Label-free, homogeneous, and ultrasensitive detection of pathogenic bacteria based on target-triggered isothermally exponential amplification
Tingting Qiua,
Yu Wang*a,
Jinghua Yub,
Su Liuc,
Hongzhi Wangb,
Yuna Guob and
Jiadong Huangab
aSchool of Biological Sciences and Technology, University of Jinan, Jinan 250022, P. R. China. E-mail: chm_huangjd@ujn.edu.cn; Fax: +86-531-82769122; Tel: +86-531-89736122
bKey Laboratory of Chemical Sensing & Analysis in Universities of Shandong, School of Chemistry and Chemical Engineering, University of Jinan, Jinan 250022, P. R. China
cSchool of Resources and Environment, University of Jinan, Jinan 250022, P. R. China
First published on 22nd June 2016
Abstract
In this work, a simple, isothermal, and ultrasensitive homogeneous colorimetric sensor for pathogenic bacteria detection has been developed on the basis of target-triggered exponential amplification reaction (EXPAR). An aptamer–primer probe (arched probe) containing anti-target aptamer and a primer sequence, which is released under the challenging of target, is used for recognizing target and triggering EXPAR-based polymerase elongation. Due to EXPAR coupled DNAzyme amplification strategy, the presence of target pathogenic bacteria leads to the formation of numerous G-quadruplex oligomers in solution, which folds into G-quadruplex/hemin complexes with the help of K+ and hemin, thus generating extremely strong catalytic activity toward H2O2 and giving a remarkably strong UV-vis absorption. This work is a novel design that EXPAR coupled DNAzyme amplification technique has been integrated into colorimetric assay for detecting pathogenic bacteria. Under optimal conditions, the proposed biosensor exhibits ultrahigh sensitivity toward target pathogenic bacteria with detection limits of 80 cfu mL−1. Besides, our biosensor also shows high selectivity toward target pathogenic bacteria and has the advantages in its low cost, simplified operations without the need of labeling steps and additional labile reagents. Hence, the EXPAR coupled DNAzyme amplification-based colorimetric method might create a useful and practical platform for detecting pathogenic bacteria and related food safety analysis and environmental monitoring.
1. Introduction
Pathogenic bacteria have always been a human health threat of global proportions and widely responsible for many foodborne diseases.1,2 According to the Center for Disease Control estimates, in the United States, 76 million cases of foodborne illness occur annually, resulting in approximately 325![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 hospitalizations and 5000 deaths.3 Pathogenic bacteria is the main cause of gastroenteritis and its infection even is fatal for some immuno-compromised patients.4 The conventional method for pathogenic bacteria detection is a culturing-based method, which is generally time-consuming and labor-intensive, and cumbersome with low throughput.5 Recent work in developing novel pathogen detection methods have involved enzyme-linked immunosorbent assay (ELISA),4 quartz crystal microbalance (QCM) assay,6 flow cytometry assay,7 protein microassay,8 and surface plasmon resonance (SPR)-based assay.9,10 Although these methods may combine some attractive features such as high sensitivity, multiplexing ability, or rapid detection, they may require laborious pretreatment of samples including cell lysis, expensive instruments and reagents, and trained technical personnel.11 Thus, there is great demand to develop convenient, low cost and highly sensitive assay methods for the determination of pathogenic bacteria.
000 hospitalizations and 5000 deaths.3 Pathogenic bacteria is the main cause of gastroenteritis and its infection even is fatal for some immuno-compromised patients.4 The conventional method for pathogenic bacteria detection is a culturing-based method, which is generally time-consuming and labor-intensive, and cumbersome with low throughput.5 Recent work in developing novel pathogen detection methods have involved enzyme-linked immunosorbent assay (ELISA),4 quartz crystal microbalance (QCM) assay,6 flow cytometry assay,7 protein microassay,8 and surface plasmon resonance (SPR)-based assay.9,10 Although these methods may combine some attractive features such as high sensitivity, multiplexing ability, or rapid detection, they may require laborious pretreatment of samples including cell lysis, expensive instruments and reagents, and trained technical personnel.11 Thus, there is great demand to develop convenient, low cost and highly sensitive assay methods for the determination of pathogenic bacteria.
Colorimetric methods, which have potential advantages over other analytical techniques such as high portability and affordability, rapid response, and low cost, are particularly appealing for pathogenic bacteria identification.12 Several colorimetric-based strategies for pathogenic bacteria detection have been reported by using nanoparticles (NPs)13 or gold nanorods14 as signal reporters. Although these methods are attractive because of their unique physical and chemical properties, catalytic DNA molecules (DNAzymes) have found growing interest as signal reporters and amplifiers for biosensing for the past few years. DNA aptamers15 and DNAzymes16 called functional nucleic acid (FNA),17 represent an emerging class of biomolecules that finds great advantages in bioassay development. DNA aptamers, selected by SELEX (systematic evolution of ligands by exponential enrichment), are single-stranded DNA molecules that can specifically bind to a non-nucleic acid target.18 Bacterium-based aptamer selection techniques targeting whole live bacteria in suspension have been carried out to create ssDNA aptamers. The selection of aptamers against live S. typhimurium bacterial cells meets the needs of the practical detection more than aptamer against dead bacteria in public health safety.19 As the DNA molecules with catalytic functions,20 DNAzymes have been extensively studied in the recent years, which are dependent on the advantages of simple preparation, low cost, and high thermal stability.21,22 In the paper, the G-quadruplex/hemin DNAzyme, which catalyzes the oxidation of 2,2′-azino-bis(3-ethylbenzothiazoline)-6-sulfonic acid (ABTS2−) by H2O2 to a colored product ABTS˙−,23,24 has been used as a signal reporter to design colorimetric biosensor for pathogenic bacteria detection.
In addition, FNA have high affinity, specificity, and chemical stability, and FNA-based assays can be integrated with a variety of established molecular biology tools such as exponential amplification reaction (EXPAR),25,26 which is an alternative amplification technique and has high amplification efficiency by combination of polymerase strand extension and single-strand nicking.27 Indeed, the simplicity and versatility have made FNA an emerging candidate in recent years for bioassay platform development including fluorescent,28 electrochemical,29 colorimetric,30 and others. The colorimetric FNA assays can be readily applied to homogeneous reaction, which holds great potential in point-of-care applications.
Herein, a novel colorimetric biosensing strategy for highly selective and ultrasensitive detection of pathogenic bacteria based on target-triggered EXPAR by the property of polymerase and nicking activity of restriction endonuclease has been reported. And G-quadruplex/hemin DNAzyme was used as an amplifying label to detect the formation of ssDNA–substrate complexes. Our method features with several significant aspects. First, improved sensitivity for pathogenic bacteria quantification is achieved by the utilization of the EXPAR coupled DNAzyme amplification-based strategy. Second, the DNAzyme is used as the catalytic tag, which makes our approach technically label-free, and avoids the need of labeling steps and the addition of additional labile reagents. Third, the colorimetric assay displays a rapid response that results from instant color change and operation of the assay is convenient and low cost. Hence, our biosensor might provide a useful and practical platform for highly sensitive and selective pathogenic bacteria determination and related food safety analysis and environmental monitoring.
2. Materials and methods
2.1. Materials and reagents
Phi29 DNA polymerase, Nb.BbvCI and dNTPs were obtained from New England Biolabs, Inc. (Ipswich, MA, USA). Salmonella typhimurium (CMCC50115), E. coli (KCTC 2571), Bacillus subtilis (KCTC 1028) and Listeria (KCTC 3569) were obtained from Institute of Microbiology Chinese Academy (Beijing, China). Peptone, beef extract powder, bacto-tryptone and bacto-yeast extract were obtained from Qingdao Biological Technology Co. Ltd. (Qingdao, China). Hemin, dimethylsulfoxide (DMSO), H2O2 and ABTS were purchased from Sigma Aldrich Chemical Co. (St. Louis, MO, USA). Other chemicals were of analytical grade and were used without further purification. Ultrapure water obtained from a Millipore filtration system was used throughout. A stock solution of hemin (0.3 μM) was prepared with DMSO and stored at −20 °C in the dark.All oligonucleotides were HPLC-purified and synthesized by Sangon Biotechnology Co. Ltd. (Shanghai, China). The primer had the sequence: 5′ TCTTTTCCTTTTCGGGCATTACT 3′. The aptamer had the sequence: 5′ AGTAATGCCCGGTAGTTATTCAAAGATGAGTAGGAAAAGA 3′. The programmed hairpin (HP) had the sequence: 5′ ![[A with combining low line]](https://www.rsc.org/images/entities/b_char_0041_0332.gif)
![[G with combining low line]](https://www.rsc.org/images/entities/b_char_0047_0332.gif)
![[T with combining low line]](https://www.rsc.org/images/entities/b_char_0054_0332.gif)
![[A with combining low line]](https://www.rsc.org/images/entities/b_char_0041_0332.gif)
![[A with combining low line]](https://www.rsc.org/images/entities/b_char_0041_0332.gif)
![[T with combining low line]](https://www.rsc.org/images/entities/b_char_0054_0332.gif)
![[G with combining low line]](https://www.rsc.org/images/entities/b_char_0047_0332.gif)
![[C with combining low line]](https://www.rsc.org/images/entities/b_char_0043_0332.gif) CCGAAAAGGAAAAGA AAC
CCGAAAAGGAAAAGA AAC![[G with combining low line]](https://www.rsc.org/images/entities/b_i_char_0047_0332.gif)
![[C with combining low line]](https://www.rsc.org/images/entities/b_i_char_0043_0332.gif)
![[A with combining low line]](https://www.rsc.org/images/entities/b_i_char_0041_0332.gif)
![[T with combining low line]](https://www.rsc.org/images/entities/b_i_char_0054_0332.gif)
![[T with combining low line]](https://www.rsc.org/images/entities/b_i_char_0054_0332.gif)
![[A with combining low line]](https://www.rsc.org/images/entities/b_i_char_0041_0332.gif)
![[C with combining low line]](https://www.rsc.org/images/entities/b_i_char_0043_0332.gif)
![[T with combining low line]](https://www.rsc.org/images/entities/b_i_char_0054_0332.gif) 3′, where the short underlined sequences were complementary to each other to make the probe form a hairpin structure. The bold letters were complementary to the whole primer, and the italic bold letters were complementary to the sequence of the helper. The helper had the sequence: 5′ CCCAACCCGCCCTACCCCCTCA ↓GCTTTT AGTAATGCG 3′. The italic letters were completely complementary with the G-quadruplex sequence and the italic bold letters were complementary to the sequence of the HP. The CCTCAGC sequence highlighted in red was the recognition site for Nb.BbvCI, and the arrow indicated the nicking position. The arched probe was performed by adding aptamer and primer in the reaction buffer containing 50 mM Tris–HCl (pH 7.5), 10 mM MgCl2. The mixture was heated to 90 °C for 5 min and slowly cooled to room temperature before use.
3′, where the short underlined sequences were complementary to each other to make the probe form a hairpin structure. The bold letters were complementary to the whole primer, and the italic bold letters were complementary to the sequence of the helper. The helper had the sequence: 5′ CCCAACCCGCCCTACCCCCTCA ↓GCTTTT AGTAATGCG 3′. The italic letters were completely complementary with the G-quadruplex sequence and the italic bold letters were complementary to the sequence of the HP. The CCTCAGC sequence highlighted in red was the recognition site for Nb.BbvCI, and the arrow indicated the nicking position. The arched probe was performed by adding aptamer and primer in the reaction buffer containing 50 mM Tris–HCl (pH 7.5), 10 mM MgCl2. The mixture was heated to 90 °C for 5 min and slowly cooled to room temperature before use.
2.2. Agarose gel electrophoresis analysis
In the gel electrophoresis assay, a sample contains 6 μL of each reaction sample and 1 μL 6× loading buffer. The 4% (w/v) agarose electrophoresis was carried out in 1× TBE (9 mM pH 7.9 Tris–HCl with 9 mM boric acid and 0.2 mM EDTA) at a 140 V constant voltage for 30 min.2.3. Bacterial strains and growth conditions
The bacterial culture was routinely grown at 37 °C in Luria–Bertani medium for 12 h. The cells at logarithmic growth were collected and separated from the medium by centrifugation at 600 rpm and then washed with phosphate-buffered saline (PBS) (10 mM, pH 7.4) twice. The sediment was resuspended in the PBS to obtain a homogeneous cell suspension. The cell number was determined by plate counting method.2.4. Target-triggered isothermally EXPAR
The arched probe was incubated with different concentrations of target for 30 min at 37 °C. Single-stranded primer was released from the arched probe in the process, due to the specific recognization of aptamer toward target. The DNA machine amplification analysis was performed by mixing the above solution, HP solution (100 μM, 2 μL), helper solution (100 μM, 2 μL), dNTPs (10 mM, 2 μL), phi29 DNA polymerase (20 U), Nb.BbvCI endonuclease (20 U), ultrapure water (2 μL), S. typhimurium of a given concentration (0 to 1.0 × 106 cfu mL−1) and incubating the mixture at 37 °C for 60 min.2.5. Absorbance measurements
The resulting reaction mixture was incubated with hemin solution (0.3 μM) at 37 °C for 30 min. Followed by the addition of the freshly prepared ABTS (2 mM) and H2O2 (2 mM), the mixture was transferred to the cuvette to monitor UV-vis absorption spectra. The UV absorbance measurements were performed using a Lambda 35 Spectrometer (Perkin Elmer, USA). The absorption spectra were recorded in the wavelength range from 200 to 700 nm.3. Results and discussion
3.1. Analytical principle for highly sensitive pathogenic bacteria detection
Scheme 1 illustrates the analytical principle of the proposed target-triggered isothermally EXPAR for label-free, homogeneous, and ultrasensitive detection of pathogenic bacteria, and Salmonella typhimurium is selected as a model organisms. The arched probe31,32 contained S. typhimurium aptamer and a universal primer sequence. In the presence of S. typhimurium, it challenged to the arched probe and associated with the aptamer, leading to the release of primer (highlight in blue). Then the HP was unfolded via the released primer binding to its 5′-end, then helper could hybridize to 3′-end of the opened HP and functioned as an initiative strand to initiate an extension reaction and replicated the template part in the HP in the presence of polymerase and dNTPs, in which the primer was displaced. Remarkably, in the system, the HP also functioned as an initiative strand to initiate another extension reaction and replicated the template part in the helper simultaneously. The simultaneous extensions resulted in the displacement of primer and the formation of dsDNA. A new such cycle was initiated when the displaced primer combined to a second HP. As a result, primer was recycled and a new DNA duplex was synthesized at each new cycle (named cycle I). Meanwhile, as a recognition sequence for Nb.BbvCI was generated in the DNA duplex produced from the first cycle, Nb.BbvCI could nick the DNA duplex to release a short ssDNA fragment, which was the aptamer of hemin. Cleavage reaction, replication reaction and strand-displacement reaction were carried out repeatedly, generating millions of short ssDNA products (named cycle II). The released ssDNA sequence could bind hemin with the help of K+ to yield G-quadruplex/hemin DNAzyme, which could catalyze the ABTS–H2O2 reaction to produce strong UV absorption (highlight in pink). As a result, a remarkable UV absorption was achieved owing to the electro-reduction of H2O2 reaction catalyzed by DNAzyme. Remarkably, the use of EXPAR techniques contributed to significantly amplified UV absorption, which improved substantially the sensitivity of the biosensor. Furthermore, our approach does not involve any labeling steps, which is simple, cost-effective and cuts the detection time down to 100 min. | ||
| Scheme 1 Schematic illustration of pathogenic bacteria assay using the target-triggered isothermally EXPAR strategy. | ||
3.2. Gel electrophoresis image of target-triggered isothermally exponential amplification
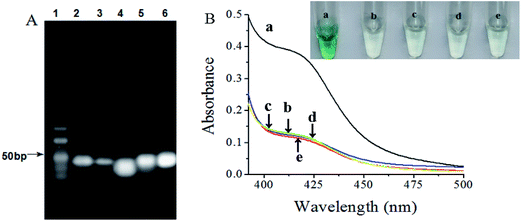
More direct proof of the biosensor mechanism could be acquired through gel electrophoresis analysis, as shown in Fig. 1A. Lane 2 displayed a bright band of our designed arched probe on the image. After incubation with target, the brightness of the band at the position of arched probe was significantly weaker, indicating that the binding of target and arched probe resulted in its structure switch and most of arched probe were consumed (lane 3). As seen from lane 4, the HP and the helper gave a wide band33 because of the same molecular weight almost. The specific binding of the primer to HP triggers its conformational change and the helper binds to the opened HP, the brightness of the band at the position of HP was significantly weaker, a new band was observed on lane 5. The top band demonstrated that the hybridized HP with the primer and helper could prime the polymerization reaction in the presence of phi29, while the band at the bottom indicated that the repeated process of cleavage reaction, replication reaction and strand-displacement reaction could produce numerous short ssDNA fragments (lane 6). These results suggested that only the addition of target could result in the structure switch of HP and trigger the EXPAR with the help of phi29 and Nb.BbvCI.3.3. UV-vis spectra characterization of the EXPAR-based colorimetric biosensor
To verify the feasibility of the designed target–aptamer binding triggered EXPAR strategy for label-free and ultrasensitive UV-vis spectra detection of S. typhimurium in Scheme 1, several important issues should be investigated and evaluated. As shown in Fig. 1B, in the presence of S. typhimurium, a significant UV absorption is generated. This clearly revealed that S. typhimurium triggered enzymatic amplification with the aid of phi29 and Nb.BbvCI, and produced abundant ssDNA fragments binding hemin to form highly active DNAzyme (curve a), in contrast, it was observed that a very weak signal was obtained for the blank sample in the absence of S. typhimurium, suggesting almost no G-quadruplex/hemin complex was formed in the system (curve b). In the absence of phi29 polymerase (curve c) or Nb.BbvCI (curve d) is the same as the background. This signified that the target-triggered amplification was dependent on the phi29-catalyzed extension reaction and Nb.BbvCI-catalyzed strand displacement amplification. Furthermore, a low UV absorption was observed in the presence of phi29, Nb.BbvCI, and nontarget E. coli instead of S. typhimurium, demonstrating that the significantly decreased UV absorption was induced by the specific recognition of S. typhimurium (curve e). On the basis of these results, it strongly indicated that only target bacteria can trigger the EXPAR process by phi29 and Nb.BbvCI to release the DNAzyme sequence for the amplified UV absorption.3.4. Optimization of conditions for the EXPAR-based colorimetric biosensor
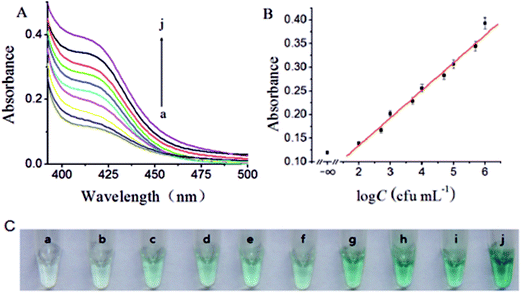
The effect of the incubation time of target S. typhimurium was investigated. As shown in Fig. 2A, the UV absorption increased with the increase of incubation time and reached equilibrium when the incubation time was 30 min. Therefore, the optimal incubation time of target S. typhimurium was 30 min.Fig. 2B depicted the effect of the enzymatic amplification reaction time in the presence of 1.0 × 106 cfu mL−1 S. typhimurium. As anticipated, the UV absorption increased with the increasing of the EXPAR reaction time, and then reached the maximum value at 60 min. When the reaction time varied over the range of 60–100 min, the absorbance and color of the solution remained unchanged, because of the decrease of enzyme activity. So, 60 min was adopted as the optimum enzymatic amplification reaction time and employed for all other investigations.
We further investigated the effect of the concentration of HP on the UV absorption, and the results were shown in Fig. 2C. It was observed that the UV absorption increased with the increase of HP concentration and reached equilibrium when the concentration was 100 μM. Therefore, 100 μM was chosen for the optimal HP concentration in the following experiments.
Considering the important role of hemin in the process of DNAzyme catalytic reaction and the electron transfer between hemin and H2O2, we also investigated the effect of the concentration of hemin on the UV-vis spectra. As shown in Fig. 2D, with increasing amount of hemin in solution, the color change from grayish to green was observed. When the concentration of hemin varied over the range of 0.1–0.5 μM, the absorbance reached equilibrium. Thus, 0.3 μM was selected for the optimal concentration of hemin.
3.5. Analytical performance of the EXPAR-based colorimetric biosensor
Under the optimal experimental conditions, the constructed biosensor was adopted for the determination of a series of different concentrations of S. typhimurium. Fig. 3A depicted typical UV absorption of the biosensor to S. typhimurium of varying concentration. It was observed that the UV absorption increased with the increasing concentration of S. typhimurium. The plot of UV absorption intensity versus the logarithm value of S. typhimurium concentration displayed a good linear relationship in the range from 100 to 1.0 × 106 cfu mL−1 (Fig. 3B), and the limit of detection (LOD) was calculated to be 80 cfu mL−1 in terms of the rule of three times standard deviation over the blank response. Furthermore, the proposed biosensor showed very desirable reproducibility. The relative standard deviations (RSDs) of signal intensity at 410 nm were 2.70%, 3.20%, 2.53%, and 2.44%, respectively, in four repetitive assay of 1.0 × 102, 1.0 × 103, 1.0 × 104 and 1.0 × 105 cfu mL−1 of S. typhimurium. These data signified our biosensor held great potential for the determination of S. typhimurium with high sensitivity and desirable reproducibility (Table 1).| Detection methods | Detection range (cfu mL−1) | Detection limit (cfu mL−1) | Reference |
|---|---|---|---|
| ELISA | 103 to 105 | 103 | 34 |
| SPR | 20 to 102 | 20 | 35 |
| Flow cytometry | 105 to 109 | 105 | 36 |
| QCM | 102 to 104 | 102 | 37 |
| Electrochemical | 6 × 102 to 6 × 106 | 6 × 102 | 38 |
| Chemiluminescence | 5.0 × 102 to 5.0 × 105 | 1.2 × 102 | 39 |
| Colorimetric | 102 to 106 | 80 | This work |
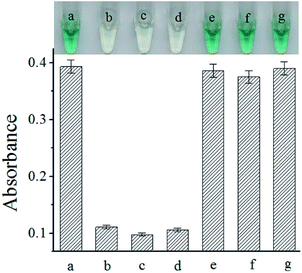
The selectivity of the proposed biosensor for S. typhimurium assay was also examined by using three non-target bacteria. The results were shown in Fig. 4. The presence of even 10-fold excess of the non-target bacteria caused minimal UV-vis absorption spectra. These results implied that our biosensing strategy could be used for the assay of S. typhimurium.
3.6. Real sample analysis
To demonstrate the validity of our biosensor to real samples, the quantitative assay of spiked milk samples was considered. Different concentrations of S. typhimurium were spiked into the milk samples and measured directly without any pretreatments except for dilution with a buffer solution. Table 2 showed the data in the assay for the spiked samples. It was observed that the recovery of our method was in the range of 93.8–102.6%. In addition, the spiked samples were further quantified by a classic plate count method and compared with the results obtained by our method. We found the data obtained by our method were in good agreement with those obtained using plate count method, and the discrepancies between two methods were all smaller than 6.43%. These data clearly demonstrated our biosensor could be applied to complex samples.| Samples | Spiked amount (cfu mL−1) | Our method (cfu mL−1) | Recovery (%) | Plate count method (cfu mL−1) |
|---|---|---|---|---|
| Milk samples | 1.0 × 103 | (1.026 ± 0.046) × 103 | 102.6 | 0.964 × 103 |
| 1.0 × 104 | (0.949 ± 0.039) × 104 | 94.9 | 0.971 × 104 | |
| 1.0 × 105 | (0.938 ± 0.041) × 105 | 93.8 | 0.960 × 105 |
4. Conclusion
A target-triggered isothermally exponential amplification for label-free, homogeneous, and ultrasensitive detection of pathogenic bacteria has been demonstrated in this study. In the presence of phi29 and Nb.BbvCI, target bacteria can trigger the EXPAR to product massive G-quadruplex sequences for production of numerous DNAzyme to achieve signal amplification. Due to the significant signal amplification and the intrinsically high sensitivity of UV-vis spectra detection, as low as 80 cfu mL−1 target bacteria can be detected in 100 min, which is sensitive and efficient far more than most of the pathogenic bacteria assays reported so far.10,40,41 The method presented here was homogeneous, label-free, low cost, simple, and highly sensitive. Thus, it opens a promising avenue to develop the EXPAR-based ultrasensitive UV-vis spectra method for pathogenic bacteria detection.Acknowledgements
This work was supported by NSFC (31171700, 31471644, and 21405060), National High Technology Research and Development Program of China (2012AA101604), Shandong Province Natural Science Funds for Distinguished Young Scholars (JQ201410), and Promotive Research Fund for Excellent Young and Middle-aged Scientists of Shandong Province (BS2014SW033).References
- C. Y. Wen, J. Hu, Z. L. Zhang, Z. Q. Tian, G. P. Ou, Y. L. Liao, Y. Li, M. Xie, Z. Y. Sun and D. W. Pang, Anal. Chem., 2013, 85, 1223–1230 CrossRef CAS PubMed.
- Y. N. Guo, Y. Wang, S. Liu, J. H. Yu, H. Z. Wang, M. Cui and J. D. Huang, Analyst, 2015, 140, 551–559 RSC.
- P. Kannan, H. Y. Yong, L. Reiman, C. Cleaver, P. Patel and A. A. Bhagwat, Foodborne Pathog. Dis., 2010, 7, 1551–1558 CrossRef CAS PubMed.
- J. Chenau, F. Fenaille, S. Simon, S. Filali, H. Volland, C. Junot, E. Carniel and F. Becher, Anal. Chem., 2014, 86, 6144–6152 CrossRef CAS PubMed.
- S. C. Donhauser, R. Niessner and M. Seidel, Anal. Chem., 2011, 83, 3153–3160 CrossRef CAS PubMed.
- J. Wang, M. J. Morton, C. T. Elliott, N. Karoonuthaisiri, L. Segatori and S. L. Biswal, Anal. Chem., 2014, 86, 1671–1678 CrossRef CAS PubMed.
- L. Wu, T. Luan, X. Yang, S. Wang, Y. Zheng, T. Huang, S. Zhu and X. Yan, Anal. Chem., 2014, 86, 907–912 CrossRef CAS PubMed.
- M. A. Vorobjeva, V. V. Krasitskaya, A. A. Fokina, V. V. Timoshenko, G. A. Nevinsky, A. G. Venyaminova and L. A. Frank, Anal. Chem., 2014, 86, 2590–2594 CrossRef CAS PubMed.
- D. Kovář, Z. Farka and P. Skládal, Anal. Chem., 2014, 86, 8680–8686 CrossRef PubMed.
- H. Shiigi, T. Kinoshita, M. Fukuda, D. Q. Le, T. Nishino and T. Nagaoka, Anal. Chem., 2015, 87, 4042–4046 CrossRef CAS PubMed.
- T. Özlem, H. B. İsmail, T. Erhan and T. Uğur, Biosens. Bioelectron., 2012, 37, 53–60 CrossRef PubMed.
- J. O. Seung, H. P. Byung, H. J. Jae, C. Goro, C. L. Doh, H. K. Do and S. S. Tae, Biosens. Bioelectron., 2016, 75, 293–300 CrossRef PubMed.
- W. S. Chan, B. S. F. Tang, V. B. Maureen, C. Chow and P. H. M. Leung, Biosens. Bioelectron., 2014, 53, 105–111 CrossRef CAS PubMed.
- Y. S. Wang, D. H. Zhang, W. Liu, X. Zhang, S. X. Yu, T. Liu, W. T. Zhang, W. X. Zhu and J. L. Wang, Biosens. Bioelectron., 2014, 55, 242–248 CrossRef CAS PubMed.
- C. Tuerk and L. Gold, Science, 1990, 249, 505–510 CrossRef CAS PubMed.
- R. R. Breaker and G. F. Joyce, Chem. Biol., 1994, 1, 223–229 CrossRef CAS PubMed.
- J. Liu, Z. Cao and Y. Lu, Chem. Rev., 2009, 109, 1948–1998 CrossRef CAS PubMed.
- D. S. Wilson and J. W. Szostak, Annu. Rev. Biochem., 1999, 68, 611–647 CrossRef CAS PubMed.
- N. Duan, S. J. Wu, X. J. Chen, Y. K. Huang, Y. Xia, X. Y. Ma and Z. P. Wang, J. Agric. Food Chem., 2013, 61, 3229–3234 CrossRef CAS PubMed.
- A. D. Ellington and J. W. Szostak, Nature, 1990, 346, 818–822 CrossRef CAS PubMed.
- S. L. Zeng, H. K. Huang, Y. Huang, X. Q. Liu, J. Qin, S. L. Zhao, Z. F. Chen and L. Hong, RSC Adv., 2015, 5, 43105–43109 RSC.
- W. Yun, J. L. Jiang, D. Z. Cai, X. F. Wang, G. Sang, J. S. Liao, T. C. Lu and K. P. Yan, RSC Adv., 2016, 6, 3960–3966 RSC.
- Y. J. Guo, L. Deng, J. Li, S. J. Guo, E. K. Wang and S. J. Dong, ACS Nano, 2011, 5, 1282–1290 CrossRef CAS PubMed.
- M. Deng, D. Zhang, Y. Zhou and X. Zhou, J. Am. Chem. Soc., 2008, 130, 13095–13102 CrossRef CAS PubMed.
- H. X. Jia, Z. B. Wang, C. L. Wang, L. J. Chang and Z. P. Li, RSC Adv., 2014, 4, 9439–9444 RSC.
- F. Ma, Y. Yang and C. Y. Zhang, Anal. Chem., 2014, 86, 6006–6011 CrossRef CAS PubMed.
- X. Q. Zhou, Y. Q. Liang, Y. Z. Xu, X. L. Lin, J. Chen, Y. J. Ma, L. Zhang, D. P. Chen, F. Y. Song, Z. Dai and X. Y. Zou, Biosens. Bioelectron., 2016, 80, 378–384 CrossRef CAS PubMed.
- P. S. Song, Y. Xiang, H. Xing, Z. J. Zhou, A. J. Tong and Y. Lu, Biosens. Bioelectron., 2012, 84, 2916–2922 CAS.
- H. Z. Wang, Y. Wang, S. Liu, J. H. Yu, W. Xu, Y. N. Guo and J. D. Huang, Chem. Commun., 2015, 51, 8337 Search PubMed.
- R. Freeman, X. Liu and I. Willner, J. Am. Chem. Soc., 2011, 133, 11597–11604 CrossRef CAS PubMed.
- F. L. Gao, J. P. Lei and H. X. Ju, Chem. Commun., 2013, 49, 4006 RSC.
- Y. Y. Yu, Z. Chen, L. Shi, F. Yang, J. Pan, B. Zhang and D. Sun, Anal. Chem., 2014, 86, 8200–8205 CrossRef CAS PubMed.
- C. Shi, X. T. Shen, S. Y. Niu and C. P. Ma, J. Am. Chem. Soc., 2015, 137, 13804–13806 CrossRef CAS PubMed.
- E. Nazemi, S. Aithal, W. M. Hassen, E. H. Frost and J. J. Dubowski, Sens. Actuators, B, 2015, 207, 556–562 CrossRef CAS.
- E. Bulard, A. B. Spinelli, P. Chaud, A. Roget, R. Calemczuk, S. Fort and T. Livache, Anal. Chem., 2015, 87, 1804–1811 CrossRef CAS PubMed.
- C. Piñero-Lambea, G. Bodelón, R. Fernández-Periáñez, A. M. Cuesta, L. Vallina and L. Fernández, ACS Synth. Biol., 2015, 4, 463–473 CrossRef PubMed.
- V. C. Ozalp, G. Bayramoglu, Z. Erdem and M. Y. Arica, Anal. Chim. Acta, 2015, 853, 533–540 CrossRef CAS PubMed.
- X. M. Wang, P. Zhu, F. W. Pi, H. Jiang, J. D. Shao, Y. Z. Zhang and X. L. Sun, Biosens. Bioelectron., 2016, 81, 349–357 CrossRef CAS PubMed.
- Z. Li, H. Yang, L. Sun, H. Qi, Q. Gao and C. Zhang, Sens. Actuators, B, 2015, 210, 468–474 CrossRef CAS.
- T. Raweewan, J. Kulachart, T. Pramuan and P. Duangporn, Anal. Chem., 2013, 85, 5996–6002 CrossRef PubMed.
- B. Emilie, B. S. Aurelie, C. Patricia, R. Andre, C. Roberto, F. Sebastien and L. Thierry, Anal. Chem., 2015, 87, 1804–1811 CrossRef PubMed.
| This journal is © The Royal Society of Chemistry 2016 |