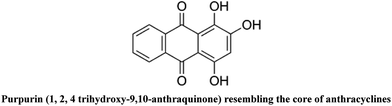
Synthesis, crystal structure from PXRD of a MnII(purp)2 complex, interaction with DNA at different temperatures and pH and lack of stimulated ROS formation by the complex†
Bitapi
Mandal
a,
Soumen
Singha
b,
Sanjay Kumar
Dey
b,
Swagata
Mazumdar
c,
Tapan Kumar
Mondal
a,
Parimal
Karmakar
c,
Sanjay
Kumar
b and
Saurabh
Das
*a
aDepartment of Chemistry (Inorganic Section), Jadavpur University, Kolkata-700032, India. E-mail: sdas@chemistry.jdvu.ac.in; Fax: +91 33 24146223; Tel: +91 89 02087756
bDepartment of Physics, Jadavpur University, Kolkata-700032, India
cDepartment of Life Science and Biotechnology, Jadavpur University, Kolkata-700032, India
First published on 18th May 2016
Abstract
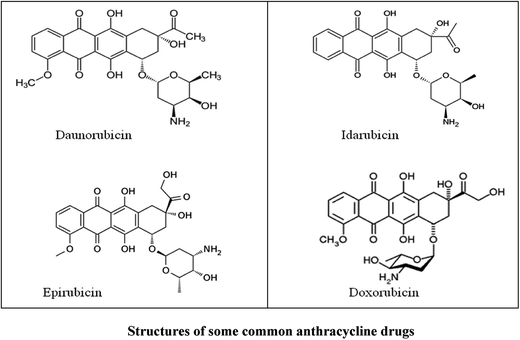
The formation of reactive oxygen species (ROS) by anthracycline anticancer drugs is essential for their antitumor activity but they also make these drugs cardiotoxic. When complexed with metal ions there is a decrease in ROS formation and therefore in cardiotoxicity. Interestingly, in spite of producing fewer ROS, some of the complexes are effective antitumor agents, often better than the parent anthracycline. Purpurin (LH3), a hydroxy-9,10-anthraquinone, resembles doxorubicin at the core. An MnII complex of LH3 [MnII(LH2)2] was synthesized to see the extent to which the complex resembles metal–anthracyclines with regard to structure and function. The crystal structure was determined by Rietveld refinement of PXRD data using an appropriate structural model developed on the basis of spectroscopic information. This is only the second report on the crystal structure of a hydroxy-9,10-anthraquinone with a 3d-transition metal ion. Bond lengths and bond angles were obtained by structural refinement. The structure is supported by DFT calculations. DNA binding of the complex is slightly better than for purpurin but more importantly unlike purpurin, the binding constant values remained constant even with an increase in the pH of the medium. The NADH dehydrogenase assay and the DCFDA-ROS generation assay showed that generation of superoxide in the former and ROS in general in the latter were significantly less for the complex than for purpurin. Even with decreased ROS formation, the complex is able to maintain the biological activity of purpurin.
1. Introduction
Hydroxy-9,10-anthraquinones present at the core of an anthracycline antibiotic are crucial for antitumor activity for more than one reason.1–5 Besides being responsible for generating reactive oxygen species (ROS) essential for cytotoxic activity, by being planar it allows the molecule to intercalate into DNA leading to disruption in replication and transcription.6–8 Studies reveal it is also responsible for inhibiting the activity of topoisomerase enzymes.5,9,10 A number of studies on various hydroxy-9,10-anthraquinones were done to realize the difference in activity of this unit when either present alone or as part of an anthracycline drug dealing in physicochemical, electrochemical, biophysical, biochemical and cell line experiments.10–15 Findings indicate while for the first two aspects there was not much difference whether the hydroxy-9,10-anthraquinone was on its own or within an anthracycline antibiotic, for the remainder the difference was significant. In most cases, anthracyclines were more effective than hydroxy-9,10-anthraquinones present on their own.5,10,11,14Probably the other units present in anthracyclines assist hydroxy-9,10-anthraquinones in discharging its duties effectively.14–16 A study with carminic acid, having a sugar unit directly bound to a hydroxy-9,10-anthraquinone showed significant improvement in biophysical interactions and in experiments on cancer cells. This supports the general belief sugars play an important role in anticancer activity although a direct scientific justification is scarce. Sugar(s), for example, is(are) believed to assist in drug binding to DNA and in recognition of cancer cells for which this moiety is often incorporated in a molecule during drug design.16–18
A good number of metal complexes of anthracyclines have also been prepared.19–23 The reason behind their formation is well known.19–23 What is important is to show with suitably designed experiments that metal complexes (either of anthracyclines or of hydroxy-9,10-anthraquinones) are actually more effective cytotoxic agents than compounds from which they are prepared. While there are examples showing complex formation besides decreasing cardiotoxic side effects compromise on cytotoxic action, several studies have also shown some of the complexes are even better anticancer agents.22,23 In fact many metal complexes are being used as drugs today and the metal ion present in these complexes has a substantial role in this regard.24–27 For studies on cancer, Cu(II) complexes are usually very effective, the reason attributed to the affinity of cancer cells for Cu(II).28–33 Inherent toxicity of the metal ion is another important aspect that requires serious consideration. Manganese is an important element in this regard.34 MnII for example, is bio-friendly when present in low concentration.35–39 Mn-containing metallo-proteins are known for functions that range from photosynthesis to controlling oxidative stress.40 Bacterial protein-based uptake systems for manganese are also known.34 Some studies reveal how these proteins contribute to manganese homeostasis.41,42 Small molecules are reported to be exported by cells to bind Mn(II) with high affinity facilitating cellular uptake.43 Numerous micro-organisms catalyse the oxidation of Mn(II) but whether it serves any biological role is still not known with certainty.44 Enzyme-catalysed oxidation occurs through sequential one-electron transfer steps, generating Mn(III) intermediates.45–47 Thus for this study, an Mn(II) complex of 1,2,4-trihydroxy-9,10-anthraquinone (purpurin) was prepared and some of its parameters compared with known metal complexes of anthracyclines or hydroxy-9,10-anthraquinones.12,14,15,20–23
The novel aspects of this study are the structure of the complex obtained from powder X-ray diffraction data since single crystals were not obtained, determination of thermodynamic parameters pertaining to interaction of purpurin and its complex with calf thymus DNA and lack of stimulated ROS formation by the complex. The structure is novel since the same is rare for metal complexes of hydroxy-9,10-anthraquinones, with only one report so far from single crystal X-ray diffraction.48 This is only the second structure of a metal complex of a hydroxy-9,10-anthraquinone with a 3d transition metal ion.14 The importance of the thermodynamic study is that it helps to explain trends observed in binding of a hydroxy-9,10-anthraquinone and its metal complex with DNA that puts our results in proper perspective with that of established anthracyclines and their metal complexes. Decrease in ROS generation by the complex is important for it indicates that the complex is likely to be less cardiotoxic.
2. Experimental
2.1 Materials and instruments used
Purpurin (1,2,4-trihydroxy-9,10-anthraquinone) was purchased from Sigma-Aldrich, USA and purified by recrystallization from ethanol. Manganese(II) acetate-4-hydrate, sodium bicarbonate, sodium chloride, ethanol and dimethyl sulphoxide were purchased from E. Merck, India. Tris buffer [i.e. tris(hydroxy methyl)amino methane] was obtained from Spectro Chem. (India) Pvt. Ltd. Calf thymus (c t) DNA was purchased from Sisco Research Laboratory, India. Triple distilled water was prepared in the laboratory by distillation of double distilled water. The tris buffer solution was prepared in triple distilled water. Stock solution of purpurin [10−3 M] was prepared in ethanol while manganese acetate [10−3 M] was prepared in water.![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2 in aqueous solution at pH ∼ 6.5 after adjusting appropriately with a solution of sodium bicarbonate. The mixture was stirred for 4 hours at 40 °C and allowed to stand overnight. A reddish brown solid settled to the bottom which was filtered and washed with 10% hot ethanol–water mixture. The product was dried in a vacuum desiccator. Distinct changes in the UV-vis spectrum (Fig. S2, ESI†), recorded on a JASCO V-630 spectrophotometer and IR spectrum (Fig. S3 and S4, ESI†), recorded on Perkin Elmer Spectrum Two FTIR spectrophotometer suggest complex formation. Mass spectrum (Fig. S5, ESI†) recorded on Micromass Q-Tofmicro™, Waters Corporation and TGA (Fig. S6, ESI†) on Mettler Toledo TGA/SDTA 851e also suggest complex formation. Elemental analysis was carried out on Perkin Elmer 2400 Series-II CHN analyzer. Magnetic susceptibility measurements of powdered samples at room temperature (298 K) were recorded by the Gouy method using Magway MSB MK1, Sherwood Scientific Ltd.
2 in aqueous solution at pH ∼ 6.5 after adjusting appropriately with a solution of sodium bicarbonate. The mixture was stirred for 4 hours at 40 °C and allowed to stand overnight. A reddish brown solid settled to the bottom which was filtered and washed with 10% hot ethanol–water mixture. The product was dried in a vacuum desiccator. Distinct changes in the UV-vis spectrum (Fig. S2, ESI†), recorded on a JASCO V-630 spectrophotometer and IR spectrum (Fig. S3 and S4, ESI†), recorded on Perkin Elmer Spectrum Two FTIR spectrophotometer suggest complex formation. Mass spectrum (Fig. S5, ESI†) recorded on Micromass Q-Tofmicro™, Waters Corporation and TGA (Fig. S6, ESI†) on Mettler Toledo TGA/SDTA 851e also suggest complex formation. Elemental analysis was carried out on Perkin Elmer 2400 Series-II CHN analyzer. Magnetic susceptibility measurements of powdered samples at room temperature (298 K) were recorded by the Gouy method using Magway MSB MK1, Sherwood Scientific Ltd.
Yield: ∼70%. Anal calc. (%) for MnC28H14O10: C, 59.47%; H, 2.48%. Experimentally found: C, 59.85%; H, 2.52%, UV-vis spectra: λmax at 513 nm. MS (m/z): 565.88 [M]+.
DNA preparation. Calf thymus DNA was dissolved in triple distilled water and allowed to stand for 24 h. A molar extinction coefficient of 6600 M−1 cm−1 at 260 nm was used for calculating concentration. Absorbance was also measured at 280 nm. The ratio of A260/A280 was found to be in between 1.8 and 1.9 suggesting the DNA was sufficiently free of protein and ready for use. Quality of c t DNA was also verified from the CD spectra at 260 nm using a CD spectropolarimeter, J815, JASCO.
2.2 Methods
The space group was obtained from statistical analysis of the powder patterns with the help of the find space module of the EXPO 2009 software package.49 Statistical analysis shows that the most probable space group is P21. For this unit cell and space group, full pattern decompositions were performed using Le Bail method giving good fit between calculated and experimental powder X-ray patterns. The structure solutions from PXRD data were carried out using the simulated annealing technique (parallel tempering mode) as implemented in the program FOX,50 a Monte Carlo based software. The initial molecular structure was first drawn using ACD/ChemSketch and the geometry of the structure was optimized with the help of MOPAC 2009 to arrive at the reference structural model for FOX.51 After a large number of successful cycles we obtained the atomic coordinates from FOX. These were used as the input of the starting model for Rietveld structure refinement using the program GSAS52 with an EXPGUI53 interface. Peak shapes were described as pseudo-Voigt functions and the backgrounds were fitted by the shifted Chebyshev function of first kind with 36 points regularly distributed over the entire 2θ range.
Initially the lattice parameters, profile parameters, and background coefficients were refined. After applying the soft constraints on bond lengths and bond angles and planar restraints on the aromatic rings, the positions of the atoms were refined. A fixed isotropic displacement parameter of 0.04 Å2 for non-hydrogen atoms and 0.06 Å2 for hydrogen atoms was maintained. At the final stage of the refinement, preferred orientation correction was applied using the generalized spherical harmonic model. The final Rietveld plot is shown in Fig. 1.
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 915 µM till saturation was reached. Titration was performed at four different pH at 303 K. Results were analyzed using eqn (S1)–(S5) (ESI†).
915 µM till saturation was reached. Titration was performed at four different pH at 303 K. Results were analyzed using eqn (S1)–(S5) (ESI†).
ln![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Kapp = (−ΔH0/RT) + (ΔS0/R) Kapp = (−ΔH0/RT) + (ΔS0/R) | (1) |
ΔG0 = −RT![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) ln ln![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Kapp Kapp | (2) |
R and T are the universal gas constant and absolute temperature respectively. The apparent binding constant [Kapp = Kd−1] was determined at each temperature using eqn (S1)–(S5) (ESI†). ΔH0 and ΔS0 were determined from the slope and intercept of eqn (1) and ΔG0 from eqn (2).54 We wish to mention here that we did not have access to an Isothermal Titration Calorimeter (ITC) that prevented us from determining the thermodynamic parameters that are model independent.
3. Results and discussion
3.1 Description of the X-ray crystal structure of [C28H14O10Mn]
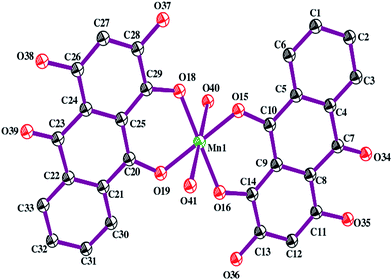
Structural analysis using the PXRD data indicates the MnII complex crystallizes in monoclinic P21 space group and that the asymmetric unit of the complex contains one MnII ion, two mono-anionic LH2− units and two coordinated water molecules. The final crystallographic data and Rietveld refinement parameters are depicted in Table 1 while the asymmetric unit of the complex is depicted in Fig. 2.| Formula | C28H14MnO10·2(H2O) |
| Formula weight | 601.36 |
| Crystal system | Monoclinic |
| Space group | P 21 |
| a/Å | 15.748(12) |
| b/Å | 12.735(9) |
| c/Å | 7.2909(17) |
| α/° | 90.00 |
| β/° | 106.47(8) |
| γ/° | 90.00 |
| V/Å3 | 1402.2(16) |
| Z | 2 |
| ρ calc./g cm−3 | 1.424 |
| Temperature/K | 293 |
| Radiation/Å | 1.54184 |
| 2θ range/° | 5–80 |
| R wp | 0.0483 |
| R p | 0.0328 |
| χ | 2.91 |
Each MnII centre shows a distorted octahedral geometry with coordination number six. The coordination environment of MnII is fulfilled by two deprotonated phenolic –OH groups (O16 and O18) of two different LH3 units and two carbonyl oxygen atoms (O15 and O19). Two water molecules (O40 and O41) occupy the fifth and sixth coordination sites. Selected bond lengths and bond angles are depicted in Table 2. In the absence of a single crystal, we were forced to arrive at the structure from X-ray powder diffraction data. For this reason we also performed DFT calculations to arrive at an optimized structure of the complex (ESI†). Of the three structures obtained in the process, i.e. a high spin octahedral form, a low spin octahedral form and a tetrahedral form, the high spin octahedral geometry having minimum energy was considered the most stable structure for the complex (ESI†). Magnetic susceptibility measurements support DFT calculations since the magnetic moment was found to be 5.78 B.M. Hence all evidence leads us to the octahedral structure as obtained from PXRD.
| Bonds | Bond lengths (Å) |
|---|---|
| Mn1–O15 | 1.8503(15) |
| Mn1–O16 | 1.8733(12) |
| Mn1–O18 | 1.8765(12) |
| Mn1–O19 | 1.8532(15) |
| Mn1–O40 | 2.3679(10) |
| Mn1–O41 | 2.3706(10) |
| Bonds | Bond angles (°) |
|---|---|
| O15–Mn1–O16 | 100.004(32) |
| O15–Mn1–O18 | 79.704(31) |
| O15–Mn1–O19 | 180.0 |
| O16–Mn1–O18 | 180.0 |
| O16–Mn1–O19 | 80.198(32) |
| O18–Mn1–O19 | 100.092(32) |
3.2 Determination of stability constant of MnII(LH2)2 in solution
A spectrophotometric titration was performed taking purpurin and MnII in the ratio 2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 in the pH range 2.08 to 10.0 (Fig. 3). The change in absorbance at 513 nm for a change in pH was fitted to eqn (3) that helped to determine the stability constant of the complex in solution (details in ESI†).
1 in the pH range 2.08 to 10.0 (Fig. 3). The change in absorbance at 513 nm for a change in pH was fitted to eqn (3) that helped to determine the stability constant of the complex in solution (details in ESI†).| Aobs = A1/(1 + 10pH−pK) + A2/(1 + 10pK−pH) | (3) |
 | ||
Fig. 3 Spectra obtained following the gradual addition of NaOH to a mixture of MnII and purpurin that was taken in the ratio 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2. pH of each solution is shown in the inset of the figure. 2. pH of each solution is shown in the inset of the figure. | ||
A 1 and A2 are the absorbance of LH3 and LH2− respectively, in presence of MnII. A proton from purpurin (from the phenolic –OH at C1) having pKa 4.90 ± 0.11 (Fig. S1d, ESI†) dissociated in the pH range 2.0 to 7.0. In the presence of MnII, the dissociation of –OH at C2 of purpurin occurs at a pH beyond 8.75.55,61 Hence, in the physiological pH range the complex does not dissociate to form anionic species. This has a biological significance pertaining to complex formation (discussed later) and also explains why it is less soluble than purpurin in aqueous medium. The stability constant of MnII(LH2)2 is 1.1 × 1018 (ESI†).
3.3 Interaction of the compounds with calf thymus DNA
| Experimental pH | K app × 10−4 (M−1) from double-reciprocal plot | K app × 10−4 (M−1) from non-linear curve fitting | n b from mole ratio plot | K′ × 10−5 (M−1) = Kapp × nb | K′ × 10−5 (M−1) Scatchard plot | n b Scatchard plot |
|---|---|---|---|---|---|---|
| 7.19 | 0.60 ± 0.05 | 0.76 ± 0.03 | 14.0 | 0.95 | 1.06 ± 0.10 | 12.0 |
| 7.44 | 1.10 ± 0.10 | 1.65 ± 0.07 | 12.0 | 1.65 | 1.14 ± 0.13 | 12.0 |
| 8.18 | 2.52 ± 0.37 | 4.63 ± 0.42 | 7.0 | 2.50 | 2.53 ± 0.28 | 8.0 |
| 8.25 | 1.74 ± 0.15 | 3.02 ± 0.18 | 10.0 | 2.38 | 2.54 ± 0.54 | 8.0 |
The ratio of the change in absorbance ΔA to ΔAmax was plotted against concentration of DNA and fitted to eqn (S4).† This provided another set of values for Kd from where Kapp was evaluated (Fig. 5 and Table 3). Fig. S8 (ESI†), indicates the site size of interaction (nb) of the complex with c t DNA at different pH.
The titrimetric data was also fitted to a modified form of the Scatchard equation (eqn (S5), ESI†) that provides values for the overall binding constant (K*) (Fig. 6).62 The same is also obtained by multiplying Kapp determined earlier with nb. Values for binding constants obtained using different equations were compared (Table 3). A comparison of the site size of interaction (nb) of purpurin and its MnII complex with c t DNA (Table 3) reveals the value obtained for the complex at each pH is approximately double that of purpurin binding to c t DNA supporting the fact two molecules of purpurin are bound to the metal ion in the complex.12
 | ||
| Fig. 6 Scatchard plot showing the interaction of MnII(LH2)2 with c t DNA. [MnII(LH2)2] = 50 µM, [NaCl] = 120 mM; [Tris–HCl] = 70 mM was used to maintain four different pH. T = 298 K. | ||
Hence, if the mode of binding of a particular molecule and its complex be same then stoichiometry of complex formation can also be determined from a comparison of their interaction with DNA. With the help of several studies we found this to hold good quite appreciably, particularly if the structure of the complex is planar.12,15,63 Experimental observations further suggest that the complex binds DNA better than purpurin. The more interesting aspect is that unlike purpurin, overall binding constant values of the complex did not decrease with an increase in the pH of the medium. Thus the results suggest an increased affinity of the complex for DNA with an added advantage that they remain more or less constant over the physiological pH range (Table 3).
It was earlier observed in case of purpurin and another hydroxy-9,10-anthraquinone that as the pH of the medium increased the wavelength at which the interaction was followed underwent a gradual red shift (by 20 nm for a change in pH from 7.2 to 8.1).13,14 However, in case of the MnII complex such a thing did not happen and the wavelength at which the titration was followed remained almost constant changing only by 4 nm (from 510 nm at pH = 7.19 to 514 nm at pH = 8.25).14 This aspect supports the fact the complex unlike purpurin does not generate anionic species in solution at physiological pH (mentioned earlier with regard to physicochemical experiments), a big advantage keeping in mind the fact generation of anions by compounds has a negative influence on their interaction with DNA.13,14,17 From the above mentioned facts, it may be said that the MnII complex seems to be a potential candidate for development as a good DNA binding or DNA damaging agent that might be used to treat cancer patients given the already established anticancer activity of hydroxy-9,10-anthraquinone unit in anthracyclines. The advantage in case of the complex would be that it would have a reasonably equal affinity for DNA even if there are fluctuations of pH in body fluids, a common phenomenon in cancer patients.
Binding constant and site size of interaction were evaluated using eqn (S3) and (S4)† for both compounds at different temperatures (Fig. 8 and Table 4). Analysis of the data provides a trend for the tendency of the compounds to bind to c t DNA following an increase in temperature.
 | ||
| Fig. 8 Binding isotherms for the interaction of purpurin and MnII(purp)2 complex with c t DNA at different temperatures where ΔA/ΔAmax was plotted against concentration of c t DNA. Dark line is the fitted data obeying eqn (S4).† Inset: plots for the normalized increase in absorbance as a function of mole-ratio of c t DNA to the compounds at different temperatures. [LH3] = 50 µM; [MnII(LH2)2] = 32 µM; [NaCl] = 120 mM; [Tris buffer] = 70 mM, pH ∼ 7.4; T = 298 K. | ||
| Monitoring technique | Temp (in °C) | Apparent binding constants Kapp × 10−3 (M−1) | Site size (nb) from mole ratio plot | Overall binding constant K* × 10−4 (M−1) [K* = Kapp × nb] | Overall binding constant K′ × 10−4 (M−1) (from Scatchard plot) | Site size (nb) Scatchard plot | ||
|---|---|---|---|---|---|---|---|---|
| From double reciprocal plot | From non linear – fit | |||||||
| UV-vis | MnII(LH2)2 | 20 | 3.48 ± 0.12 | 2.82 ± 0.06 | 13.0 | 4.09 | 3.53 ± 0.42 | 10.0 |
| 25 | 3.30 ± 0.14 | 3.20 ± 0.13 | 12.0 | 3.90 | 5.59 ± 0.12 | 12.6 | ||
| 30 | 6.10 ± 0.16 | 7.15 ± 0.30 | 10.0 | 6.63 | 7.04 ± 1.06 | 10.2 | ||
| 35 | 8.87 ± 0.17 | 9.21 ± 0.23 | 14.0 | 12.66 | 10.29 ± 1.07 | 11.6 | ||
| LH3 | 20 | 1.49 ± 0.05 | 1.50 ± 0.04 | 7.0 | 1.05 | 2.02 ± 0.55 | 6.4 | |
| 25 | 5.96 ± 0.35 | 4.90 ± 0.26 | 5.8 | 3.02 | 5.38 ± 0.23 | 5.2 | ||
| 30 | 6.66 ± 0.26 | 6.48 ± 0.32 | 8.0 | 5.26 | 6.62 ± 1.55 | 8.5 | ||
| 35 | 13.0 ± 1.56 | 11.2 ± 1.11 | 7.5 | 9.08 | 8.60 ± 1.11 | 6.0 | ||
Thermodynamic parameters for the interaction were evaluated (Table 5). Fig. 9 is a representative van't Hoff plot for purpurin and its MnII complex considering values for apparent binding constants as obtained from their respective titration with c t DNA at four different temperatures (eqn (S2)–(S4),† and Table 4). The plot clearly indicates binding affinity increases with increase in temperature. A comparison of the interaction of purpurin and the complex with c t DNA further revealed binding free energy is actually comparable, this being the case when the complex is actually almost twice as large as compared to purpurin. Binding of both compounds to DNA is characterized by a positive enthalpy change implying that the association is entropy driven. Increase in enthalpy of the process is compensated by an increase in entropy. The positive change in entropy leading to an entropy driven nature of the association could be attributed to the release of minor groove bound water from the hydration region as well as the release of counter ions from the bound surface of both the interacting partners.64–67 On the other hand, the positive change could arise from an energetically unfavourable distortion of the DNA backbone at the binding site under going a B to A type transition characterized by the widening of the minor groove to accommodate the complex.64–67 Besides in such entropically driven DNA binding, hydrophobic interactions also play a major role.68,69
| Compounds | ΔG0 (kcal mol−1) | ΔH0 (kcal mol−1) | ΔS0 (e.u) |
|---|---|---|---|
| LH3 | −4.858 | 23.451 | 95.00 |
| MnII(LH2)2 | −4.919 | 13.473 | 61.72 |
 | ||
| Fig. 9 Representative van't Hoff plot obtained for the interaction of purpurin and its MnII complex to c t DNA in 70 mM Tris–HCl buffer, pH ∼ 7.4. | ||
Structural and thermodynamic studies are both essential for understanding the molecular basis of the binding process.70 The linear van't Hoff plots indicate the small value of heat capacity changes.71 These may be attributed to the absence of any major conformational alterations involving the DNA. In trying to develop a correlation between thermodynamic and structural data, it is essential to consider that enthalpy-entropy compensation leads to the observed free energy change. The evaluated thermodynamic parameters are able to explain the binding parameters obtained for the two compounds when they separately interact with c t DNA.
3.4 Generation of ROS by the compounds
Since superoxide radical anions are formed when semiquinones interact with molecular oxygen,14,15,19,72–80 in case of the complex, with one of the carbonyls of each purpurin involved in coordinating the metal ion, semiquinone formation decreases further. Therefore, in case of the complex only one carbonyl on each purpurin is free to form semiquinone. MnII present in the complex transfers an electron from the semiquinone to the metal centre thus further decreasing the possibility of its interaction with molecular oxygen.14,15,19,81 Such decreased semiquinone or O2−˙ formation due to the complex is a blessing since the complex is likely to be less cardiotoxic as reported earlier for different metal-anthracyclines.19,21–23 In order to see whether the complex is able to maintain the efficacy on cancer cells we performed a study of purpurin and MnII(purp)2 on ALL MOLT-4 cells followed by the MTT assay (Fig. S9, ESI†). It was seen that the performance of MnII(purp)2 was comparable to that of purpurin.
4. Conclusions
A mononuclear complex of MnII with purpurin [MnII(LH2)2] was characterized by physicochemical and spectroscopic techniques. The structure of the complex was solved from powder XRD. Physicochemical attributes of [MnII(LH2)2] were similar to metal complexes of some standard anthracycline drugs reported earlier. Interaction with c t DNA suggest while in case of purpurin binding constant values decreased with increase in pH, for the complex the values were not only higher but they also remained constant over a wide range of pH. This fact is significant, as the complex may be used on cancer patients for whom fluctuation of pH of body fluids is a common phenomenon. Evaluation of thermodynamic parameters on the interaction of purpurin and its MnII complex with DNA justifies observed trends in binding parameters of the complex with respect to purpurin. The NADH dehydrogenase assay on O2−˙ generation and the DCFDA assay on ROS formation reveals inspite of lower values in case of MnII(LH2)2, the complex was found to be slightly more potent in killing ALL MOLT-4 cells than purpurin. This goes to indicate MnII(LH2)2, even with decreased ROS formation shows almost comparable anticancer activity as purpurin. Moreover since reduced ROS formation is often correlated to the molecule being less cardiotoxic, complex formation of purpurin with MnII may be viewed in this perspective as well. Hence, there exists a good possibility that MnII(LH2)2 or MnII(purp)2 might be active on cancer cells and yet have less cardiotoxic side effects. It could be developed into a promising, useful and less costly alternative.Conflict of interest
The authors declare no competing financial interests.Abbreviations
| LH3 | Purpurin or 1,2,4-trihydroxy-9,10-anthraquinone |
| MnII(LH2)2 or MnII(purp)2 | The MnII complex of purpurin or LH3 |
| ROS | Reactive oxygen species |
| DOX | Doxorubicin |
| DNR | Daunorubicin |
| c t DNA | Calf thymus DNA |
| SOD | Superoxide dismutase |
Acknowledgements
Financial support from DST, Govt. of West Bengal [794(Sanc.)1(10) ST/P/S&T/9G-23/2013] in the form of a research project to S. D. is gratefully acknowledged. BM expresses his gratitude to the U G C, New Delhi for a Junior & Senior Research Fellowship. SD, BM, SS, SKD and SK are grateful to DST, Government of India, New Delhi for the special grant provided to the Department of Chemistry, Jadavpur University in 2011 on the occasion of “The International Year of Chemistry” with which the powder X-ray diffractometer and the HRMS facility was purchased. Authors are grateful to Dr Arup Gayen and his scholars for help in procuring PXRD data and wish to thank Dr M. Ali and his research scholars for their help in procuring mass spectrum data. SD is grateful to Dr Shouvik Chattopadhyay of the Department of Chemistry, Jadavpur University for kindly providing the IR data of the compounds used. BM expresses her gratitude to Dr Piyal Das, her lab senior, for help during enzyme assay experiments.References
- G. Minotti, P. Menna, E. Salvatorelli, G. Cairo and L. Gianni, Pharmacol. Rev., 2004, 56, 185–229 CrossRef CAS PubMed.
- K. C. Chow, T. L. Macdonald and W. E. Ross, Mol. Pharmacol., 1988, 34, 467–473 CAS.
- J. C. Wang, Nat. Rev. Mol. Cell Biol., 2002, 3, 430–440 CrossRef CAS PubMed.
- D. A. Koster, A. Crut, S. Shuman, M. Bjornsti and N. H. Dekker, Cell, 2010, 142, 519–530 CrossRef CAS PubMed.
- F. Arcamone and S. Penco, in Anthracycline and Anthracenedione-Based AnticancerAgents, ed. J. W. Lown, Elsevier, Amsterdam, 1988 Search PubMed.
- T. Terasaki, T. Iga, Y. Sugiyama, Y. Sawada and M. Hanano, J. Pharmacobiodyn, 1984, 7, 269–277 CrossRef CAS PubMed.
- J. Cummings and C. S. McArdle, Br. J. Cancer, 1986, 53, 835–838 CrossRef CAS PubMed.
- M. Gigli, S. M. Doglia, J. M. Millot, L. Valentini and M. Manfait, Biochim. Biophys. Acta, 1988, 950, 13–20 CrossRef CAS.
- (a) Y. Pommier, Curr. Med. Chem.: Anti-Cancer Agents, 2004, 5, 429–434 CrossRef; (b) Y. Pommier, Nat. Rev. Cancer, 2006, 6, 789–802 CrossRef CAS PubMed; (c) Y. Pommier, Chem. Rev., 2009, 109, 2894–2902 CrossRef CAS PubMed; (d) Y. Pommier, E. Leo, H. L. Zhang and C. Marchand, Chem. Biol., 2010, 17, 421–433 CrossRef CAS PubMed.
- C.-C. Wu, Y.-C. Li, Y.-R. Wang, T.-K. Li and N.-L. Chan, Nucleic Acids Res., 2013, 41(22), 10630–10640 CrossRef CAS PubMed.
- (a) E. J. Fox, Neurology, 2004, 63, S15–S18 CrossRef CAS PubMed; (b) L. J. Scott and D. P. Figgitt, CNS Drugs, 2004, 18(6), 379–396 CrossRef CAS PubMed.
- P. S. Guin, S. Das and P. C. Mandal, J. Inorg. Biochem., 2009, 103, 1702–1710 CrossRef CAS PubMed.
- (a) P. S. Guin, S. Das and P. C. Mandal, J. Phys. Org. Chem., 2010, 23, 477–482 CrossRef CAS; (b) P. Das, P. S. Guin, P. C. Mandal, M. Paul, S. Paul and S. Das, J. Phys. Org. Chem., 2011, 24, 774–785 CAS; (c) S. Mukherjee, P. Das and S. Das, J. Phys. Org. Chem., 2012, 25, 385–393 CrossRef CAS.
- P. Das, C. K. Jain, S. K. Dey, R. Saha, A. D. Choudhury, S. Roychowdhury, S. Majumder, H. K. Kumar and S. Das, RSC Adv., 2014, 4, 59344–59357 RSC.
- S. Roy, P. Mondal, P. S. Sengupta, D. Dhak, R. C. Santra, S. Das and P. S. Guin, Dalton Trans., 2015, 44, 5428–5440 RSC.
- W. Priebe, Molecules, 2000, 5, 299–301 CrossRef CAS.
- S. Mukherjee, P. Gopal, S. Paul and S. Das, J. Anal. Oncol., 2014, 3, 122–129 CrossRef.
- T. Banerjee and R. Mukhopadhyay, Biochem. Biophys. Res. Commun., 2008, 374, 264–268 CrossRef CAS PubMed.
- (a) M. M. L. Fiallo and A. G. Suillerot, Biochim. Biophys. Acta, Gen. Subj., 1985, 840, 91–98 CrossRef CAS; (b) H. Beraldo, A. Garnier-Suillerot, L. Tosi and F. Lavelle, Biochemistry, 1985, 24, 284–289 CrossRef CAS PubMed.
- I. D. Liegro, A. Cestelli, B. F. Matzanke, E. Bill and A. X. Trautwein, BioMetals, 1996, 9, 121–130 CrossRef PubMed.
- C. M. Pellei, V. Gandin, M. Porchia, F. Tisato and C. Marzano, Chem. Rev., 2014, 114, 815–862 CrossRef PubMed.
- A. Kheirolomoom, L. M. Mahakian, C. Y. Lai, H. A. Lindfors, J. W. Seo, E. E. Paoli, K. D. Watson, E. M. Haynam, E. S. Ingham, L. Xing, R. H. Cheng, A. D. Borowsky, R. D. Cardiff and K. W. Ferrara, Mol. Pharm., 2010, 7, 1948–1958 CrossRef CAS PubMed.
- M. M. L. Fiallo, A. Garnier-Suillerot, B. Matzanke and H. Kozlowski, J. Inorg. Biochem., 1999, 75, 105–115 CrossRef CAS PubMed.
- D. L. Ma, D. S. Chan and C. H. Leung, Acc. Chem. Res., 2014, 47, 3614–3631 CrossRef CAS PubMed.
- D. L. Ma, L. J. Liu, K. H. Leung, Y. T. Chen, H. J. Zhong, D. S. Chan, H. M. Wang and C. H. Leung, Angew. Chem., Int. Ed., 2014, 53, 9178–9182 CrossRef CAS PubMed.
- C. H. Leung, H. J. Zhong, D. S.-H. Chan and D. L. Ma, Coord. Chem. Rev., 2013, 257, 1764–1776 CrossRef CAS.
- C. H. Leung, S. Lin, H. J. Zhong and D. L. Ma, Chem. Sci., 2015, 6, 871–884 RSC.
- C. M. Pellei, F. Tisato and C. Santini, Adv. Anticancer Agents Med. Chem., 2009, 9, 185–211 CrossRef.
- A. A. Kumbhar, A. T. Franks, R. J. Butcher and K. Franz, Chem. Commun., 2013, 49, 2460–2462 RSC.
- S. Tardito and L. Marchiò, Curr. Med. Chem., 2009, 16, 1325–1348 CrossRef CAS PubMed.
- L. Finney, S. Mandava, L. Ursos, W. Zhang, D. Rodi, S. Vogt, D. Legnini, J. Maser, F. Ikpatt, O. I. Olopade and D. Glesne, Proc. Natl. Acad. Sci. U. S. A., 2007, 104, 2247–2252 CrossRef CAS PubMed.
- S. A. Lowndes and A. L. Harris, Journal of Mammary Gland Biology and Neoplasia, 2005, 10, 299–310 CrossRef PubMed.
- S. Zhai, L. Yang, Q. C. Cui, Y. Sun, Q. P. Dou and B. Yan, J. Biol. Inorg. Chem., 2010, 15, 259–269 CrossRef CAS PubMed.
- T. C. Johnstone and E. M. Nolan, Dalton Trans., 2015, 44, 6320–6339 RSC.
- R. J. Debus, The polypeptides of Photosystem II and their influence on mangano tyrosyl-based oxygen evolution, in Metal ions in biological systems: Manganese and its role in biological processes, ed. A. Sigel and H. Sigel, Dekker Marcel, New York, 2000, pp. 657–711 Search PubMed.
- M. Aschner, T. R. Guilarte, J. S. Schneider and W. Zheng, Toxicol. Appl. Pharmacol., 2007, 221, 131–147 CrossRef CAS PubMed.
- A. Takeda, Brain Res. Rev., 2003, 41, 79–87 CrossRef CAS PubMed.
- R. S. Magliozzo and J. A. Marcinkeviciene, J. Biol. Chem., 1997, 272, 8867–8870 CrossRef CAS PubMed.
- O. P. Soldin and M. Aschner, Neurotoxicology, 2007, 28, 951–956 CrossRef CAS PubMed.
- N. S. Jakubovics and H. F. Jenkinson, Microbiology, 2001, 147, 1709–1718 CrossRef CAS PubMed.
- D. G. Kehres and M. E. Maguire, FEMS Microbiol. Rev., 2003, 27, 263–290 CrossRef CAS PubMed.
- K. M. Papp-Wallace and M. E. Maguire, Annu. Rev. Microbiol., 2006, 60, 187–209 CrossRef CAS PubMed.
- R. E. Trouwborst, B. G. Clement, B. M. Tebo, B. T. Glazer and G. W. Luther III, Science, 2006, 313, 1955–1957 CrossRef CAS PubMed.
- T. G. Spiro, J. R. Bargar, G. Sposito and B. M. Tebo, Acc. Chem. Res., 2010, 43, 2–9 CrossRef CAS PubMed.
- R. Caspi, B. M. Tebo and M. G. Haygood, Appl. Environ. Microbiol., 1998, 64, 3549–3555 CAS.
- J. P. M. de Vrind, G. J. Brouwers, P. L. A. M. Corstjens, J. den Dulk and E. W. de Vrind-de Jong, Appl. Environ. Microbiol., 1998, 64, 3556–3562 CAS.
- G.-J. Brouwers, J. P. M. de Vrind, P. L. A. M. Corstjens, P. Cornelis, C. Baysse and E. W. de Vrind-de Jong, Appl. Environ. Microbiol., 1999, 65, 1762–1768 CAS.
- M. D. Vaira, P. Orioli, F. Piccioli, B. Bruni and L. Messori, Inorg. Chem., 2003, 42, 3157–3159 CrossRef PubMed.
- A. Altomare, M. Camalli, C. Cucci, C. Giacovazzo, A. Moliterni and R. Rizzi, J. Appl. Crystallogr., 2009, 42, 1197–1202 CrossRef CAS.
- V. Favre-Nicolin and R. Cerny, J. Appl. Crystallogr., 2002, 35, 734–743 CrossRef CAS.
- J. J. Stewart, J. Mol. Model., 2007, 13, 1173–1213 CrossRef CAS PubMed.
- A. C. Larson and R. B. Von Dreele, General Structure Analysis System (GSAS), Los Alamos National Laboratory Report LAUR, Los Alamos, NM, 2000, p. 86 Search PubMed.
- B. H. Toby, J. Appl. Crystallogr., 2001, 34, 210–213 CrossRef CAS.
- G. W. Castellan, in Physical Chemistry, Addison-Wesley/Narosa Publishing House, New Delhi, 3rd edn, 1989, p. 799, Indian Student Edition Search PubMed.
- S. Das, A. Saha and P. C. Mandal, Talanta, 1996, 43, 95–102 CrossRef CAS PubMed.
- H. R. Mahler, in Methods in Enzymology, ed. S. P. Colowick and N. O. Kaplan, Academic Press, New York, 1955, vol. 11, pp. 668–672 Search PubMed.
- H. Wang and J. A. Joseph, Free Radical Biol. Med., 1999, 27, 612–616 CrossRef CAS PubMed.
- O. Myhre, J. M. Andersen, H. Aarnes and F. Fonnum, Biochem. Pharmacol., 2003, 65, 1575–1582 CrossRef CAS PubMed.
- A. Gomes, E. Fernandes and J. L. F. C. Lima, J. Biochem. Biophys. Methods, 2005, 65, 45–80 CrossRef CAS PubMed.
- A. Pramanik, D. Laha, S. Chattopadhyay, S. K. Dash, S. Roy, P. Pramanik and P. Karmakar, Toxicol. Res., 2015, 4, 1604–1612 RSC.
- J. Schmidt, Organic Chemistry, ed. N. Campbell, Oliver and Boyd, London, 7th edn, 1955, p. 598 Search PubMed.
- G. Scatchard, Ann. N. Y. Acad. Sci., 1949, 51, 660–672 CrossRef CAS.
- P. S. Guin, P. Das, S. Das and P. C. Mandal, Int. J. Electrochem., 2012, 183745, DOI:10.1155/2012/183745.
- P. Aich and D. Dasgupta, Biochemistry, 1995, 34, 1376–1385 CrossRef CAS PubMed.
- S. Majee, R. Sen, S. Guha, D. Bhattacharya and D. Dasgupta, Biochemistry, 1997, 36, 2291–2299 CrossRef CAS PubMed.
- S. Das and D. Dasgupta, J. Inorg. Biochem., 2005, 99, 707–715 CrossRef CAS PubMed.
- S. Roy, R. Banerjee and M. Sarkar, J. Inorg. Biochem., 2006, 100, 1320–1331 CrossRef CAS PubMed.
- J. B. Chaires, Curr. Opin. Struct. Biol., 1998, 8, 314–320 CrossRef CAS PubMed.
- I. Haq, Arch. Biochem. Biophys., 2002, 403, 1–15 CrossRef CAS PubMed.
- J. B. Chaires, Biopolymers, 1985, 24, 403–419 CrossRef CAS PubMed.
- D. P. Mascotti and T. M. Lohman, Biochemistry, 1993, 32, 10568–10579 CrossRef CAS PubMed.
- A. J. M. Ferreri, E. Campo, A. Ambrosetti, F. Ilariucci, J. F. Seymour, R. Willemze, G. Arrigoni, G. Rossi, A. Lopez-Guillermo, E. Berti, M. Eriksson, M. Federico, S. Govi, S. Cortelazzo, N. Frungillo, S. Dell'Oro, M. Lestani, S. Asioli, E. Pedrinis, M. Ungari, T. Motta, R. Rossi, T. Artusi, P. Iuzzolino, E. Zucca, F. Cavalli and M. Ponzoni, Ann. Oncol., 2004, 15, 1215–1221 CrossRef CAS PubMed.
- E. V. Barry, S. E. Lipshultz and S. E. Sallan, Anthracycline-induced cardiotoxicity: natural history, risk factors, and prevention, in American Society of Clinical Oncology 2008 Educational Book, ed. R. Govindan, American Society of Clinical Oncology, Alexandria, 2008, pp. 448–453 Search PubMed.
- K. J. Schimmel, D. J. Richel, R. B. van den Brink and H. J. Guchelaar, Cancer Treat. Rev., 2004, 30, 181–191 CrossRef CAS PubMed.
- M. I. Gharib and A. K. Burnett, Eur. J. Heart Failure, 2002, 4, 235–242 CrossRef CAS PubMed.
- L. C. M. Kremer, E. C. V. Dalen, M. Offringa and P. A. Voute, Ann. Oncol., 2002, 13, 503–512 CrossRef CAS PubMed.
- S. M. Swain, F. S. Whaley and M. S. Ewer, Cancer, 2003, 97, 2869–2879 CrossRef CAS PubMed.
- D. Iarussi, P. Indolfi, F. Casale, V. Martino, M. T. Tullio and R. Di Calabrò, Paediatr. Drugs, 2005, 7, 67–76 CrossRef PubMed.
- D. L. Keefe, Semin. Oncol., 2001, 28, 2–7 CrossRef CAS PubMed.
- D. Cardinale, A. Colombo, G. Lamantia, N. Colombo, M. Civelli, G. D. Giacomi, M. Rubino, F. Veglia, C. Fiorentini and C. M. Cipolla, J. Am. Coll. Cardiol., 2010, 55, 213–220 CrossRef CAS PubMed.
- S. Das, A. Bhattacharya, P. C. Mandal, M. C. Rath and T. Mukherjee, Radiat. Phys. Chem., 2002, 65, 93–100 CrossRef CAS.
Footnote |
| † Electronic supplementary information (ESI) available: Physicochemical experiments on complex formation in solution including equations leading to evaluation of stability constant. Spectroscopic characterization (IR, mass) of the complex. Equations for the evaluation of binding parameters for the interaction of the compounds with calf thymus DNA. CIF and check CIF pdf files of MnII(LH2)2. CCDC 1479625. For ESI and crystallographic data in CIF or other electronic format see DOI: 10.1039/c6ra09387f |
| This journal is © The Royal Society of Chemistry 2016 |