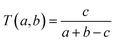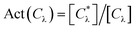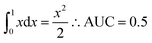A chemical space odyssey of inhibitors of histone deacetylases and bromodomains†
Fernando D. Prieto-Martínez,
Eli Fernández-de Gortari,
Oscar Méndez-Lucio and
José L. Medina-Franco *
*
Facultad de Química, Departamento de Farmacia, Universidad Nacional Autónoma de México, Avenida Universidad 3000, Mexico City 04510, Mexico. E-mail: medinajl@unam.mx; jose.medina.franco@gmail.com; Tel: +52-55-5622-3899 ext. 44458
First published on 6th June 2016
Abstract
The interest in epigenetic drug and probe discovery is growing as reflected in the large amount of structure-epigenetic activity information available. Therefore, the significance of understanding the entire or fractions of the epigenetic relevant chemical space is increasing. Major epigenetic targets are histone lysine deacetylases (HDACs), bromodomains (BRDs), and DNA methyltransferases (DNMTs). However, with the exception of DNMTs, characterization of the chemical space of these epi-targets is limited. This work is the first chemoinformatic analysis of the physicochemical properties, structural diversity, and coverage of the chemical space of compounds screened as inhibitors of HDACs and BRDs. The chemical space was compared to DNMTis, approved drugs, commercial screening compounds, and generally recognized as safe (GRAS) molecules. The structural complexity of compounds directed towards epigenetic targets was also addressed. The outcome of this analysis indicated that it is required to increase the structural diversity and molecular complexity of screening libraries tested as modulators of DNMTs, HDACs and BRDs. Results also suggested that it is feasible to develop dual inhibitors targeting HDACS and BRDs. This work has implications in repurposing of food chemicals with potential epigenetic activity and design of poly-epigenetic compounds.
1. Introduction
Every living being has the ability to inherit its genetic material. However this process is not flawless. After a few decades, the study of DNA repair lead to the discovery of higher order mechanisms and the term ‘epigenetics’ was coined.1 Initially, inhibition of epigenetic targets was considered a novel alternative for the treatment of cancer. While this approach may be true, current research showed that environmental factors such as radiation exposure, nutritional history, dietary intake, reproductive factors, among others, also play a key role on the expression of specific epigenetic modifications.1–3 Nowadays it is accepted that epi-modulation can act as a link between genotype and environment stimuli4 and may be used as a Rosetta Stone to better understand, prevent and/or cure diseases. While this is yet to be proved, many researchers consider epigenetics as the missing link on the biogenesis of chronic diseases like Alzheimer's, dementia, schizophrenia, diabetes, metabolic syndrome, to name few examples.5,6Chemical modifications are key features in epigenetics. Although the number of reactions and enzymes involved are different and comprise more than one hundred, it is possible to distinguish three functions: writers, erasers and readers. Writers add chemical groups that can be labile or stable. Erasers remove the groups added by writing enzymes. Readers are ‘effector proteins’ that identify specific chemical groups associated with epigenetic modifications and produce large scale changes such as chromatin remodeling or recruitment of other enzymes involved in DNA replication or gene expression.7
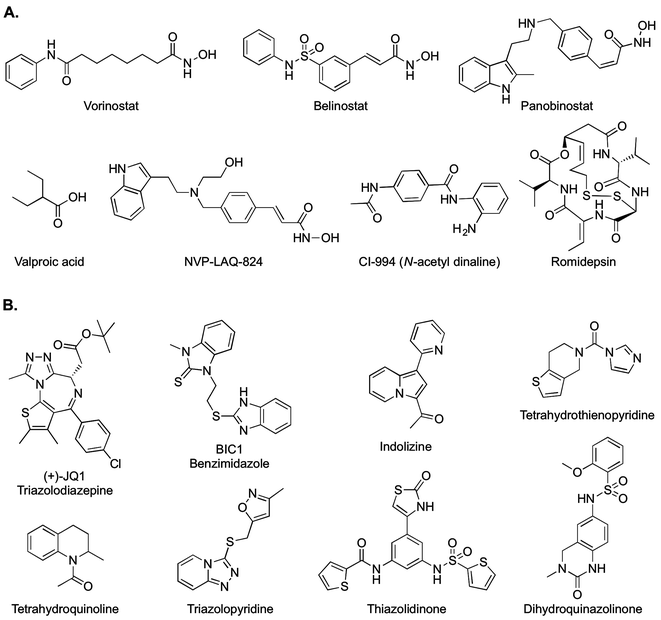
The correlation between epigenetic changes and carcinogenesis attracted attention to histone deacetylases (HDACs). Acetylation on lysine residues is one of the most common processes on epigenetics.1 Eighteen different HDACs have been identified, characterized and classified in three classes. Class I comprises HDACs 1, 2, 3 and 8, that are located on the nucleus with involvement on development of numerous cancer types.8 Class III gathers seven HDACs that are NAD+ dependent and sirtuin constituted and are known as SIRT 1–7. This class has been mainly involved with pancreas and breast cancer, nevertheless some of them (e.g. SIRT1) may be involved with type II diabetes.9 Although the removal of acetate groups from histone tails may be conceived as the first step towards transcriptional repression, it has been shown that regulation by HDACs goes beyond histones acting in a plethora of cellular pathways.10 Despite major efforts from industry and academia and the baffling amount of chemical and biochemical studies towards these enzymes, the Food and Drug Administration (FDA) of the United States has approved only four drugs so far for clinical use: vorinostat, romidepsin, belinostat, and panobinostat. Fig. 1A shows the chemical structures of different HDAC inhibitors (HDACis) including those approved for clinical use.
More recently, epigenetic drug and probe discovery has turned to other targets such as the Bromodomain and Extra-Terminal domain (BET) protein family. From this family, the bromodomains (BRDs) BRD2, BRD3 and BRD4 have become of particular therapeutic interest. BRDs are structural motifs that recognize acetylated lysines, mainly those located in histone tails.11 BRDs decide the ultimate fate of histones and their function has been correlated with cancer and inflammatory disease.12 Also BRDs are responsible for crucial steps during cell cycles.13 Fig. 1B depicts representative BRDs inhibitors (BRDis) including different scaffolds and acetyl-lysine mimicking groups.
Drug development is a daunting task and epigenetic drug discovery is not an exception. One of the main drawbacks is the high attrition rate,14 which may be associated with the traditional trend in drug discovery to design specific drugs. Indeed, there is increasing evidence that drugs may act as ‘master key compounds’,15 i.e. drugs exhibit biological activity through the interaction with a set of selected targets with reduced affinity for off-targets (at therapeutic doses). Thus, polypharmacology is largely influencing drug discovery strategies including the discovery of epigenetic drugs.16 One of the strategies to explore the development of poly-epigenetic compounds is the assessment of the diversity and chemical space coverage of compounds with known epigenetic activity. The putative intersection in chemical space by approved drugs and compounds with epigenetic activity may lead to strategies to conduct drug repurposing.17 Similarly, the intersection in chemical space of epigenetic compounds and diverse screening collections offers the possibilities to guide focused library design within novel regions in chemical space.18 Furthermore, the comparison of chemical spaces of epigenetic compounds and food-related molecules may lead to the systematic elucidation of food chemical as bioactive epigenetic molecules. As a proof-of-concept, chemoinformatic mining of generally recognized as safe (GRAS) compounds, widely used in the food industry, lead to the identification of HDACis.19
There are several chemoinformatic tools that enable the analysis of the coverage and diversity of the chemical space of public data sets of epigenetic compounds. Despite the fact that public databases do not necessarily cover the entire current knowledge of epigenetic collections, they represent a reasonable starting point to better understand the entire or fractions of the chemical space covered by epigenetic-related compounds i.e., Epigenetic Relevant Chemical Space (ERCS). In fact, the wide variety of structures and molecules currently studied by epigenetics accounts for more than 5000 compounds available in the public domain with structure–activity data. As preliminary data, the chemical space of DNMT1 inhibitors (DNMTis) has been recently reported.20
The goal of the present study is to characterize the chemical space of HDACis and BRDis currently stored in two major public databases: ChEMBL and Binding Database (see below). Therefore, in light of the emerging research area of Epi-informatics,21 this work contributes to the understanding of a fraction of ERCS. Of note, the compounds analyzed thought the study and deposited in ChEMBL and Binding Database have been developed not as part of drug discovery projects but also in the development of molecular probes. Chemoinformatic characterization of compound data sets is extensively documented by our and other groups to be an essential component of drug, lead and molecular probe discovery projects.22,23 As discussed through the study and emphasized in the Conclusions section, the outcome of the analysis provided specific information that can be used to develop novel and improved epigenetic compounds. These collections were compared to DNMTis, approved drugs, compounds in clinical trials, a general screening collection typically used in high-throughput screening (HTS), and a commercial screening collection focused on epigenetic targets. Moreover, the epigenetic-related libraries were compared to GRAS chemicals. In order to conduct the comparisons, four complementary criteria were used including physicochemical properties (PCP) of pharmaceutical relevance, molecular fingerprints, molecular scaffolds, and established measures of molecular complexity. To the best of our knowledge, this is the first analysis of the structural complexity of epigenetic databases. As discussed throughout the study, the insights of these analyses provided sound basis to conduct computer-aided drug repurposing, identify bioactive compounds from food chemicals, and guide library design. The intersection of epigenetic target spaces may also indicate the feasibility of developing dual or poly-epigenetic modulators. Of note, the findings of this work directly impact not only the development of therapeutic agents but also on molecular probes.
2. Methods
The epigenetic and reference collections were retrieved from public databases. Curated data sets were analyzed and compared to each other using complementary molecular representations: six PCP, three structural fingerprints, and molecular scaffolds. The methods used to represent the molecular structures and perform the visual and quantitative comparisons are described in this section.2.1 Datasets and data curation
The chemical structures and activity data of HDACis (against isoforms 1, 2, 3, 8 and SIRT1) and BRDis (against isoforms 2, 3, 4) were downloaded from ChEMBL24 version 20, and Binding Database (BDB).25 The HDACis analyzed in this work had been associated with the treatment of different types of cancer including colon, prostate, gastric and cervical,8 and they may also play key roles on diabetes26 and cardiovascular diseases.27 Likewise, the BRDis had been associated with activity against cancer cell lines.26 The structures were downloaded as simplified molecular input line entries (SMILES) and then converted to three-dimensional (3D) structures using Molecular Operating Environment (MOE).28 To curate the data sets, which is a major step in chemoinformatic analysis of compound data sets29 we employed the ‘Wash’ module implemented in MOE. Using this module, structures were prepared by disconnecting metal salts, remove simple components, and rebalancing protonation states. Molecules were energy minimized with the MMFF94x force field using the default stochastic sampling algorithm followed by LowModeMDsearch implemented in MOE. The stereochemistry annotated in the SMILES string was used. The resulting data sets were visually inspected and final details were corrected. Table 1 summarizes the compounds initially downloaded and the number of compounds after curation.| Size | Source | URL | ||
|---|---|---|---|---|
| Number of compounds | Unique molecules | |||
| Data set | ||||
| HDACs | >5000 | 2000 | ChEMBL | https://www.ebi.ac.uk/chembl/ |
| BRDs | ∼2000 | 207 | ChEMBL, BDB | https://www.bindingdb.org/ |
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) |
||||
| Reference sets | ||||
| DNMTs | >5000 | 565 | ChEMBL, BDB, HEMD | http://xlink.rsc.org/?DOI=C5RA19611F |
| Drugs | >5000 | 1490 | DrugBank | http://www.drugbank.ca/drugs/ |
| Clinic | 1151 | 837 | Therapeutic target database | http://bidd.nus.edu.sg/group/cjttd/ |
| General | 1224 | 1100 | Selleck | http://www.selleckchem.com |
| Epi-focused | 128 | 113 | Selleck | http://www.selleckchem.com |
| GRAS | 2200 | 2200 | FEMA | http://dx.plos.org/10.1371/journal.pone.0050798 |
2.2 Molecular representation
Since the chemical space depends on the structure representation, multiple criteria were implemented, namely; PCP of pharmaceutical relevance, structural fingerprints of different design and molecular scaffolds.20,30 Structural complexity using well-established methods was also measured.31 Details of each method are provided hereunder.![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P), rotatable bonds (RTB), hydrogen bond acceptors (HBA), hydrogen bond donors (HBD), topological surface area (TPSA), and molecular weight (MW). This set of properties is typically used to compare compound data sets in drug discovery20 and was employed to make cross-comparisons with studies that report the characterization of other data sets. The distribution of the properties was assessed with box plots, generated with R Studio using the PMCMR package,32 and summary statistics of the distributions. In order to generate a visual representation of the chemical space based on properties, principal component analysis (PCA) was conducted with MOE. Data visualization was performed with Data Warrior.33 The statistical comparison of the properties was made by assessment of homoscedasticity (data values follow a normal distribution). For this approach we made a Shapiro test, followed by a Kruskal–Nemenyi test. Allowing pairwise comparisons for which p < 0.05 for acceptance of null hypothesis i.e., there is not a statistically significant difference between the distributions in the data sets.
P), rotatable bonds (RTB), hydrogen bond acceptors (HBA), hydrogen bond donors (HBD), topological surface area (TPSA), and molecular weight (MW). This set of properties is typically used to compare compound data sets in drug discovery20 and was employed to make cross-comparisons with studies that report the characterization of other data sets. The distribution of the properties was assessed with box plots, generated with R Studio using the PMCMR package,32 and summary statistics of the distributions. In order to generate a visual representation of the chemical space based on properties, principal component analysis (PCA) was conducted with MOE. Data visualization was performed with Data Warrior.33 The statistical comparison of the properties was made by assessment of homoscedasticity (data values follow a normal distribution). For this approach we made a Shapiro test, followed by a Kruskal–Nemenyi test. Allowing pairwise comparisons for which p < 0.05 for acceptance of null hypothesis i.e., there is not a statistically significant difference between the distributions in the data sets.Structural similarity was computed with the Tanimoto coefficient:34,35
The intra-library similarity of a given data set with N compounds was measured as the distribution of the N(N − 1)/2 pairwise similarity values. The inter-library similarity was analyzed by means of nearest-neighbor curves. These curves represent the distribution of the maximum similarity values of molecules in a test set with respect to the molecules in the reference set.36 In this study, six data sets (i.e., BRDs, HDAC and DNMT1, GRAS, ‘Drugs’ and ‘Clinic’) were used as reference and test sets, e.g., they were compared to each other. The distribution of the intra- and inter-library similarity values was analyzed by means of cumulative distribution functions (CDF) generated with matplotlib.pyplot Python scripts.37
2.2.3.1 Scaffold diversity. The scaffold diversity of the BRDs and HDACs data sets was evaluated using cyclic systems recovery (CSR) curves. Briefly, a CSR curve measures the fraction of cyclic systems contained in a given fraction of the database. To generate this curve, the list of cyclic systems for each data set was ordered by frequency. Then, the fraction of cyclic systems was plotted on the X axis and the fraction of compounds containing cyclic systems was plotted on the Y axis. The CSR curve was characterized by the area under the curve (AUC) and the fraction of cyclic systems that contain 50% of the corresponding data set (F50).43 The development and application of CSR curves is discussed elsewhere.43,44
2.2.3.2 Chemotype enrichment. For the most frequent scaffolds of BRDs and HDACs, the proportion of active compounds in a given scaffold relative to the fraction of active compounds in the entire data set was analyzed. The chemotype enrichment analysis was based on the definition of scaffolds given by Johnson and Xu (vide supra). To measure scaffold enrichment we employed an established methodology.45,46 The background or baseline activity of the corresponding data set was calculated using the equation:
| Act(C) = [C*]/[C] |
The fraction of active compounds in a specific chemotype Act(Cλ) was calculated with the expression:
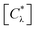 are the total number of compounds and active compounds, respectively, in the chemotype class λ.
are the total number of compounds and active compounds, respectively, in the chemotype class λ.
The enrichment factor (EF) for chemotype λ was calculated with the equation:
| EF(Cλ) = Act(Cλ)/Act(C) |
Thus, EF(Cλ) measured the proportion of active molecules of a particular chemotype relative to the proportion of active compounds in the dataset. In this manner, the molecular scaffolds with the highest EF were flagged as the most attractive. To further differentiate the most attractive cyclic systems i.e., molecular scaffolds with the highest frequency, chemotype enrichment plots were generated plotting the EF on the X-axis and the cyclic systems frequency on the Y-axis.45 Chemotype enrichment plots have been used in the scaffold analysis of compound databases, including the scaffold analysis of DNMTis.20,45,47
3. Results and discussion
Table 1 summarizes the number of compounds before and after data curation. In total, we analyzed 2000 unique compounds tested against HDACs and 207 unique molecules tested against BRDs. Of note, in this work we did not differentiate between different isoforms of HDACs and BRDs; first we aimed to obtain a first overall assessment of the chemical space coverage and distribution of the compounds that have been tested with these epigenetic targets. Despite the fact that most of the HDACs inhibitors studied in this work are from class I, a follow-up study will be conducted to distinguish the chemical space of different classes.The degree of activity of each set was explored taking as reference an IC50 value of 1 μM to define an ‘active’ compound. Following this heuristic criterion, a high percentage (79.5%) of the HDAC data set was composed of active compounds, whereas only 46.85% of the molecules in the BRDs dataset had IC50 < 1 μM. This result is in line with the amount of activity data accumulated to optimize the activity of HDACis as compared to the current development of BRDis.
3.1 Physicochemical properties
Fig. 2 shows box plots of the distribution of the six PCPs calculated for the epigenetic and reference data sets. Table S1 in the ESI† summarizes the statistics of each distribution for all data sets. BRDs and HDACs sets had similar distributions of HBA that are comparable to the distribution for the general screening collection (‘General’) and the commercial collection focused on epigenetic targets (‘Epi-focused’) (i.e. median value of 4). Overall, DNMTs had a slightly higher number of HBA while approved drugs, compounds in the clinic (‘Clinic’) and, more markedly GRAS, have fewer HBA. In a similar way, HDACs and DNMTs had slightly higher number of HBD (i.e. median of 3) compared to ‘General’ and ‘Epi-focused’ datasets that presented a median of 2 HBD, or to BRDs, ‘Drugs’, and ‘Clinic’ dataset with a median of 1 HBD. Similar trends where obtained for TPSA that is other measure associated with polarity. HDACs and DNMTs had higher values of TPSA than other reference collections.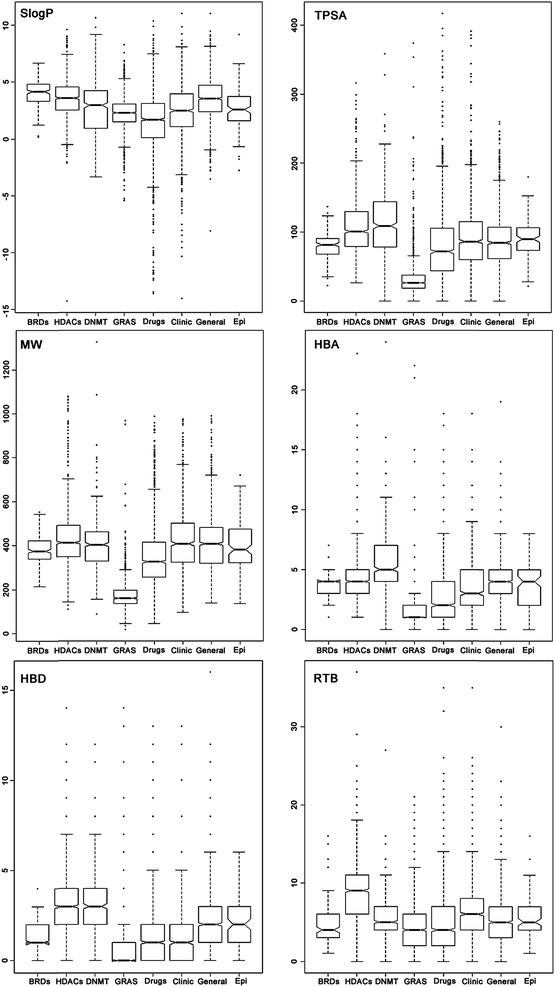 | ||
| Fig. 2 Box plots of the distribution of six physicochemical properties of pharmaceutical relevance for the BRDs, HDACs, DNMTs and reference data sets. Summary statistics are in Table S1 of the ESI.† | ||
Regarding Slog![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P as a measure of hydrophobicity, BRDs had, on average, the highest values across all the data sets. The Slog
P as a measure of hydrophobicity, BRDs had, on average, the highest values across all the data sets. The Slog![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P of HDACs was comparable to the ‘General’ set and had the second highest values. As per compound flexibility, measured by RTB, BRDs presented a similar mean value compared to the other reference collections, including GRAS. Noteworthy, HDACs had a higher mean number of RTB. This is due to the presence of peptide molecules that have been largely explored as HDACis. Overall, HDACs presented the highest mean MW, nevertheless all the other dataset (with exception of GRAS) presented comparable values of MW.
P of HDACs was comparable to the ‘General’ set and had the second highest values. As per compound flexibility, measured by RTB, BRDs presented a similar mean value compared to the other reference collections, including GRAS. Noteworthy, HDACs had a higher mean number of RTB. This is due to the presence of peptide molecules that have been largely explored as HDACis. Overall, HDACs presented the highest mean MW, nevertheless all the other dataset (with exception of GRAS) presented comparable values of MW.
Statistical analysis using the Nemenyi test (Table S3 in the ESI†) revealed that, overall, BRDs is similar to ‘General’ and ‘Epi-focused’. This result could suggest that compounds tested for BRD inhibition came from generally screening collections commercially available. Of note, the ‘Epi-focused’ set is also commercially available (Table 1) and it was assembled from a generally screening library. The HDACs set also showed to be similar to the ‘General’ and ‘Epi-focused’ sets, and it has some degree of similarity to the PCP of the DNMT set (for example in terms of HBD, TPSA, and MW).
Statistical tests were used to determine whether there is a significant difference between ‘active’ compounds (IC50 < 1 μM) in the HDACs and BRDs and the entire sets. It was found that the most active compounds were not statistically different from the inactive compounds based on PCP (data not shown).
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P had the highest contribution to PC2. Of note, all these properties are associated with compound polarity.
P had the highest contribution to PC2. Of note, all these properties are associated with compound polarity.
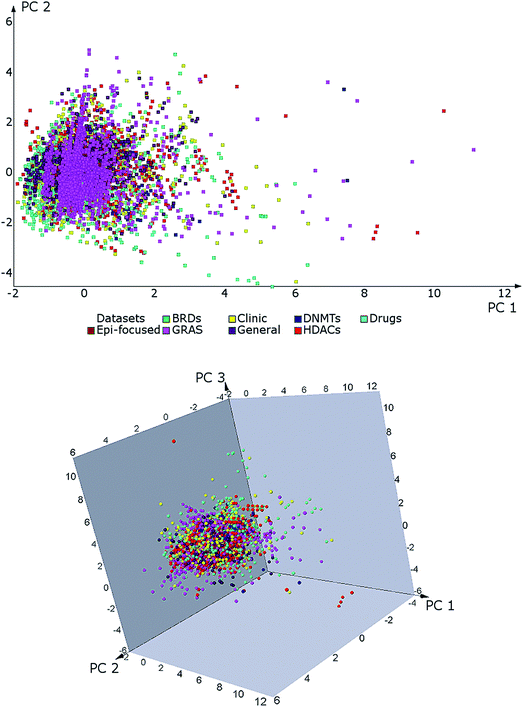 | ||
| Fig. 3 2D and 3D visual representations of the chemical space of BRDis, HDACis, DNMTis, GRAS and reference data sets. The plots were generated with principal component (PC) analysis of six physicochemical properties of pharmaceutical relevance. The first two PC recover 82% of the variance and the first three, 90%. Data sets are shown separately in Fig. S1 (ESI†). Outliers in HDACs and GRAS sets are not shown for clarity. See main text for details. | ||
Fig. 3 shows that BRDs and HDACs cover similar regions of the property space of DNMTs, ‘Drugs’, and the other reference collections. In particular, BRDs cover the smallest region of the property space (Fig. S1†), followed by ‘Epi-focused’, which is nearby in the ERCS. This may be related not only to the fewer number of compounds in these sets (207 compounds and 113, respectively), but also to the type of chemical structures. Statistical analysis (Nemenyi values shown on Table S3†) confirmed that BRDs and ‘Epi-focused’ are significantly similar in properties to each other. Compounds tested for HDAC inhibition show a broader distribution on the property space. The outliers of the HDACs set are mainly associated with peptides. Overall, this type of compounds are more flexible and more polar than small molecules used in drug discovery and may have, depending on the nature of the peptide, a large MW.
Analysis of the property distribution and visual representation of the chemical space of GRAS revealed remarkable trends. While GRAS shares common regions with the fraction of ERCS studied in this work, the coverage of property space of GRAS is scattered (Fig. 3 and S1†). Most of the outliers in GRAS are similar to the outliers in HDACs. This finding may be attributed to the nature of food additives as there are flexible molecules as sugars or even peptide derivatives. It is noteworthy the similar distribution of Slog![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P values between GRAS and ‘Epi-focused’ (Fig. 2). It has been discussed that hydrophobicity as measured by log
P values between GRAS and ‘Epi-focused’ (Fig. 2). It has been discussed that hydrophobicity as measured by log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P is one of the most important PCP to develop bioactive compounds.53 This novel finding that emerged from this comparison is related to the association between food chemicals and epigenetics.
P is one of the most important PCP to develop bioactive compounds.53 This novel finding that emerged from this comparison is related to the association between food chemicals and epigenetics.
Despite the fact that the analysis of the distribution of the PCP and visual representation of the chemical space based on properties of pharmaceutical relevance is important, they do not provide direct information on the nature of the chemical structures of the epigenetic sets. Structural aspects of the epigenetic-related compounds are addressed in the next section.
3.2 Structural fingerprints
Structural fingerprints enabled the comparison of the epigenetic data sets considering the atom connectivity and chemical structures. As detailed in the Methods section, three fingerprints of different design were employed, namely MACCS keys (166 bits), GpiDAPH3, and TGD. These representations have been used in the chemoinformatic characterization of the structural diversity of a number of compound libraries, including compounds tested with DNMT1 activity.20,49,54 Fingerprint-based representation enabled an analysis of the intra- and inter-compound set similarity that are discussed in the next two sub-sections.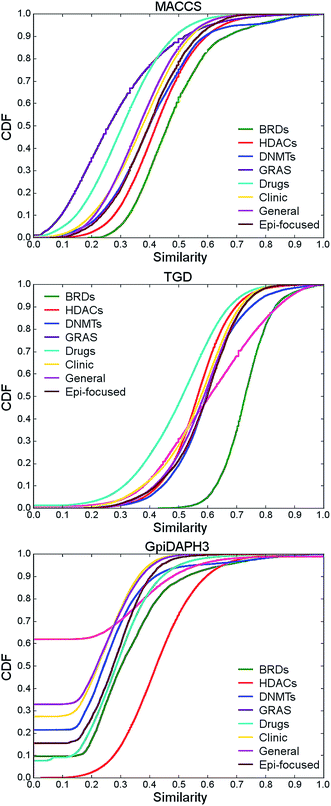 | ||
| Fig. 4 Intra-library similarity: cumulative distribution function (CDF) of the pairwise similarity values for BRDs, HDACs and reference data sets using MACCS keys, GpiDAPH3, and TGD fingerprints. The statistics of each distribution is summarized in Table S5 of the ESI.† | ||
According to MACCS keys, BRDs was the data set with the highest intra-set similarity followed by HDACs i.e., median similarity values of 0.463 and 0.423, respectively. In other words, these two collections had the lowest structural diversity as compared to all other reference data sets including DNMTs and the ‘Epi-focused’ collections. For reference, the most diverse sets were GRAS followed by ‘Drugs’; both sets had the lowest distribution of MACCS keys/Tanimoto similarity values (i.e., median values of 0.261 and 0.308, respectively). The high structural diversity of approved drugs has been reported49 and is in line with the fact that approved drugs in DrugBank cover a wide number of molecular targets and mechanisms of action. Interestingly, compounds in the ‘Clinic’ are less diverse than ‘Drugs’ and the structural diversity of ‘Clinic’ is comparable to the diversity of the general screening collection, ‘General’ (Table S5†). Similar to the conclusions obtained with MACCS keys, BRDs is one of the most similar (less diverse) sets considering GpiDAPH3 and TGD representations. Interestingly, according to GpiDAPH3, BRDs have comparable diversity to ‘Drugs’.
The structural diversity of HDACs highly depended on the molecular representation (Fig. 4 and Table S5†). According to GpiDAPH3, the HDACs set is the most similar (less diverse). But considering MACCS keys, HDACs is one of the most diverse sets with similarity comparable to ‘Clinic’.
Of note, there was not a general relationship between the size of the data sets with structural diversity. It could be anticipated that smaller data sets are less diverse than bigger ones. For instance, the low diversity of BRDs may be associated with the fewer number of molecules (207, see Table 1). Note, however, that ‘Epi-focused’ has even fewer molecules than BRDs (113) but is more diverse. This result can be rationalized considering that ‘Epi-focused’ was designed considering several epigenetic targets. A second example is the lower diversity of HDACs as compared to ‘Drugs’, ‘Clinic’, and ‘Epi-focused’ (Fig. 4) despite the fact that the size of the HDACs set (2000 compounds) is bigger than the size of the reference sets (1, 490, 837, and 113 compounds, respectively).
Taking together the results of the molecular diversity using different fingerprints it can be concluded that the BRDs set analyzed in this work does not have a large structural diversity. This result is related to the fact that research groups are developing derivatives of specific type of compounds such as benzodiazepines for BRD inhibition.11 These results clearly indicated the need to develop new chemical structures as BRDs. This far, compounds cover a limited region of chemical space and there is a large opportunity of increase novelty. The higher structural diversity of HDACs, as compared to BRDs, can be expected from the different type of compounds that have been reported as HDACis. However, one of the most interesting and active compounds are the hydroxamic acid derivatives. For these compounds, specific structural features change but keeping the same pharmacophoric features.55 This fact can be associated with the fact that HDACs have, overall, higher GpiDAPH3/Tanimoto similarity values but lower MACCS/Tanimoto similarity values (e.g., MACCS keys is more ‘sensitive’ to the changes in chemical modifications). The relative high diversity of GRAS compounds using different representation is also worth noting. The intra-set similarity results also highlighted the convenience of using multiple molecular representations for a comprehensive assessment of the molecular diversity of compound data sets.56
Fig. 5 shows CDF plots of the maximum similarity of five test sets with BRDs, HDACs, and DNMTs as reference sets using MACCS keys and TGDs. The CDFs for other reference sets are shown in Fig. S2 in the ESI.† Summary statistics from the distributions using MACCS keys and TGD are presented in Tables S6 and S7 (ESI†), respectively.
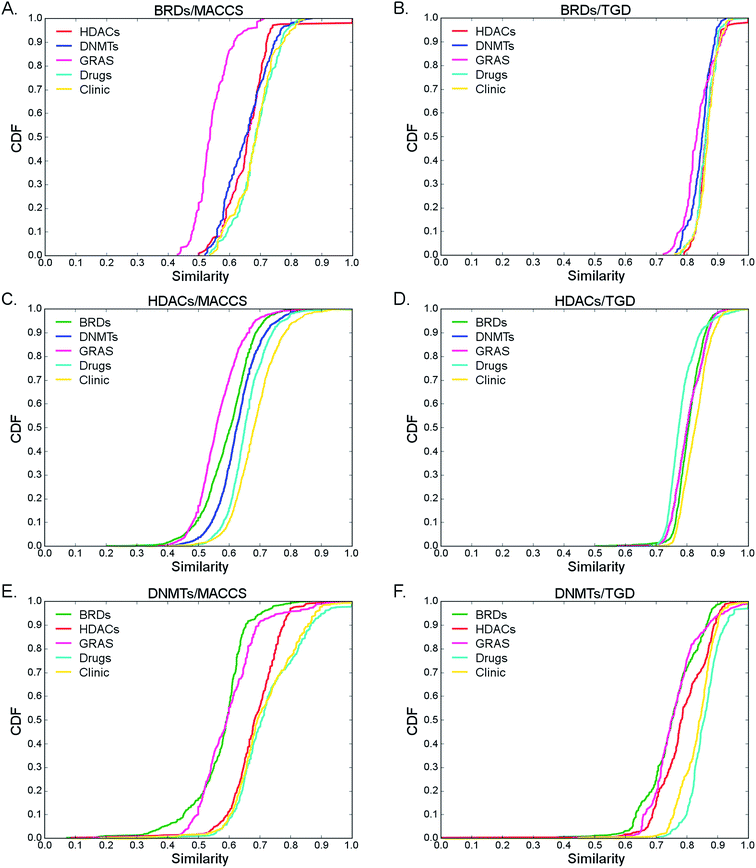 | ||
| Fig. 5 Inter-library similarity: cumulative distribution function (CDF) of the maximum structure similarity calculated with MACCS keys, TDG and the Tanimoto coefficient comparing three epigenetic data sets, ‘Drugs’ and ‘Clinic’ with BRDs, HDACs and DNMTs. The reference set is indicated at the top of each graph. Summary statistics of the CDFs are presented in Tables S6 and S7 in the ESI.† | ||
The low values in the CDFs and statistics obtained with MACCS keys and TGD indicated that the epigenetic sets and the reference collections analyzed in this section have, in general, compounds with different chemical structures as compared to BRDs, HDACs and DNMTs. However, the distribution of maximum similarity values showed that there are compounds in the epigenetic-related data sets with similarity value of one. After inspection of pairs of compounds that present MACCS keys and TGD similarity value of one, we found that although there are not identical structures, they share similar motifs. For instance, the presence of biphenyl derivatives with hydroxamic acid moiety on both sets is noteworthy. In fact, a dual active HDAC/bromodomain and extra terminal (BET) small molecule tool inhibitor with a hydroxamic acid has been published.57 Other moiety shared by the two sets is the phenylsulfonamide.
The CDFs and statistics also showed that ‘Drugs’ and ‘Clinic’ are, on average, more similar to BRDs, HDACs and DNMTs than the similarity showed among the three epigenetic sets of compounds. These results further highlights that, in general, the chemical structures tested as inhibitors of BRDs, HDACs and DNMTs, are different.
According to MACCS keys, the relative order or maximum similarity values of GRAS to the epigenetic sets is DNMTs > HDACs > BRDs. In other words, in similarity searching using MACCS keys, it is more likely to identify GRAS molecules similar to DNMTs. However, comparing GRAS to the epigenetic sets using TGD fingerprints, the relative order of maximum similarity values is different: BRDs > HDACs > DNMTs. Taken together, these results suggest that in similarity searching, more than one molecular representation should be employed and then select consensus hits. A detailed discussion of the comparison of all data sets studied in this work (Table 1) with each other is beyond the scope of this work that is focused on BRDs, HDACs and DNMTs.
3.3 Molecular scaffolds
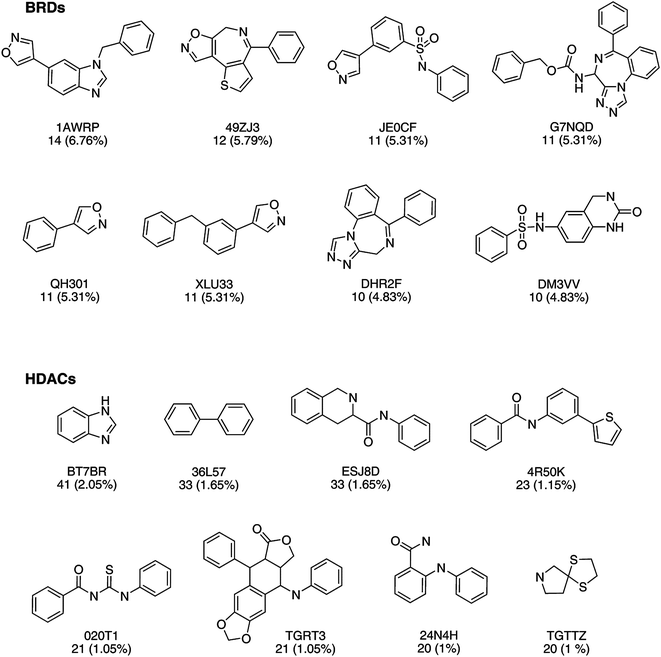
The analysis of the molecular scaffolds is organized in three major groups: scaffold content, diversity, and activity distribution.The most frequent scaffold in the BRDs set (cyclic system 1AWRP) had a frequency of 14 (6.8%) compounds followed by the cyclic system 49ZJ3 with 12 (5.8%) molecules. Most of the frequent cyclic systems had between 3 and 4 rings (Fig. 6). In contrast, the most populated cyclic system as defined by MEQI for HDACs (41 compounds, 2%) had only one ring. Interestingly this cyclic system is the benzimidazole ring which is a sub-structure of the most frequent cyclic system in the BRDs set (1AWRP, Fig. 6). The high prevalence of benzimidazole ring in bioactive compounds is well documented.54,58 Such findings increase the interest to design polyepigenetic drugs using the benzimidazole ring as a key sub-structure.
Not surprisingly, the benzene ring (cyclic system RYLFV) is highly frequent in the HDACs data set and had a frequency of 127 (6.35%) compounds. This cyclic system is highly frequent in approved drugs and several other compound collections.41,54 Surprisingly, the benzene ring is not present as a cyclic system (i.e., core scaffold) in the BRDs set although it is part of the structure.
Acyclic structures (chemotype identifier ‘00000’) are amongst the most frequent structures in the HDACs set with 43 (2.15%) molecules. Similar to the benzene ring, the number of acyclic structures is also common in other data sets.54 In contrast, no acyclic structures were found in the BRDs set.
The most frequent cyclic system identified in the BRDs and HDACS sets are not included in the ‘not wanted’ list and are not flagged as scaffold that have propensity to form multi-target activity cliffs.59,60 However one of them (DM3VV) shows a PAINS-like moiety; it has a structure that may break down causing false positives results in biological assays (Fig. 6).60
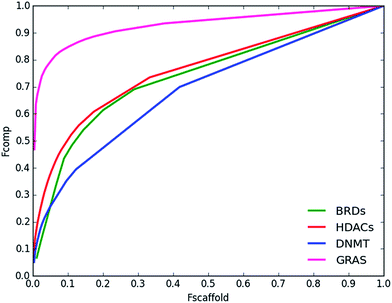 | ||
| Fig. 7 Cyclic system recovery curves for the epigenetic-relevant data sets and GRAS compounds. Summary statistics are presented in Table S8 of the ESI.† | ||
To measure the scaffold diversity with the CSR curves the following rationale was used. The fraction of a given cyclic system is compared to the fraction of the compounds in the data set contained in that group of cyclic systems. If we consider a reference ‘most-diverse-set’ i.e., a set in which every molecule has its own scaffold, the CSR ‘curve’ is a diagonal line. As y equals x AUC is the integral:
According to this expression, for the maximum diversity AUC = 0.5. It follows that as the AUC value increases (up to a maximum of one), the dataset contains higher number of compound with the same scaffold and the diversity of the data set is lower. Following this rationale, the CSR curves in Fig. 7 indicate that the scaffold diversity of the epigenetic and GRAS data sets decreases in the order: DNMTs > BRDs > HDACs > GRAS. Interestingly, according to the AUC metric, the scaffold diversity of the BRDs and HDACs (AUC = 0.74 and 0.76, respectively) is comparable to the diversity of compound tested as inhibitors of the androgen receptor, estrogen receptor agonist, glucocorticoid receptor, angiotensin-converting enzyme, and acethylcholine esterase that have reported AUC values between 0.74 and 0.76.43 The scaffold diversity of the DNMT set is similar to compounds tested with enoyl-(acyl-carrier-protein) reductase (AUC of 0.69 and 0.70, respectively).43
The same relative order of scaffold diversity was obtained with the F50 values (Table S8 in the ESI†). Half (50%) of the GRAS compounds were contained in 0.4% of the cyclic systems. In contrast, 50% of the compounds in the DNMTs set were distributed in 22% of the cyclic systems of this set. The F50 metric also indicated that the relative order of scaffold diversity of the epigenetic-related data sets is DNMT > BRDs > HDACs. The lower cyclic system diversity of the HDACs set can be influenced by research trends e.g., hydroxamic acid derivatives.
It is noteworthy that the relative order of scaffold diversity does not necessarily match the scaffold diversity measured with structural fingerprints. As discussed elsewhere, the structural diversity evaluated with fingerprints consider the entire structure, including the own nature of the cyclic systems (e.g., size, complexity) and the side chains. A clear example is the relative diversity of the GRAS set: it has a high structural diversity (MACCS keys/Tanimoto in Fig. 4) but it has lower scaffold diversity (CSR curve in Fig. 7) as compared to other data sets. Of note, this is the first study that addresses the scaffold diversity of GRAS.
3.4 Molecular complexity
The search for diverse datasets is usually made with the purpose of exploring ‘empty voids’ of chemical space. The universe of possible existing molecules is huge so the idea of entering uncharted regions may seem easy. What is more challenging is the successful navigation through novel but pharmaceutical relevant areas of chemical space.61 Specific strategies to venture further in this chemical wilderness have been discussed recently. One of such strategies depends on assessing molecular complexity. To the best of our knowledge, there are not reports that measure the molecular complexity of epigenetic data sets.Fig. 8 shows the distribution of the fraction of chiral and sp3 carbon atoms for the BRDs, HDACs, DNMTs and other reference data sets. Summary statistics are presented in Table S9 in the ESI.† The distributions were compared with Nemenyi tests for pairing and analysis of raw data. Analysis of the results revealed that HDACs and ‘Epi-focused’ have similar distribution of F-sp3 values suggesting that it is likely equal to find 3D structures (less flat compounds) in both data sets. The overall lower F-sp3 values of BRDs than HDACs (and other reference sets, except DNMTs), indicate that the structures of BRDs currently contained in ChEMBL are, in general, more flat.
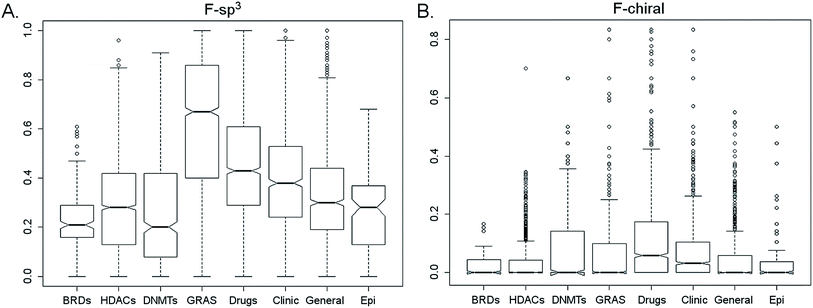 | ||
| Fig. 8 Box plots of the distribution of the fraction of sp3 carbon atoms (F-sp3) and fraction of chiral centers (F-chiral) for the BRDs, HDACs, DNMTs and reference data sets. Summary statistics are in Table S9 of the ESI.† | ||
BRDs and HDACs had comparable distributions of F-chiral values indicating similar stereochemical complexity. Moreover, their distribution of F-chiral values was also comparable to ‘General’ and ‘Epi-focused’. DNMTs showed broader range of F-chiral values.
In agreement with previous analyses, ‘Drugs’ had overall, higher F-chiral and F-sp3 values than commercial screening library ‘General’. Notably, GRAS compounds had even higher F-sp3 values than ‘Drugs’. These results suggest that it is more likely that GRAS compounds have 3D structures as compared to approved drugs and any other data sets analyzed in this work. Indeed, it is known that the activity of several approved drugs are stereospecific as only one enantiomer is active, while the other may not be active or even toxic. GRAS on the other hand, contains flavor molecules: many sugars and sweeteners are optically active. Also flavors may show “property cliffs”, that is, flavor may drastically change with small structure changes (i.e., limonene gives both their flavors to lemons and tangerines, the difference is just a chiral center).62
Taken together, the results of this analysis suggested that the compounds currently tested as inhibitors of BRDs, HDACs and DNMTs have comparable or less stereochemical complexity, and are less flat than currently approved drugs.
4. Conclusions
Epigenetics is an emerging therapeutic approach and has proven to be a very important aspect of our daily lives.63,64 As such, it is quite relevant to understand the chemical space currently covered. As part of our research program to further advance epi-drug and epi-drug discovery using computational methods,17,65 herein we discuss for the first time the characterization of the chemical space of BRDis and HDACis using structure–activity data available in two major public databases: ChEMBL and Binding Database. It was concluded that the chemical space of data sets with activity against HDACs and BRDs showed a significant overlap with the property space of drugs approved for clinical use, compounds in clinical development and commercial screening libraries. Complementary measures of structural diversity revealed that there is a need to develop and test more diverse compounds as BRDis.Based on the diversity analysis of the screening data analyzed of HDACis, BRDis and DNMTis it was found that the development of poly-epigenetic drugs has not been extensively explored. However, it was shown the feasibility of develop at least dual epigenetic inhibitors. This conclusion has been supported experimentally. These observations open up a unique avenue to develop potentially more efficient epigenetic therapies of compounds targeting multiple epigenetic targets.
Surprisingly, despite the fact that several different chemical scaffolds have been explored for BRDs and HDACs, there is not yet a unique molecular scaffold (as defined by Johnson and Xu) with a large enrichment factor. These quantitative results highlight the need to continue increasing the SAR of the most promising chemical scaffolds identified in this work.
From the quantitative analysis of the structural complexity it was concluded that, in general, the chemical structure of inhibitors of BRDs, HDACs, and DNMT have a limited structural complexity. These results encourage the development of new inhibitors with increased complexity that may lead to improved selectivity. Taking all this together, this work represents a significant contribution to further advance the understanding of the ERCS.
5. Perspectives
The first step in our ERCS odyssey was to analyze compounds targeted to several different types of BRDs (2, 3, 4) and HDACs (1, 2, 3, 5, 8 and SIRT1). The next logical step is to assess the fraction of ERCS of different BRDs and HDACs individually and expand the analysis to other public and more specialized repositories such as Chromohub.66 As part of this analysis, an in-depth characterization of the overlap between ChEMBL/Binding Database with Chromohub can be made. Furthermore, it will be of high relevance to explore the chemical space of compounds with and without cellular activity, as well as to determine structure–selectivity relationships of HDACs, BRDs, and other epigenetic modulators.During the course of this study it was found that GRAS compounds share a similar property space as the ERCS region; in particular, share similar hydrophobicity values which is one of the most important PCP related to bioactive compounds. These novel results support the systematic exploration of flavor chemicals with potential health benefits as epigenetic modulators. Furthermore, the chemoinformatic study uncovered that GRAS chemicals have a larger number of 3D (less flat) structures as compared to other general screening and ‘Epi-focused’ commercial collections. It is anticipated that GRAS chemicals may show selectivity towards epigenetic targets and may act as ‘master key’ epigenetic-compounds. Further experimental studies are required to assess this hypothesis.
List of abbreviations
| BRDs | Bromodomains |
| DNA | Deoxyribonucleic acid |
| DNMT | DNA-methyl transferase |
| EF | Enrichment factor |
| ERCS | Epigenetic relevant chemical space |
| GRAS | Generally recognized as safe |
| HDACs | Histone lysine deacetylase |
| HBA | Hydrogen bond acceptor |
| HBD | Hydrogen bond donor |
| RTB | Number of rotatable bonds |
Slog![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P P | Partition coefficient water/octanol |
| MW | Molecular weight |
| PCP | Physicochemical properties |
| PC | Principal component |
| PCA | Principal component analysis |
| SMILES | Simplified molecular input line entry |
| SAM | S-Adenosyl-methionine |
| TPSA | Topological surface area |
Acknowledgements
Fernando Prieto-Martínez and Eli Fernández-de Gortari are grateful to CONACyT for the fellowships granted No. 660465/576637 and 348291/240072, respectively. Authors thank Tatiana Enríquez Gómez and Jonathan Guerra Pérez for carefully proofreading the manuscript. This work was supported by the Universidad Nacional Autónoma de México (UNAM), grant PAPIIT IA204016. We also thank the institutional program Nuevas Alternativas de Tratamiento para Enfermedades Infecciosas (NUATEI) of the Instituto de Investigaciones Biomédicas (IIB) UNAM and Programa de Apoyo a la Investigación y el Posgrado (PAIP) 5000-9163, Facultad de Química, UNAM, for financial support.References
- D. C. Dolinoy, in Comprehensive Toxicology, ed. C. A. McQueen, Elsevier, Oxford, 2nd edn, 2010, pp. 293–309 Search PubMed.
- J. C. Jiménez-Chillarón, R. Díaz, M. Ramón-Krauel and S. Ribó, in Transgenerational Epigenetics, ed. T. Tollefsbol, Academic Press, Oxford, 2014, pp. 281–301 Search PubMed.
- N. Carey, MedChemComm, 2012, 3, 162–166 RSC.
- K. P. Nightingale, in Epigenetics for Drug Discovery, The Royal Society of Chemistry, 2016, pp. 1–19 Search PubMed.
- Q. Gao, J. Tang, J. Chen, L. Jiang, X. Zhu and Z. Xu, Drug Discovery Today, 2014, 19, 1744–1750 CrossRef CAS PubMed.
- A. O. Arguelles, S. Meruvu, J. D. Bowman and M. Choudhury, Drug Discovery Today, 2016, 21, 499–509 CrossRef PubMed.
- A. Tarakhovsky, Nat. Immunol., 2010, 11, 565–568 CrossRef CAS PubMed.
- C. Robert and F. V. Rassool, in Advances in Cancer Research, ed. G. Steven, Academic Press, 2012, vol. 116, pp. 87–129 Search PubMed.
- S. Majumder and A. Advani, Journal of Diabetes and its Complications, 2015, 29, 1337–1344 CrossRef PubMed.
- F. L. Cherblanc, R. W. M. Davidson, P. Di Fruscia, N. Srimongkolpithak and M. J. Fuchter, Nat. Prod. Rep., 2013, 30, 605–624 RSC.
- B. Pachaiyappan and P. M. Woster, Bioorg. Med. Chem. Lett., 2014, 24, 21–32 CrossRef CAS PubMed.
- C.-Y. Wang and P. Filippakopoulos, Trends Biochem. Sci., 2015, 40, 468–479 CrossRef CAS PubMed.
- S.-Y. Wu and C.-M. Chiang, J. Biol. Chem., 2007, 282, 13141–13145 CrossRef CAS PubMed.
- C. W. Lindsley, ACS Chem. Neurosci., 2014, 5, 1142 CrossRef CAS PubMed.
- J. L. Medina-Franco, M. A. Giulianotti, G. S. Welmaker and R. A. Houghten, Drug Discovery Today, 2013, 18, 495–501 CrossRef PubMed.
- G. Franci, M. Miceli and L. Altucci, Epigenomics, 2010, 2, 731–742 CrossRef CAS PubMed.
- O. Méndez-Lucio, J. Tran, J. L. Medina-Franco, N. Meurice and M. Muller, ChemMedChem, 2014, 9, 560–565 CrossRef PubMed.
- J. L. Medina-Franco, K. Martinez-Mayorga and N. Meurice, Expert Opin. Drug Discovery, 2014, 9, 151–165 CrossRef CAS PubMed.
- K. Martinez-Mayorga, T. L. Peppard, F. López-Vallejo, A. B. Yongye and J. L. Medina-Franco, J. Agric. Food Chem., 2013, 61, 7507–7514 CrossRef CAS PubMed.
- E. Fernández-de Gortari and J. L. Medina-Franco, RSC Adv., 2015, 5, 87465–87476 RSC.
- J. L. Medina-Franco, Epi-Informatics: Discovery and Development of Small Molecule Epigenetic Drugs and Probes, Academic Press, London, UK, 2016 Search PubMed.
- X. Lucas, B. A. Grüning, S. Bleher and S. Günther, J. Chem. Inf. Model., 2015, 55, 915–924 CrossRef CAS PubMed.
- J. B. Baell, J. Chem. Inf. Model., 2013, 53, 39–55 CrossRef CAS PubMed.
- G. Papadatos and J. P. Overington, Future Med. Chem., 2014, 6, 361–364 CrossRef CAS PubMed.
- T. Q. Liu, Y. M. Lin, X. Wen, R. N. Jorissen and M. K. Gilson, Nucleic Acids Res., 2007, 35, D198–D201 CrossRef CAS PubMed.
- C. H. Arrowsmith, C. Bountra, P. V. Fish, K. Lee and M. Schapira, Nat. Rev. Drug Discovery, 2012, 11, 384–400 CrossRef CAS PubMed.
- M. Wegner, D. Neddermann, M. Piorunska-Stolzmann and P. P. Jagodzinski, Diabetes Res. Clin. Pract., 2014, 105, 164–175 CrossRef CAS PubMed.
- Molecular Operating Environment (MOE), version 2013, Chemical Computing Group Inc., Montreal, Quebec, Canada, available at http://www.chemcomp.com Search PubMed.
- D. Fourches, E. Muratov and A. Tropsha, Nat. Chem. Biol., 2015, 11, 535 CrossRef CAS PubMed.
- J. L. Medina-Franco, K. Martínez-Mayorga, T. L. Peppard and A. Del Rio, PLoS One, 2012, 7, e50798 CAS.
- F. Lovering, J. Bikker and C. Humblet, J. Med. Chem., 2009, 52, 6752–6756 CrossRef CAS PubMed.
- R Development Core Team, R Foundation for Statistical Computing, Vienna, Austria, 2011 Search PubMed.
- T. Sander, J. Freyss, M. von Korff and C. Rufener, J. Chem. Inf. Model., 2015, 55, 460–473 CrossRef CAS PubMed.
- P. Jaccard, Bull. Soc. Vaudoise Sci. Nat., 1901, 37, 547–579 Search PubMed.
- J. L. Medina-Franco and G. M. Maggiora, in Chemoinformatics for Drug Discovery, ed. J. Bajorath, John Wiley & Sons, Inc., 2014, ch. 15, pp. 343–399 Search PubMed.
- J. L. Medina-Franco, G. M. Maggiora, M. A. Giulianotti, C. Pinilla and R. A. Houghten, Chem. Biol. Drug Des., 2007, 70, 393–412 CAS.
- M. P. scripts, Matplotlib Python scripts, available at http://dx.doi.org/10.5281/zenodo.15423.
- Y. J. Xu and M. Johnson, J. Chem. Inf. Comput. Sci., 2002, 42, 912–926 CrossRef CAS PubMed.
- Y. Xu and M. Johnson, J. Chem. Inf. Comput. Sci., 2001, 41, 181–185 CrossRef CAS PubMed.
- F. López-Vallejo, A. Nefzi, A. Bender, J. R. Owen, I. T. Nabney, R. A. Houghten and J. L. Medina-Franco, Chem. Biol. Drug Des., 2011, 77, 328–342 Search PubMed.
- A. B. Yongye, J. Waddell and J. L. Medina-Franco, Chem. Biol. Drug Des., 2012, 80, 717–724 CAS.
- A. B. Yongye and J. L. Medina-Franco, Chem. Biol. Drug Des., 2013, 82, 367–375 CAS.
- J. L. Medina-Franco, K. Martínez-Mayorga, A. Bender and T. Scior, QSAR Comb. Sci., 2009, 28, 1551–1560 CAS.
- A. H. Lipkus, Q. Yuan, K. A. Lucas, S. A. Funk, W. F. Bartelt, R. J. Schenck and A. J. Trippe, J. Org. Chem., 2008, 73, 4443–4451 CrossRef CAS PubMed.
- J. L. Medina-Franco, J. Petit and G. M. Maggiora, Chem. Biol. Drug Des., 2006, 67, 395–408 CAS.
- J. Pérez-Villanueva, O. Méndez-Lucio, O. Soria-Arteche and J. Medina-Franco, Mol. Diversity, 2015, 19, 1021–1035 CrossRef PubMed.
- J. Pérez-Villanueva, J. L. Medina-Franco, O. Méndez-Lucio, J. Yoo, O. Soria-Arteche, T. Izquierdo, M. C. Lozada and R. Castillo, Chem. Biol. Drug Des., 2012, 80, 752–762 Search PubMed.
- F. Lovering, MedChemComm, 2013, 4, 515–519 RSC.
- F. López-Vallejo, M. A. Giulianotti, R. A. Houghten and J. L. Medina-Franco, Drug Discovery Today, 2012, 17, 718–726 CrossRef PubMed.
- S. H. Bertz, J. Am. Chem. Soc., 1981, 103, 3599–3601 CrossRef CAS.
- R. Barone and M. Chanon, J. Chem. Inf. Comput. Sci., 2001, 41, 269–272 CrossRef CAS PubMed.
- H. Chen, O. Engkvist, N. Blomberg and J. Li, MedChemComm, 2012, 3, 312–321 RSC.
- A. Ganesan, Curr. Opin. Chem. Biol., 2008, 12, 306–317 CrossRef CAS PubMed.
- N. Singh, R. Guha, M. A. Giulianotti, C. Pinilla, R. A. Houghten and J. L. Medina-Franco, J. Chem. Inf. Model., 2009, 49, 1010–1024 CrossRef CAS PubMed.
- Y. Chen, H. Li, W. Tang, C. Zhu, Y. Jiang, J. Zou, Q. Yu and Q. You, Eur. J. Med. Chem., 2009, 44, 2868–2876 CrossRef CAS PubMed.
- A. M. Wassermann, E. Lounkine, J. W. Davies, M. Glick and L. M. Camargo, Drug Discovery Today, 2015, 20, 422–434 CrossRef PubMed.
- S. J. Atkinson, P. E. Soden, D. C. Angell, M. Bantscheff, C.-w. Chung, K. A. Giblin, N. Smithers, R. C. Furze, L. Gordon, G. Drewes, I. Rioja, J. Witherington, N. J. Parr and R. K. Prinjha, MedChemComm, 2014, 5, 342–351 RSC.
- S. R. Langdon, N. Brown and J. Blagg, J. Chem. Inf. Model., 2011, 51, 2174–2185 CrossRef CAS PubMed.
- Y. Hu and J. Bajorath, J. Chem. Inf. Model., 2010, 50, 500–510 CrossRef CAS PubMed.
- J. Baell and M. A. Walters, Nature, 2014, 513, 481–483 CrossRef CAS PubMed.
- T. I. Oprea and J. Gottfries, J. Comb. Chem., 2001, 3, 157–166 CrossRef CAS PubMed.
- J. L. Medina-Franco, G. Navarrete-Vázquez and O. Méndez-Lucio, Future Med. Chem., 2015, 7, 1197–1211 CrossRef CAS PubMed.
- I. R. Miousse, R. Currie, K. Datta, H. Ellinger-Ziegelbauer, J. E. French, A. H. Harrill, I. Koturbash, M. Lawton, D. Mann, R. R. Meehan, J. G. Moggs, R. O'Lone, R. J. Rasoulpour, R. A. R. Pera and K. Thompson, Toxicology, 2015, 335, 11–19 CrossRef CAS PubMed.
- M. Szyf, Eur. Neuropsychopharmacol., 2015, 25, 682–702 CrossRef CAS PubMed.
- J. L. Medina-Franco, O. Méndez-Lucio, J. Yoo and A. Dueñas, Drug Discovery Today, 2015, 20, 569–577 CrossRef CAS PubMed.
- L. Liu, X. T. Zhen, E. Denton, B. D. Marsden and M. Schapira, Bioinformatics, 2012, 28, 2205–2206 CrossRef CAS PubMed.
Footnote |
| † Electronic supplementary information (ESI) available. See DOI: 10.1039/c6ra07224k |
| This journal is © The Royal Society of Chemistry 2016 |