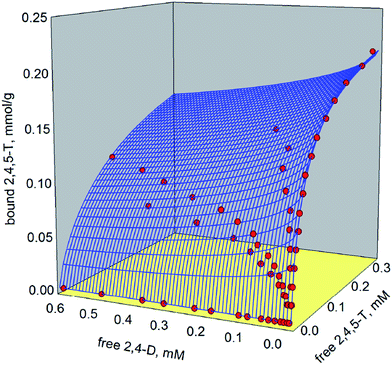
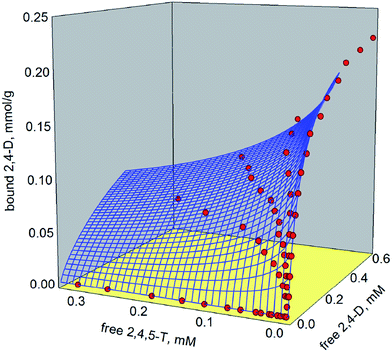
Full vs. partial competitive binding behaviour in molecularly imprinted polymers. The case for a chlorinated phenoxyacids-binding polymer
G. Spano,
C. Giovannoli,
F. Di Nardo,
L. Anfossi and
C. Baggiani *
*
Department of Chemistry, University of Torino, Torino 10125, Italy. E-mail: claudio.baggiani@unito.it; Fax: +39-11-6707615; Tel: +39-11-6705266
First published on 8th August 2016
Abstract
The assessment of the molecular recognition properties of a molecularly imprinted polymer (MIP) towards its target molecule requires the determination of its binding thermodynamic parameters through the measurement of the binding isotherm that governs the binding equilibrium. This measurement is usually performed assuming that the binding properties of a MIP are not affected by the simultaneous presence of two or more ligands of comparable affinity for the binding sites (full competitivity). The validity of this requirement is particularly critical when the selectivity of a MIP is evaluated. In fact, if conditions of full competitiveness fail to exist, the selectivity measured in the presence of one ligand at a time can be markedly different from the selectivity measured in the simultaneous presence of several different ligands. The goal of this work is to test whether the binding parameters of a MIP for its template molecule result are modified by the presence of a competitor of comparable affinity. As template and competitor molecules the herbicides 2,4,5-trichlorophenoxyacetic acid (2,4,5-T) and 2,4-dichlorophenoxyacetic acid (2,4-D) were chosen. The binding data obtained by partition experiments were fitted by using the general Freundlich–Langmuir competitive isotherm model for multiple ligands. The results show that the equilibrium constant (Keq) and the maximum binding site concentration (Bmax) calculated in both conditions are statistically indistinguishable. Thus, it can be concluded that the thermodynamics that govern the binding behavior of a MIP works in full competitive conditions.
Introduction
Molecularly imprinted polymers (MIPs) show selective molecular recognition properties towards the template molecule and, frequently, towards other strictly structurally related molecules too.As documented in the literature, the most important application of MIPs is in the field of solid phase extraction where they are used as selective extraction materials in the well-known MISPE technique.1–4 However, the interest has recently moved towards the use of MIPs as plastic antibodies to set up MIP-based assays in a competitive format where a competition is achieved between it and a labelled analyte analogue.5–8 The feasibility and the analytical performance of a competitive assay strictly depend on the validity of some theoretical assumptions. Among them, one of the most important is the occurrence of complete competitive conditions which warrant a balanced competition between the analyte and the tracer used for the same class of binding sites. If this assumption is not valid the competition fails because the binding sites bound by the analyte are not the same as those bound by the tracer used. This does not allow a quantitative relationship between the analytical signal and the concentration of the analyte taking part in the competition. This prevents to develop a competitive MIP-based assay and consequently to think MIPs as potential substituted of natural antibodies. For this reason, the experimental evidence of a behaviour like that of the antibodies is a fundamental step for considering the development of MIP-based assay feasible.
In order to quantitatively evaluate such properties, one of the most popular method is based on the equilibrium batch rebinding which is based on the experimental measurement of the partition coefficient for the template molecule in the presence of a suspension of the binding polymer.9 However, even if very popular, such method can be misleading because it can be potentially lead to large errors of evaluation on the true binding properties of MIPs. In fact, the numerical value of the partition coefficient heavily depends on the relative amounts of ligand and polymer put to the equilibrium. As recently shown,10 such dependence cannot be avoided or neglected, as it is due to the non-linear nature of the binding isotherm which describes the binding behaviour of the polymer. It is therefore clear that to have a correct assessment of the molecular recognition properties of a MIP, it is necessary to measure the binding thermodynamic parameters, i.e. the equilibrium constant (Keq) and the concentration (density) of binding sites (Bmax) related to its binding isotherm. These parameters can be obtained by evaluating the ligand binding process through the experimental measurement of the dynamic equilibrium between a fixed amount of polymer and varying amounts of ligand distributed in a wide concentration range. It follows that, once a given binding isotherm model is considered suitable to describe the binding between the ligand and the MIP, it is possible to define the binding properties of such polymer regardless of the stoichiometric concentration of the ligands and the amount of the polymer itself.
As regards the measurement of the ligand–polymer equilibrium, it should be noted that it is usually performed with a single ligand, i.e. in non-competitive conditions.9 In such experiments it is implied that the binding properties of a MIP are not influenced by the simultaneous presence of two or more ligands of comparable affinity for the binding sites. This assumption is valid only if all the ligands are capable of binding – with less or more strength – to all the binding sites, without any preference because only in this case a condition of full competition between different ligands exists for all the binding sites. On the contrary, if some ligands are not able to bind to all the binding sites, but they exhibit a binding preference for some sub-classes of them, then a condition of partial competition is present between different ligands. This becomes particularly critical when the selectivity of a MIP has to be assessed since selectivity is usually measured by the evaluation of the binding isotherms for several ligands through a set of separate partition experiments. But, when a condition of partial competition exists, selectivity measured in the presence of one ligand at a time can be markedly different from selectivity measured in the simultaneous presence of several different ligands.
The presence of binding behaviours based on full or partial competitive conditions has outstanding implications in the development of MIP-based assays mimicking the traditional immunoassay which is based on the competition between the analyte and the tracer molecules. In fact, as such assays should work in deficiency of binding sites with respect to the amount of competing molecules, only a fully indistinguishable behaviour of the binding properties towards both the analyte and tracer molecules, can assure a correct analytical response.6,7
It is quite remarkable that despite the very large number of papers dealing with the measurement and evaluation of MIP binding properties, at the present no information is available about the possible presence of partial competitive conditions during the measurement of ligand–polymer equilibrium in the presence of multiple binding ligands. For this reason, the goal of this work is to experimentally test whether the binding parameters of a MIP for its template molecule can be modified by the presence of a competitor. As the related MIP has been extensively studied by our group,11,12 the herbicides 2,4,5-trichlorophenoxyacetic acid (2,4,5-T) and 2,4-dichlorophenoxyacetic acid (2,4-D) were chosen as template and competitor, respectively.
Experimental
Materials
2,4-Dichlorophenoxyacetic acid (2,4-D) and 2,4,5-trichlorophenoxyacetic acid (2,4,5-T), were from Sigma-Aldrich-Fluka (Milan, Italy). Acetonitrile (HPLC-gradient grade) and acetic acid were from VWR International (Milan, Italy).Phenoxyacid stock solutions were prepared by dissolving 100 mg of substance in 25 mL of dry acetonitrile and stored in the dark at −20 °C. Working solutions were prepared by diluting the standard immediately before use. The water used was ultra-purified in Purelab Prima System from Elga (Marlow, UK). The 0.22 μm nylon membranes used for filtration were from Alltech Italia (Milano, Italy).
The 2,4,5-T imprinted polymer was prepared as previously reported in literature.11 The HPLC apparatus LaChrom Elite system composed of a programmable binary pump L-2130, an auto-sampler L-2200, a UV detector L-2400, and provided with EZChrom Elite software for the instrumental programming, data acquisition and data processing was from Merck-Hitachi (Milano, Italy).
Reverse-phase liquid chromatography
Reversed-phase HPLC with UV detection at 280 nm was used to measure phenoxyacids. The analytical column was a 100 × 4.6 mm, 5 μm, LichroCART RP-C18 PUROSPHERE from VWR (Milano, Italy). The mobile phase was composed of acetonitrile–water–acetic acid (75 + 25 + 1 v/v/v) and elutions were performed under isocratic conditions at a flow rate of 0.5 mL min−1. The sample volume injected was 5 μL. Reference standard solutions of phenoxyacids at concentrations of 0.50, 0.75, 1.0, 2.5, 5.0, 7.5, 10, 25, 50 and 75 μg mL−1 were analysed three times consecutively and peak areas were plotted against concentration. Calibration plots were drawn by using weighted linear regression (weight = 1/conc).Equilibrium batch rebinding
About 40 mg of polymer were exactly weighed in 4 mL flat bottom amber glass vials. Then, 2.00 mL of acetonitrile solutions containing increasing amounts of phenoxyacids ranging from 100 to 1500 μg, alone or in binary mixture (2,4,5-T + 2,4-D, 1 + 2, 1 + 1 and 2 + 1 mol mol−1) were added and sonicated for 10 min. Then, the vials were incubated overnight at room temperature under continuous agitation on a horizontal rocking table. Therefore, the solutions were filtered on 0.22 μm nylon membranes, and the free amounts of phenoxyacids were measured by HPLC analysis. Each experimental point of the binding isotherms was assessed as the average of three repeated measures.The binding isotherms were calculated by using SigmaPlot 12 (Systat Software Inc., Richmond, CA, USA). Non-linear least square fitting was applied to the averaged experimental data, by using a Freundlich–Langmuir (FL) isotherm model generalized for k simultaneous ligands:13
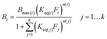 | (1) |
Results and discussion
It has been previously shown that a polymer imprinted with 2,4,5-T exhibit good molecular recognition properties towards both the template and the related ligand 2,4-D.5,6 Since such properties were evaluated in terms of chromatographic behaviour only, we decided to measure the binding parameters for 2,4,5-T and 2,4-D by equilibrium batch rebinding in non-competitive conditions. By the fitting of the experimental data with the isotherm model described by eqn (1) (k = 1). The affinity towards 2,4,5-T resulted to be about twice the affinity towards 2,4-D (Keq = 18.9 ± 2.0 and 10.2 ± 1.4 μM−1 for 2,4,5-T and 2,4-D, respectively), confirming the good selectivity of the MIP towards its template molecule. Interestingly, the binding site densities resulted to be nearly the same for 2,4,5-T and 2,4-D (Bmax = 246 ± 9 and 267 ± 12 μmol g−1 for 2,4,5-T and 2,4-D, respectively). This can be considered an indirect indication that both the ligands are recognized by the same binding sites. Thus, the ligands can effectively compete for the same binding sites if present simultaneously.Subsequently, the equilibrium batch rebinding experiments were performed in the presence of both the ligands, with 2,4,5-T/2,4-D molar ratios ranging from 1 + 2 to 2 + 1. It is rather obvious that the fitting of the experimental data through the binding isotherm model with k = 1 (thus deliberately ignoring the presence of the additional competitive ligand) produces apparent binding parameters whose numerical values should be affected by the presence of a competitive ligand. In fact, it is possible to observe in Fig. 1 and 2 that the apparent affinity constants increase and the apparent binding site densities decrease in proportion with the increase of the amount of the competitive ligand. More interestingly, the numerical ratio between the apparent binding site densities, Bmax(2,4-D)/Bmax(2,4,5-T) (Fig. 1, insert) is essentially constant, thus not affected by the presence of a competitive ligand. On the contrary, the numerical ratio between the apparent equilibrium binding constants, Keq(2,4-D)/Keq(2,4,5-T) (Fig. 2, insert), seems to be influenced by the presence of a competitive ligand, as its value increases with increased amount of the competitive ligand itself. However, such increase is significant for the last binding isotherm only, where the competitive ligand is in a molar excess of 2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1. To sum up, the numerical results obtained by using a Freundlich–Langmuir isotherm model suitable for non-competitive conditions only (k = 1) show that such model is able to give some hints, but it is not sufficient to definitively decide whether the binding parameters of a MIP for its template molecule are influenced by the presence of a competitive ligand, and a more comprehensive model should be adopted in any case.
1. To sum up, the numerical results obtained by using a Freundlich–Langmuir isotherm model suitable for non-competitive conditions only (k = 1) show that such model is able to give some hints, but it is not sufficient to definitively decide whether the binding parameters of a MIP for its template molecule are influenced by the presence of a competitive ligand, and a more comprehensive model should be adopted in any case.
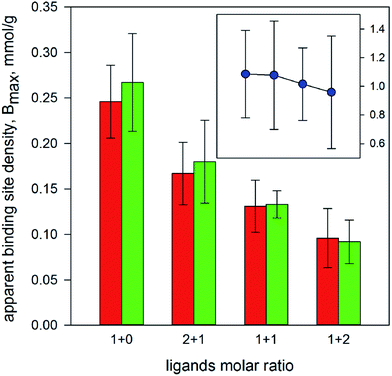 | ||
| Fig. 1 Apparent binding site density measured for 2,4,5-T (red bars) and 2,4-D (green bars) vs. ligands molar ratio. In the insert: numerical ratio between the apparent binding site densities. | ||
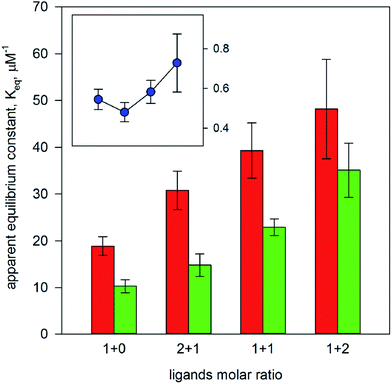 | ||
| Fig. 2 Apparent equilibrium constant measured for 2,4,5-T (red bars) and 2,4-D (green bars) vs. ligands molar ratio. In the insert: numerical ratio between the apparent equilibrium constants. | ||
The model described by eqn (1) can be potentially expanded to take into account an arbitrarily large number of ligands, generating k different isotherm models for k different ligands. However, its main drawback becomes apparent when the number of involved ligands is greater than 2 (k > 2). In fact, the resulting k binding isotherms are graphically described by k different hypersurfaces with dimensionality k + 1, which obviously cannot be visually represented. Furthermore, to achieve acceptable low values for the standard fit error, it is necessary to dramatically increase the number of experimental points, that is quite impractical. For these reasons, we limited the number of competing ligands to k = 2, by performing equilibrium binding experiment with binary mixtures of 2,4-D and 2,4,5-T. In these conditions, eqn (1) generates two isotherm models, describing the equilibrium binding of 2,4,5-T in the presence of 2,4-D (2) and 2,4-D in the presence of 2,4,5-T (3), respectively:
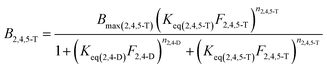 | (2) |
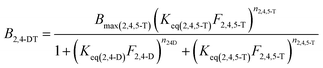 | (3) |
The dataset generated by the combination of the equilibrium batch rebinding data obtained in the presence and in the absence of competing ligands was fitted with these isotherm models, obtaining two 3-D binding surfaces, reported in Fig. 3 and 4. The corresponding binding parameters are reported in Table 1. The statistical t-test used to check the presence of a statistically significant difference between the binding parameters calculated in the absence and in the presence of a competing ligand shows that the differences in the values of the parameters are not great enough to reject the possibility that these differences are due to data random variability (Keq(2,4,5-T): t = 0.609, P = 0.544: Keq(2,4-D): t = 0.215, P = 0.830; Bmax(2,4,5-T): t = 0.631, P = 0.530; Bmax(2,4-D): t = 0.117, P = 0.907; n2,4,5-T: t = 1.263, P = 0.210; n2,4-D: t = 0.290, P = 0.773). Thus, there is not a statistically significant difference between the binding parameters calculated in the absence (k = 1) and in the presence (k = 2) of a competing ligand.
| Binding isotherm model | Ligand | Competitor | Bmax, μmol g−1 | Keq, μM−1 | n | Adj. r2 | SEE |
|---|---|---|---|---|---|---|---|
| FL, k = 1 | 2,4,5-T | — | 246 ± 9 | 18.9 ± 2.0 | 0.872 ± 0.045 | 0.996 | 3.58 |
| FL, k = 1 | 2,4-D | — | 267 ± 12 | 10.2 ± 0.4 | 0.834 ± 0.053 | 0.994 | 4.73 |
| FL, k = 2 | 2,4,5-T | 2,4-D | 264 ± 14 | 15.7 ± 2.5 | 0.772 ± 0.037 | 0.991 | 5.34 |
| FL, k = 2 | 2,4-D | 2,4,5-T | 268 ± 6 | 9.9 ± 0.7 | 0.820 ± 0.020 | 0.997 | 3.05 |
Conclusions
The binding data obtained by the partition experiments under competitive and non-competitive conditions and fitted by using a general Freundlich–Langmuir competitive isotherm model for multiple ligands show that the numerical values calculated for the equilibrium binding constant, Keq, and the binding site density, Bmax, are statistically indistinguishable. As a consequence, it can be concluded that the thermodynamic parameters governing the binding behaviour of a MIP towards a given ligand are not significantly conditioned by the presence of one or more competitors. This result, for the first time clearly demonstrated in the literature, has deep implications on the evaluation of the binding selectivity, which is related to the binding properties of the polymer. In fact, the substantial independence of the binding parameters by the number of the effectively present ligands makes possible the correct measurement of the selectivity by performing simple batch rebinding experiments with one ligand at a time. Moreover, such results are relevant for the correct set-up of whatever analytical assay based on the competition between an analyte and a tracer for the imprinted binding sites, confirming the feasibility of immunoassay-like methods based on imprinted polymers.Notes and references
- J. Haginaka, J. Sep. Sci., 2009, 32, 1548 CrossRef CAS PubMed.
- E. Turiel and A. Martin-Esteban, Anal. Chim. Acta, 2010, 668, 87 CrossRef CAS PubMed.
- J. Fu, L. Chen, J. Li and Z. Zhang, J. Mater. Chem. A, 2015, 3, 13598 CAS.
- L. Chen, X. Wang, W. Lu, X. Wu and J. Li, Chem. Soc. Rev., 2016, 45, 2137 RSC.
- Y. Ge and A. P. F. Turner, Chem.–Eur. J., 2009, 15, 8100 CrossRef CAS PubMed.
- M. C. Moreno-Bondi, M. E. Benito-Pena, J. L. Urraca and G. Orellana, in Molecular Imprinting, Topics in Current Chemistry, ed. K. Haupt, Springer Verlag, Hilderberg, 2012, vol. 325, pp. 165–266 Search PubMed.
- Z.-X. Xu, H.-J. Gao, L.-M. Zhang, X.-Q. Chen and X.-G. Qiao, J. Food Sci., 2011, 76, R69 CrossRef CAS PubMed.
- T. S. Bedwell and M. J. Whitcombe, Anal. Bioanal. Chem., 2016, 408, 1735 CrossRef CAS PubMed.
- J. A. Garcia-Calzon and M. E. Diaz-Garcia, Sens. Actuators, B, 2007, 123, 1180 CrossRef CAS.
- O. K. Castell, D. A. Barrow, A. R. Kamarudin and C. J. Allender, J. Mol. Recognit., 2011, 24, 1115 CrossRef PubMed.
- C. Baggiani, G. Giraudi, C. Giovannoli, F. Trotta and A. Vanni, J. Chromatogr. A, 2000, 883, 119 CrossRef CAS PubMed.
- C. Baggiani, L. Anfossi, C. Giovannoli and C. Tozzi, Talanta, 2004, 62, 1029 CrossRef CAS PubMed.
- U. Rusch, M. Borkovec, J. Daicic and W. H. van Riemsdijk, J. Colloid Interface Sci., 1997, 191, 247 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2016 |