (GC-MS). It is worth mentioning that the ligninolytic enzyme played an important role in dye degradation, particularly laccase and MnP.
O was fractured (FTIR) and some new peaks were found (GC-MS). It is worth mentioning that the ligninolytic enzyme played an important role in dye degradation, particularly laccase and MnP.
4. Conclusions
The microalgae named C. sorokiniana XJK used in this study was exposed to disperse blue 2BLN under various operating conditions. C. sorokiniana XJK played a vital role in dye degradation, in addition to the accumulation of bioenergy. Under the optimal conditions, XJK can remove the dye with a maximum 83% decolorization rate and obtain 570 mg L−1 of biomass and 43% of lipid content at same time. Above all, during the enzyme production function as well dye removal, laccase and MnP activity played essential roles during the decolorization of the dye by C. sorokiniana XJK. Moreover, the results of UV-vis, FTIR and GC-MS analysis observed that the anthraquinone structure was actually broken down into small molecules and some products were formed during the degradation process.Acknowledgements
This study was financially supported by the National Natural Science Foundation of China (21466032) and Scientific Research Foundation for Changjiang Scholars of Shihezi University.References
- L. Liu, J. Zhang, Y. Tan, Y. Jiang, M. Hu, S. Li and Q. Zhai, Chem. Eng. J., 2014, 244, 9–18 CrossRef CAS.
- D. A. Yaseen and M. Scholz, Ecol. Eng., 2016, 94, 295–305 CrossRef.
- S. Rangabhashiyam, E. Suganya, N. Selvaraju and L. A. Varghese, World J. Microbiol. Biotechnol., 2014, 30, 1669–1689 CrossRef CAS PubMed.
- N. Colin, A. Maceda-Veiga, N. Flor-Arnau, J. Mora, P. Fortuno, C. Vieira, N. Prat, J. Cambra and A. de Sostoa, Ecotoxicol. Environ. Saf., 2016, 132, 295–303 CrossRef CAS PubMed.
- E. Baldev, D. MubarakAli, A. Ilavarasi, D. Pandiaraj, K. A. Ishack and N. Thajuddin, Colloids Surf., B, 2013, 105, 207–214 CrossRef CAS PubMed.
- S. C. D. Sharma, Q. Sun, J. Li, Y. Wang, F. Suanon, J. Yang and C. P. Yu, Int. Biodeterior. Biodegrad., 2016, 112, 88–97 CrossRef CAS.
- V. Gopalakrishnan and D. Ramamurthy, BioMed Res. Int., 2014 DOI:10.1155/2014/529560.
- G. Boduroglu, N. K. Kilic and G. Donmez, Environ. Technol., 2014, 35, 2410–2415 CrossRef CAS PubMed.
- M. Cerboneschi, M. Corsi, R. Bianchini, M. Bonanni and S. Tegli, Appl. Microbiol. Biotechnol., 2015, 99, 8235–8245 CrossRef CAS PubMed.
- C. R. Holkar, A. B. Pandit and D. V. Pinjari, Bioresour. Technol., 2014, 173, 342–351 CrossRef CAS PubMed.
- J. P. G. Rodriguez, D. E. Williams, I. D. Sabater, R. C. Bonugli-Santos, L. D. Sette, R. J. Andersen and R. G. S. Berlinck, RSC Adv., 2015, 5, 66360–66366 RSC.
- R. Lu, L. Ma, F. He, D. Yu, R. Fan, Y. Zhang, Z. Long, X. Zhang and Y. Yang, Bioprocess Biosyst. Eng., 2015, 39, 381–390 CrossRef PubMed.
- X. Yang, J. Zheng, Y. Lu and R. Jia, Environ. Sci. Pollut. Res., 2016, 23, 9585–9597 CrossRef CAS PubMed.
- V. V. Pathak, R. Kothari, A. K. Chopra and D. P. Singh, J. Environ. Manage., 2015, 163, 270–277 CrossRef CAS PubMed.
- H. C. Kim, W. J. Choi, S. K. Maeng, H. J. Kim, H. S. Kim and K. G. Song, Bioresour. Technol., 2014, 159, 128–135 CrossRef CAS PubMed.
- P. S. Kumar, J. Pavithra, S. Suriya, M. Ramesh and K. A. Kumar, Desalin. Water Treat., 2015, 55, 1342–1358 CAS.
- D. G. Ülküye and D. Gönül, Desalin. Water Treat., 2013, 51, 3597–3603 CrossRef.
- S. Katia, A. L. Lewis, V. Elie, M. Isabella and L. R. Nicoletta, J. Phycol., 2015, 51, 1172–1188 CrossRef PubMed.
- N. Hatvani and I. Mécs, Process Biochem., 2001, 37, 491–496 CrossRef.
- H. M. N. Iqbal, M. Asgher and H. N. Bhatti, BioResources, 2011, 6, 1273–1287 CAS.
- H. Bermek, İ. Gülseren, K. Li, H. Jung and C. Tamerler, World J. Microbiol. Biotechnol., 2004, 20, 345–349 CrossRef CAS.
- E. G. Bligh and W. J. Dyer, Can. J. Biochem. Physiol., 1959, 37, 911–917 CrossRef CAS PubMed.
- D. Rajkumar, B. J. Song and J. G. Kim, Dyes Pigm., 2007, 72, 1–7 CrossRef.
- R. Waqas, M. Arshad, H. N. Asghar and M. Asghar, Int. J. Agric. Biol., 2015, 17, 803–808 CrossRef CAS.
- Q. He, H. Yang and C. Hu, Bioresour. Technol., 2015, 198, 876–883 CrossRef CAS PubMed.
- M. T. Ale, M. Pinelo and A. S. Meyer, Prep. Biochem. Biotechnol., 2014, 44, 242–256 CrossRef CAS PubMed.
- P. Feng, Z. Deng, Z. Hu and L. Fan, Bioresour. Technol., 2011, 102, 10577–10584 CrossRef CAS PubMed.
- S. Mahalakshmi, D. Lakshmi and U. Menaga, Int. J. Environ. Res., 2015, 9, 735–744 CAS.
- S. V. Mohan, Y. V. Bhaskar and J. Karthikeyan, Int. J. Environ. Pollut., 2004, 21, 211–222 CrossRef CAS.
- K. V. Kumar, S. Sivanesan and V. Ramamurthi, Process Biochem., 2005, 40, 2865–2872 CrossRef CAS.
- S. Xue, Z. Su and W. Cong, J. Biotechnol., 2011, 151, 271–277 CrossRef CAS PubMed.
- Y. Xiao, J. Zhang, J. Cui, X. Yao, Z. Sun, Y. Feng and Q. Cui, Algal Res., 2015, 11, 55–62 CrossRef.
- G. Yildiz and Ş. Dere, Acta Oceanol. Sin., 2015, 34, 125–129 CrossRef CAS.
- K. G. Schulz, J. Barcelos e Ramos, R. E. Zeebe and U. Riebesell, Biogeosciences Discuss, 2009, 6, 4441–4462 CrossRef.
- J. Fan, H. Xu, Y. Luo, M. Wan, J. Huang, W. Wang and Y. Li, Appl. Microbiol. Biotechnol., 2015, 99, 2451–2462 CrossRef CAS PubMed.
- J. M. Fanchiang and D. H. Tseng, Chemosphere, 2009, 77, 214–221 CrossRef CAS PubMed.
- C. W. M. Yuen, S. K. A. Ku, P. S. R. Choi, C. W. Kan and S. Y. Tsang, Res. J. Text. Apparel, 2005, 9, 26–38 CrossRef.
- F. N. Memon and S. Memon, Pak. J. Anal. Environ. Chem., 2012, 13, 148–158 CAS.
- R. Dosdall, V. Hahn, F. Preuß, H. Kreisel, J. Miersch and F. Schauer, Int. Biodeterior. Biodegrad., 2014, 96, 174–180 CrossRef.
- G. McMullan, C. Meehan, A. Conneely, N. Kirby, T. Robinson, P. Nigam, I. M. Banat, R. Marchant and W. F. Smyth, Appl. Microbiol. Biotechnol., 2001, 56, 81–87 CrossRef CAS PubMed.
- M. M. Martorell, H. F. Pajot and L. I. C. de Figueroa, Int. Biodeterior. Biodegrad., 2012, 66, 25–32 CrossRef CAS.
- S. Saraf and B. Thomas, Process Saf. Environ. Prot., 2007, 85, 360–364 CrossRef CAS.
- J. S. Ye, H. Yin, J. Qiang, H. Peng, H. M. Qin, N. Zhang and B. Y. He, J. Hazard. Mater., 2011, 185, 174–181 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2016 |