The effect of POPC acyl chains packing by aromatic amino acid methyl esters investigated by ATR-FTIR combined with QM calculations†
Bo Penga,
Xiao-Yan Dinga,
Chao Suna,
Wei Liub,
John Z. H. Zhangbc and
Xin Zhao*a
aShanghai Key Laboratory of Magnetic Resonance, Department of Physics, School of Physics and Materials Science, East China Normal University, Shanghai 200062, P. R. China. E-mail: xzhao@phy.ecnu.edu.cn; Fax: +86 21 62234329; Tel: +86 21 62234329
bState Key Laboratory of Precision Spectroscopy, Department of Physics, School of Physics and Materials Science, East China Normal University, Shanghai 200062, P. R. China
cNYU-ECNU Center for Computational Chemistry at NYU Shanghai, Shanghai 200062, P. R. China
First published on 5th May 2016
Abstract
Aromatic residues play important roles in modulating the function of proteins and binding of proteins to biomembranes. However, the effect of small molecules which contain aromatic residues on the packing of unsaturated lipid acyl chains and the dynamics properties in membrane bilayers remains unclear. The membrane interactions of a series of amino acid methyl esters have been investigated to establish how different amino acids influence lipid acyl chains packing. Attenuated total reflectance-Fourier transform infrared spectroscopic (ATR-FTIR) measurements show that the ![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH and C
CH and C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C stretching vibrational bands of 2-oleoyl-1-pamlitoyl-sn-glyecro-3-phosphocholine (POPC) present evident red-shifts when aromatic amino acid methyl esters are mixed into membrane bilayers. Quantum mechanical (QM) calculations further demonstrate that the red shifts of the
C stretching vibrational bands of 2-oleoyl-1-pamlitoyl-sn-glyecro-3-phosphocholine (POPC) present evident red-shifts when aromatic amino acid methyl esters are mixed into membrane bilayers. Quantum mechanical (QM) calculations further demonstrate that the red shifts of the ![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH and C
CH and C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C stretching modes for POPC acyl chains are not through a direct interaction between the side chain groups and the HC
C stretching modes for POPC acyl chains are not through a direct interaction between the side chain groups and the HC![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH motif but are mainly caused by the conformational change of POPC acyl chains, in which the HC
CH motif but are mainly caused by the conformational change of POPC acyl chains, in which the HC![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH motif is packed closed to the other one of an adjacent acyl chain with enhancement by dispersion interactions. The implication of these findings is that the packing of POPC acyl chains can be significantly influenced by aromatic amino acid methyl esters. This work may provide valuable insight into the effect on the lipid acyl chains packing by small molecules interacting with membranes and offer possibilities for the dynamic control of membrane fusion and binding of proteins to the membrane.
CH motif is packed closed to the other one of an adjacent acyl chain with enhancement by dispersion interactions. The implication of these findings is that the packing of POPC acyl chains can be significantly influenced by aromatic amino acid methyl esters. This work may provide valuable insight into the effect on the lipid acyl chains packing by small molecules interacting with membranes and offer possibilities for the dynamic control of membrane fusion and binding of proteins to the membrane.
1. Introduction
Biological membranes are involved in many essential processes, including biomolecule transportation, distribution, signal transduction, cell recognition, enzymatic reactions, membrane protein structure and function stabilization, etc.1–3 Phospholipids provide structural support for membranes and are responsible for semi-permeable and stability properties of the membranes. Lipid bilayers are formed by phospholipids in two layers, with polar headgroups along the two layers and the lipid acyl chains forming the apolar domain in between as the most common arrangement.4 Lipid bilayers form the underlying structure of every animal and plant cell membrane. The special structure of the cell membrane provides a relatively stable internal environment for a cell. Changes of lipid bilayer dynamics and structure play crucial roles in many membrane-associated processes, and corresponding factors influencing the structure and flexibility of lipid bilayers are various, such as temperature,5 biomolecule interactions with the membrane6 and lipid composition7 etc. Thus, it seems to be very important to study the stability of phospholipid bilayers and the perturbation of membranes by other molecules.Biological membranes are composed of various phospholipids with different polar head groups, degrees of unsaturation and acyl chain length. The degree of unsaturation is determined by the number of double bonds in lipid tails. With the unsaturated double bonds presenting in the lipid acyl chains, the thermodynamic properties of membrane may change dramatically. The fluidity of unsaturated lipids increases, and the gel-phase transition temperature is lower than normal temperatures.8,9 Lipid components include glycerophospholipids, sterols and sphingolipids. The glycerophospholipids is the most abundant lipid species in biological membranes, such as phosphatidylcholine (PC), phosphatidylethanolamine (PE), phosphatidylserine (PS), phosphatidylglycerol (PG), phosphatidylinositol (PI) and phosphatidic acid (PA).7 The PC family has been extensively studied as the model membrane system,10 especially for 2-oleoyl-1-pamlitoyl-sn-glyecro-3-phosphocholine (POPC) is a monounsaturated lipid.
Aromatic residues play vital roles in the function of protein and binding of protein to biomembranes. Sequence motifs that contain aromatic residues widely occur in transmembrane,11,12 peripheral,13,14 and membrane-associated proteins.15,16 However, the effects of aromatic residues on the membrane composed of unsaturated phospholipids are of particular specific and complex. On the basis of fluorescence emission spectra, Williams et al. reported that tryptophan in the juxtamembrane domain of vesicle-associated membrane protein 2 (VAMP2), which can bind to the adjacent lipid bilayers in insulin granules or other transport vesicles, may insert into the hydrophobic interior of PC or 3![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 PC/PS bilayers.17 Merlino et al. suggested that the octapeptide (C8), which contains three tryptophan residues, binds to the POPC bilayers surface and decreases the bilayer thickness. This discovery was supported by neutron reflectivity, electron spin resonance, circular dichroism, fluorescence spectroscopy and molecular dynamics simulations.18 Using fluorescence correlation spectroscopy, high resolution 31P field cycling NMR spectroscopy and X-ray crystallography, Cheng et al. found that the choline cation–aromatic π interactions may be a specific motif to anchor proteins to PC-rich bilayers.13 The findings discussed above suggest that the packing of unsaturated lipid acyl chains underlying the interaction of the motif which contains aromatic residue penetration in membrane bilayers remains poorly understood. Thus, it seems to be important to elucidate the effect on the packing of unsaturated lipid acyl chains in membrane bilayers by motif which contains aromatic residue interaction with lipids in the model membrane.
1 PC/PS bilayers.17 Merlino et al. suggested that the octapeptide (C8), which contains three tryptophan residues, binds to the POPC bilayers surface and decreases the bilayer thickness. This discovery was supported by neutron reflectivity, electron spin resonance, circular dichroism, fluorescence spectroscopy and molecular dynamics simulations.18 Using fluorescence correlation spectroscopy, high resolution 31P field cycling NMR spectroscopy and X-ray crystallography, Cheng et al. found that the choline cation–aromatic π interactions may be a specific motif to anchor proteins to PC-rich bilayers.13 The findings discussed above suggest that the packing of unsaturated lipid acyl chains underlying the interaction of the motif which contains aromatic residue penetration in membrane bilayers remains poorly understood. Thus, it seems to be important to elucidate the effect on the packing of unsaturated lipid acyl chains in membrane bilayers by motif which contains aromatic residue interaction with lipids in the model membrane.
Fourier transform infrared (FTIR) spectroscopy is increasingly used to analyse the conformational order of lipid19 and organization of lipid membranes in different phases,5,20 to study the structure21 and conformation22 of proteins and to investigate interaction mechanisms between small molecules and lipid membranes.6 Attenuated total reflection (ATR)-FTIR is a powerful tool to study the structure, orientation and tertiary structure changes in peptides and membrane proteins in biologically relevant model membrane bilayers conveniently.23 This sensitive technique requires only a very small sample without the need to introduce an additional probe. Besides, ATR-FTIR spectroscopy with polarisation can be used to probe the different orientation of proteins in cell membranes24 and orientation of membrane molecules.25 In the current study, we have sought to gain an insight into the effect of the motif which contains aromatic residues mixing into membrane bilayers on unsaturated lipid acyl chains packing and membrane structure. As the membrane environment is extraordinarily complex, in order to simplify the system, we used polarised ATR-FTIR spectroscopy to systematically investigate the effect of a series of designed amino acid methyl esters containing small hydrophobic and aromatic residues on the POPC membrane bilayers packing. In addition, we have used quantum mechanical (QM) calculations to present more information of the possible interaction modes of aromatic rings/HC![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH motif with HC
CH motif with HC![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH motif directly at the molecular level.
CH motif directly at the molecular level.
2. Methods
2.1. Experimental
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 10. The hydrated pellets were then lyophilised, rehydrated with D2O to 50% (w/v) and incubated at 37 °C for 2 days.
10. The hydrated pellets were then lyophilised, rehydrated with D2O to 50% (w/v) and incubated at 37 °C for 2 days.2.2. Calculations
As the water molecules can not penetrate into the interior of POPC bilayers in the gel and liquid-crystalline state,26 the salt ions also have negligible effect on the structure of POPC bilayer,27 so the QM calculations were performed using the Gaussian 09 computational chemistry program28 in the gas phase. The geometry optimisation and frequency calculation were performed at the ωB97X-D/6-31G (d,p) level29 for POPC–side chain group complexes. Model systems contained only a 3-hexene chain was used instead of a full length lipid molecule, as well as the side chain groups of amino acid methyl esters. The geometry optimisation and frequency calculation were also performed at the Møller–Plesset second-order perturbation theory (MP2)/6-31G (d,p) level30 for model systems and were performed using the “opt = modredundant” command after freezing and increasing the distance of the C9 and C27 of interest two 3-hexene motif from 3.27 Å to 3.87 Å every 0.2 Å. The factor of 0.950 for ωB97X-D/6-31G (d,p) and 0.933 for MP2/6-31G (d,p) were used to scale the vibrational frequencies to correct for the harmonic approximation, respectively.31 Normal modes and IR spectra were displayed using the GaussView program.3. Results and discussion
3.1. FTIR spectra of POPC bilayers and POPC bilayer–amino acid methyl ester complexes
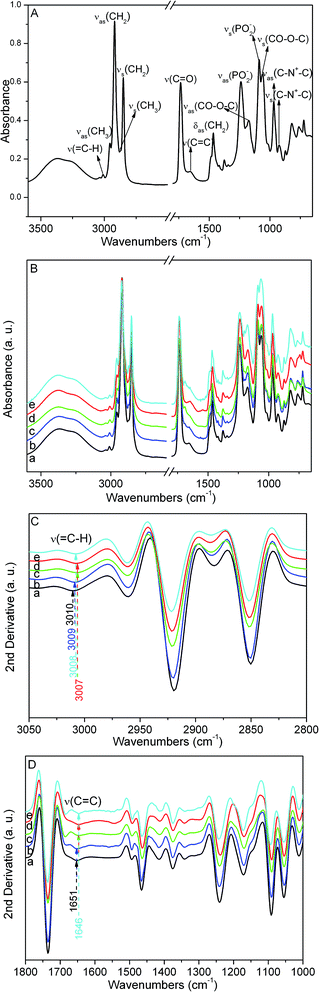
A typical infrared spectrum for POPC bilayers and FTIR spectra of POPC bilayer–amino acid methyl ester complexes in the 3600–650 cm−1 region are presented in Fig. 2A and B, respectively. Most of the vibrational peaks could be unambiguously assigned,32 as listed in Table S1 in the ESI.† For POPC bilayers, the band of the![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH stretching mode at 3010 cm−1 and the C
CH stretching mode at 3010 cm−1 and the C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C stretching mode at 1651 cm−1 can be clearly identified. For POPC bilayer–amino acid methyl ester complexes, the vibrational frequencies of CH2 antisymmetric and symmetric stretching modes show blue shifts to some extent. Besides, the vibrational frequencies of
C stretching mode at 1651 cm−1 can be clearly identified. For POPC bilayer–amino acid methyl ester complexes, the vibrational frequencies of CH2 antisymmetric and symmetric stretching modes show blue shifts to some extent. Besides, the vibrational frequencies of ![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH and C
CH and C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C stretching modes show evident changes in POPC bilayer–amino acid methyl ester complexes. The second-derivative FTIR spectra of POPC bilayers and POPC bilayer–amino acid methyl ester complexes in the
C stretching modes show evident changes in POPC bilayer–amino acid methyl ester complexes. The second-derivative FTIR spectra of POPC bilayers and POPC bilayer–amino acid methyl ester complexes in the ![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH stretching region are presented in Fig. 2C. An obvious red shift of 3 cm−1 for the
CH stretching region are presented in Fig. 2C. An obvious red shift of 3 cm−1 for the ![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH stretching band in POPC acyl chains was observed when tryptophan methyl ester (Trp-OMe) and phenylalanine methyl ester (Phe-OMe) were added into pure POPC bilayers, and a red shift of 2 cm−1 was observed when tyrosine methyl ester (Tyr-OMe) was added. However, there was only a small red shift of 1 cm−1 when leucine methyl ester (Leu-OMe) was mixed. Furthermore, the second-derivative FTIR spectra in the C
CH stretching band in POPC acyl chains was observed when tryptophan methyl ester (Trp-OMe) and phenylalanine methyl ester (Phe-OMe) were added into pure POPC bilayers, and a red shift of 2 cm−1 was observed when tyrosine methyl ester (Tyr-OMe) was added. However, there was only a small red shift of 1 cm−1 when leucine methyl ester (Leu-OMe) was mixed. Furthermore, the second-derivative FTIR spectra in the C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C stretching region are presented in Fig. 2D. The C
C stretching region are presented in Fig. 2D. The C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C stretching band of POPC bilayers was shifted from 1651 cm−1 to 1646 cm−1 (a red shift of 5 cm−1) when Trp-OMe, Phe-OMe and Tyr-OMe were added, respectively. However, a red shift of 0 cm−1 was observed when Leu-OMe was mixed, which shows almost no change in the C
C stretching band of POPC bilayers was shifted from 1651 cm−1 to 1646 cm−1 (a red shift of 5 cm−1) when Trp-OMe, Phe-OMe and Tyr-OMe were added, respectively. However, a red shift of 0 cm−1 was observed when Leu-OMe was mixed, which shows almost no change in the C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C stretching frequency of POPC bilayers. In addition, the PO2− antisymmetric stretching band at 1241 cm−1 and PO2− symmetric stretching band at 1091 cm−1 show different variations when different amino acid methyl esters were added, and the C–N+–C antisymmetric stretching band at 968 cm−1 shows a small red shift of 1 cm−1 when four amino acid methyl esters were added. These bands are mainly associated with the interface region of POPC membrane. In this paper, we aim at investigating the effect of POPC acyl chains packing. Therefore, alterations in the frequency values of
C stretching frequency of POPC bilayers. In addition, the PO2− antisymmetric stretching band at 1241 cm−1 and PO2− symmetric stretching band at 1091 cm−1 show different variations when different amino acid methyl esters were added, and the C–N+–C antisymmetric stretching band at 968 cm−1 shows a small red shift of 1 cm−1 when four amino acid methyl esters were added. These bands are mainly associated with the interface region of POPC membrane. In this paper, we aim at investigating the effect of POPC acyl chains packing. Therefore, alterations in the frequency values of ![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH and C
CH and C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C stretching vibrations of POPC can be used to probe the effect of amino acid methyl esters on the POPC acyl chains packing when they were mixed into the membrane bilayers.
C stretching vibrations of POPC can be used to probe the effect of amino acid methyl esters on the POPC acyl chains packing when they were mixed into the membrane bilayers.
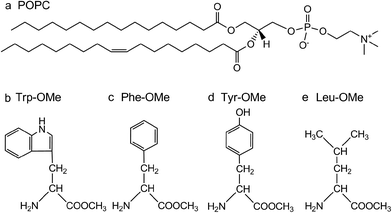
The chemical structures of POPC and amino acid methyl esters are shown in Scheme 1. POPC has one glycerol, one phosphate, two carbonyl groups and two acyl chains, one of which contains a C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C bond in the centre. Trp-OMe, Phe-OMe, Tyr-OMe and Leu-OMe have the same NH2 and COOCH3 groups, and different side chain groups. Among these four amino acid methyl esters, the experimental results show that the bathochromic shifts of the
C bond in the centre. Trp-OMe, Phe-OMe, Tyr-OMe and Leu-OMe have the same NH2 and COOCH3 groups, and different side chain groups. Among these four amino acid methyl esters, the experimental results show that the bathochromic shifts of the ![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH and C
CH and C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C stretching bands of POPC acyl chains are obvious for POPC bilayer complexes with Trp-OMe, Phe-OMe and Tyr-OMe. A very small red shift of
C stretching bands of POPC acyl chains are obvious for POPC bilayer complexes with Trp-OMe, Phe-OMe and Tyr-OMe. A very small red shift of ![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH stretching band was observed when Leu-OMe was mixed in POPC bilayers. According to White et al.'s work,33 the free energy of partition of Trp, Phe, Tyr and Leu from water to POPC interface are around −1.85, −1.13, −0.94 and −0.56 kcal mol−1, respectively. It indicates that the location process of three aromatic amino acids which have the highly hydrophobic aromatic groups is more favored energetically than Leu in interface scale.
CH stretching band was observed when Leu-OMe was mixed in POPC bilayers. According to White et al.'s work,33 the free energy of partition of Trp, Phe, Tyr and Leu from water to POPC interface are around −1.85, −1.13, −0.94 and −0.56 kcal mol−1, respectively. It indicates that the location process of three aromatic amino acids which have the highly hydrophobic aromatic groups is more favored energetically than Leu in interface scale.
Each band in an infrared spectrum has its defining characteristic, denoted by its frequency, peak width at half height, and absorption maximum. When the frequency of a vibrational peak changes, the force constant of corresponding group changes based on the harmonic oscillator model, indicating that the corresponding microenvironment of the group also changes. The bathochromic shifts of the C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C and
C and ![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH stretching vibrational bands may be associated with several possible mechanism, such as the direct interactions between the side chain groups of amino acid methyl esters and the C
CH stretching vibrational bands may be associated with several possible mechanism, such as the direct interactions between the side chain groups of amino acid methyl esters and the C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C bond of the POPC acyl chains, or the packing and conformational change of POPC acyl chains.
C bond of the POPC acyl chains, or the packing and conformational change of POPC acyl chains.
3.2. QM calculation results of POPC, POPC–side chain group complexes, 3-hexene and 3-hexene–side chain group complexes
Our experimental results deduce that the red shifts of![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH and C
CH and C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C stretching bands in HC
C stretching bands in HC![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH motif of POPC acyl chain may associate with the mixing of amino acid methyl esters into the lipids membrane. As the physicochemical properties of lipid membranes are complex, and the configuration and property of amino acid clusters differ in different amino acid methyl esters. Trp has a side chain with a large indole ring, which can interact with other molecules through π–π interactions34 or dipolar interactions35 because of its aromatic character. Furthermore, although it is a very hydrophobic residue, Trp is also involved in hydrogen bonding interactions36 and cation–π interactions11 in numerous phenomena in structural biology. Phe has a hydrophobic benzene ring structure, which can also be involved in interactions with other aromatic groups via stacking interactions37 and π–π interactions.34 In addition, the benzene ring can interact with hydrogen bond donors.38 Tyr has a phenol ring, which can also be involved in hydrogen bonding interactions39 and aromatic π–π interactions.40 Leu has a hydrophobic aliphatic side chain that is sensitive to helix packing interactions in peptides that oligomerise.41 To obtain more detailed information about the phenomena that side chain groups interact with the HC
CH motif of POPC acyl chain may associate with the mixing of amino acid methyl esters into the lipids membrane. As the physicochemical properties of lipid membranes are complex, and the configuration and property of amino acid clusters differ in different amino acid methyl esters. Trp has a side chain with a large indole ring, which can interact with other molecules through π–π interactions34 or dipolar interactions35 because of its aromatic character. Furthermore, although it is a very hydrophobic residue, Trp is also involved in hydrogen bonding interactions36 and cation–π interactions11 in numerous phenomena in structural biology. Phe has a hydrophobic benzene ring structure, which can also be involved in interactions with other aromatic groups via stacking interactions37 and π–π interactions.34 In addition, the benzene ring can interact with hydrogen bond donors.38 Tyr has a phenol ring, which can also be involved in hydrogen bonding interactions39 and aromatic π–π interactions.40 Leu has a hydrophobic aliphatic side chain that is sensitive to helix packing interactions in peptides that oligomerise.41 To obtain more detailed information about the phenomena that side chain groups interact with the HC![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH motif directly, QM calculations were performed not only in POPC–side chain group complexes but also in truncated systems of 3-hexene and side chain groups of amino acid methyl ester complexes. As the internal acyl side chains of POPC bilayers are protected from interacting with water molecules and salt ions, the QM calculations are reasonable.
CH motif directly, QM calculations were performed not only in POPC–side chain group complexes but also in truncated systems of 3-hexene and side chain groups of amino acid methyl ester complexes. As the internal acyl side chains of POPC bilayers are protected from interacting with water molecules and salt ions, the QM calculations are reasonable.
The IR vibration frequencies of ![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH and C
CH and C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C stretching bands of POPC that were involved in special interactions with side chain groups at the ωB97X-D/6-31G (d,p) level are presented in Table 1. The corresponding optimised geometries are shown in Fig. S1 in the ESI.† The frequency of the C
C stretching bands of POPC that were involved in special interactions with side chain groups at the ωB97X-D/6-31G (d,p) level are presented in Table 1. The corresponding optimised geometries are shown in Fig. S1 in the ESI.† The frequency of the C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C stretching mode of POPC bathochromically shifted by about 3.09 cm−1 in the benzene–POPC complex, and the
C stretching mode of POPC bathochromically shifted by about 3.09 cm−1 in the benzene–POPC complex, and the ![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH stretching band hypochromatically shifted by about 8.18 cm−1. The calculated red shift of C
CH stretching band hypochromatically shifted by about 8.18 cm−1. The calculated red shift of C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C stretching band was lower than the experimental value (5 cm−1). The calculated blue shift of
C stretching band was lower than the experimental value (5 cm−1). The calculated blue shift of ![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH stretching band did not match the experimental red shift value (3 cm−1). In the indole–POPC complex, a red shift of 8.27 cm−1 in the frequency of the C
CH stretching band did not match the experimental red shift value (3 cm−1). In the indole–POPC complex, a red shift of 8.27 cm−1 in the frequency of the C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C stretching mode and a red shift of 4.14 cm−1 in the frequency of the
C stretching mode and a red shift of 4.14 cm−1 in the frequency of the ![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH stretching mode were observed. The calculated red shift of C
CH stretching mode were observed. The calculated red shift of C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C stretching band was higher than the experimental value (5 cm−1). And the calculated red shift of
C stretching band was higher than the experimental value (5 cm−1). And the calculated red shift of ![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH stretching band was higher than the experimental value (3 cm−1). Also, the
CH stretching band was higher than the experimental value (3 cm−1). Also, the ![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH stretching band bathochromically shifted by about 5.62 cm−1, and the C
CH stretching band bathochromically shifted by about 5.62 cm−1, and the C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C stretching band hypochromatically shifted by about 0.14 cm−1 in the phenol–POPC complex. The calculated red shift of C
C stretching band hypochromatically shifted by about 0.14 cm−1 in the phenol–POPC complex. The calculated red shift of C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C stretching band was higher than the experimental value (5 cm−1). And the calculated blue shift of
C stretching band was higher than the experimental value (5 cm−1). And the calculated blue shift of ![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH stretching band did not match the experimental red shift value (2 cm−1). In addition, a red shift of 4.65 cm−1 in the frequency of the C
CH stretching band did not match the experimental red shift value (2 cm−1). In addition, a red shift of 4.65 cm−1 in the frequency of the C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C stretching mode and a blue shift of 3.15 cm−1 in the frequency of the
C stretching mode and a blue shift of 3.15 cm−1 in the frequency of the ![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH stretching mode were observed in the propane–POPC complex. This calculated red shift of C
CH stretching mode were observed in the propane–POPC complex. This calculated red shift of C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C stretching band was higher than the experimental value (0 cm−1). And the calculated blue shift of
C stretching band was higher than the experimental value (0 cm−1). And the calculated blue shift of ![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH stretching band did not match the experimental red shift value (1 cm−1). These results demonstrate that the calculated shifts of C
CH stretching band did not match the experimental red shift value (1 cm−1). These results demonstrate that the calculated shifts of C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C and
C and ![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH stretching bands in POPC–side chain groups did not match the experimental red shifts value.
CH stretching bands in POPC–side chain groups did not match the experimental red shifts value.
![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C) and ν(
C) and ν(![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH) for POPC and POPC–side chain group complexes at the ωB97X-D/6-31G (d,p) level
CH) for POPC and POPC–side chain group complexes at the ωB97X-D/6-31G (d,p) level
| Sample | ν(C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C) C) |
ν(![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH) CH) |
ν(C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C) (scaled) C) (scaled) |
ν(![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH) (scaled) CH) (scaled) |
|---|---|---|---|---|
| POPC | 1775.32 | 3190.76 | 1686.55 | 3031.22 |
| POPC and benzene | 1772.06 | 3199.37 | 1683.46 | 3039.40 |
| POPC and indole | 1766.61 | 3195.01 | 1678.28 | 3035.26 |
| POPC and phenol | 1769.40 | 3190.91 | 1680.93 | 3031.36 |
| POPC and propane | 1770.42 | 3194.03 | 1681.90 | 3034.33 |
As the acyl chains may not affect the frequency of the HC![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH motif significantly, the simplified theoretical model is reasonable. So the geometry optimisation and frequency calculation were performed at the MP2/6-31G (d,p) level for model systems further. The IR vibration frequencies of
CH motif significantly, the simplified theoretical model is reasonable. So the geometry optimisation and frequency calculation were performed at the MP2/6-31G (d,p) level for model systems further. The IR vibration frequencies of ![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH and C
CH and C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C stretching bands of 3-hexene that were involved in special interactions with side chain groups at the MP2/6-31G (d,p) level are presented in Table 2. The frequency of the C
C stretching bands of 3-hexene that were involved in special interactions with side chain groups at the MP2/6-31G (d,p) level are presented in Table 2. The frequency of the C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C stretching mode of 3-hexene bathochromically shifted by about 3.63 cm−1 in the benzene–3-hexene complex, and the
C stretching mode of 3-hexene bathochromically shifted by about 3.63 cm−1 in the benzene–3-hexene complex, and the ![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH stretching band hypochromatically shifted by about 0.68 cm−1. The calculated red shift of C
CH stretching band hypochromatically shifted by about 0.68 cm−1. The calculated red shift of C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C stretching band was lower than the experimental value (5 cm−1). The calculated blue shift of
C stretching band was lower than the experimental value (5 cm−1). The calculated blue shift of ![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH stretching band did not match the experimental red shift value (3 cm−1). Besides, a red shift of 14.46 cm−1 in the frequency of the C
CH stretching band did not match the experimental red shift value (3 cm−1). Besides, a red shift of 14.46 cm−1 in the frequency of the C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C stretching mode and a red shift of 5.83 cm−1 in the frequency of the
C stretching mode and a red shift of 5.83 cm−1 in the frequency of the ![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH stretching mode were observed in the indole–3-hexene complex. The calculated red shift of C
CH stretching mode were observed in the indole–3-hexene complex. The calculated red shift of C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C stretching band was higher than the experimental value (5 cm−1). And the calculated red shift of
C stretching band was higher than the experimental value (5 cm−1). And the calculated red shift of ![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH stretching band was higher than the experimental value (3 cm−1). Also, the
CH stretching band was higher than the experimental value (3 cm−1). Also, the ![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH stretching band bathochromically shifted by about 4.9 cm−1, and the C
CH stretching band bathochromically shifted by about 4.9 cm−1, and the C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C stretching band hypochromatically shifted by about 2.07 cm−1 in the phenol–3-hexene complex. The calculated red shift of C
C stretching band hypochromatically shifted by about 2.07 cm−1 in the phenol–3-hexene complex. The calculated red shift of C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C stretching band was lower than the experimental value (5 cm−1). And the calculated blue shift of
C stretching band was lower than the experimental value (5 cm−1). And the calculated blue shift of ![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH stretching band did not match the experimental red shift value (2 cm−1). In addition, a red shift of 3.2 cm−1 in the frequency of the C
CH stretching band did not match the experimental red shift value (2 cm−1). In addition, a red shift of 3.2 cm−1 in the frequency of the C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C stretching mode and a red shift of 1.91 cm−1 in the frequency of the
C stretching mode and a red shift of 1.91 cm−1 in the frequency of the ![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH stretching mode were observed in the propane–3-hexene complex. This calculated red shift of C
CH stretching mode were observed in the propane–3-hexene complex. This calculated red shift of C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C stretching band was higher than the experimental value (0 cm−1). And the calculated red shift of
C stretching band was higher than the experimental value (0 cm−1). And the calculated red shift of ![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH stretching band was higher than the experimental value (1 cm−1).
CH stretching band was higher than the experimental value (1 cm−1).
![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C) and ν(
C) and ν(![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH) for 3-hexene and 3-hexene–side chain group complexes at the MP2/6-31G (d,p) level
CH) for 3-hexene and 3-hexene–side chain group complexes at the MP2/6-31G (d,p) level
| Sample | ν(C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C) C) |
ν(![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH) CH) |
ν(C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C) (scaled) C) (scaled) |
ν(![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH) (scaled) CH) (scaled) |
|---|---|---|---|---|
| 3-Hexene | 1748.01 | 3206.75 | 1630.89 | 2991.90 |
| 3-Hexene and benzene | 1744.12 | 3207.48 | 1627.26 | 2992.58 |
| 3-Hexene and indole | 1732.51 | 3200.50 | 1616.43 | 2986.07 |
| 3-Hexene and phenol | 1742.75 | 3208.97 | 1625.99 | 2993.97 |
| 3-Hexene and propane | 1744.58 | 3204.71 | 1627.69 | 2989.99 |
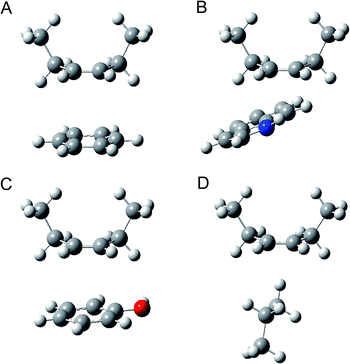
The corresponding optimised geometries at the MP2/6-31G (d,p) level are shown in Fig. 3. The calculation evidenced that the benzene ring and the phenol ring are preferentially located below the C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C bond of 3-hexene in favor of the parallel π–π stacking mode. The indole ring preferentially interacts with the C
C bond of 3-hexene in favor of the parallel π–π stacking mode. The indole ring preferentially interacts with the C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C bond of 3-hexene at the π–π stacking mode. Besides, the N–H bond of indole ring locates towards the C
C bond of 3-hexene at the π–π stacking mode. Besides, the N–H bond of indole ring locates towards the C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C bond of 3-hexene. So the NH–π interaction may also exist between the indole ring and the HC
C bond of 3-hexene. So the NH–π interaction may also exist between the indole ring and the HC![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH motif of 3-hexene. While the propane is preferentially located below the C
CH motif of 3-hexene. While the propane is preferentially located below the C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C bond of 3-hexene at the σ–π mode.
C bond of 3-hexene at the σ–π mode.
These theoretical calculation results of 3-hexene interacting with aromatic rings mainly at the π–π stacking mode are not consistent with the FTIR spectroscopic measurements of aromatic amino acid methyl ester–POPC bilayer complexes. It indicates that these models of the 3-hexene and side chain groups of amino acid methyl esters complexes differ from the real interaction mechanism of amino acid methyl ester–POPC bilayer complexes. Therefore, it can be concluded that the red shifts of the ![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH and C
CH and C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C stretching modes for POPC acyl chains are not through a direct interaction between the side chain groups and the C
C stretching modes for POPC acyl chains are not through a direct interaction between the side chain groups and the C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C bond. Besides, as the hydrophilic group of amino acid interacts with the choline group of lipids,42 the phenylalanine interacts with the interface of DPPC model membrane by hydrophobic interaction, the dipole interaction and the hydrogen bonding,43 the amino acid methyl esters may interact with the lipid bilayer surface without deep penetration. It further infers that the red shifts of the
C bond. Besides, as the hydrophilic group of amino acid interacts with the choline group of lipids,42 the phenylalanine interacts with the interface of DPPC model membrane by hydrophobic interaction, the dipole interaction and the hydrogen bonding,43 the amino acid methyl esters may interact with the lipid bilayer surface without deep penetration. It further infers that the red shifts of the ![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH and C
CH and C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C stretching bands of POPC acyl chains are caused by the packing and conformational change of POPC acyl chains in membrane with the side chain groups of amino acid methyl esters insertion.
C stretching bands of POPC acyl chains are caused by the packing and conformational change of POPC acyl chains in membrane with the side chain groups of amino acid methyl esters insertion.
3.3. Dichroic properties of POPC bilayers and POPC bilayer–amino acid methyl ester complexes
Dichroic properties are useful to characterise the membrane order,44 the orientation of lipids45 and the conformation of proteins.46 For lipids, when the acyl chains in the membrane bilayers become more disordered, the frequency values of asymmetric and symmetric CH2 stretching vibrations are blue-shifted.44 Therefore, the frequencies of the CH2 stretching vibrations provide useful information about the conformational order of lipid acyl chains as a sensitive qualitative measurement.RATR parameters for lipids and lipid–amino acid methyl ester complexes are presented in Table 3. The RATR of lipids is determined from the peak intensity of the asymmetric CH2 stretching mode because the intensity of νasym(CH2) peak is stronger than that of νsym(CH2) peak. The band for νasym(CH2) appearing at 2920 cm−1 suggests that bilayered membranes are in a liquid crystalline phase.47 Table 3 shows that pure POPC bilayers are well ordered because the orientation factor f of the hydrocarbon chains of POPC bilayers was 0.40. However, the RATR significantly increased when Trp-OMe was added in POPC bilayers. Correspondingly, the orientation factor f decreased to 0.32 ± 0.01 and the γ increased to 42.3° ± 0.4°. Thus the average angle of lipid chains relative to the membrane normal increases. This result indicates that the addition of Trp-OMe induces an obvious decrease in the order of acyl chains. The RATR also increased significantly when Phe-OMe and Tyr-OMe were added with similar magnitude with the case of Trp-OMe. However, the RATR shows less increment when Leu-OMe was added. These results indicate that the packing and conformation of POPC acyl chains can be influenced significantly when aromatic amino acid methyl esters were mixed into POPC bilayers, and the amplitude of conformational order decrease for POPC acyl chains mainly depends on the steric hindrance of the insertion of side chain groups.
| Sample | RATR of νas(CH2) | f | γ |
|---|---|---|---|
| a Derivations mean the upper and lower limits calculated values for three measurements. | |||
| POPC | 1.18 | 0.40 | 39.2° |
| POPC and Trp-OMe | 1.32 ± 0.02 | 0.32 ± 0.01 | 42.3° ± 0.4° |
| POPC and Phe-OMe | 1.33 ± 0.02 | 0.32 ± 0.01 | 42.3° ± 0.4° |
| POPC and Tyr-OMe | 1.32 ± 0.02 | 0.32 ± 0.01 | 42.3° ± 0.4° |
| POPC and Leu-OMe | 1.24 ± 0.03 | 0.37 ± 0.02 | 40.4° ± 0.8° |
3.4. QM calculation results of 3-hexene and 3-hexene–3-hexene complexes
From the above discussion, it suggests that the C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C and
C and ![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH bond of POPC bilayers in POPC bilayer–amino acid methyl ester complexes become different than those in pure POPC bilayers, with corresponding FTIR vibrational frequencies of the
CH bond of POPC bilayers in POPC bilayer–amino acid methyl ester complexes become different than those in pure POPC bilayers, with corresponding FTIR vibrational frequencies of the ![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH and C
CH and C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C stretching bands for POPC acyl chains showing bathochromic shifts. It is mainly due to the packing and conformational change of POPC acyl chains in membrane when amino acid methyl esters were mixed into POPC bilayers. Besides, the steric hindrance causing by the side chain groups of amino acid methyl ester may become the primary factor affecting the conformational change of POPC acyl chains. As previous publication reported that the acyl chains are packed close to each other and the dispersion interactions between the CH2 groups of adjacent chains are enhanced in gel state, thus the frequency of CH2 group is lower in the gel than in the fluid state.48 We can presume that the conformational change of POPC acyl chains result in the phenomena that the HC
C stretching bands for POPC acyl chains showing bathochromic shifts. It is mainly due to the packing and conformational change of POPC acyl chains in membrane when amino acid methyl esters were mixed into POPC bilayers. Besides, the steric hindrance causing by the side chain groups of amino acid methyl ester may become the primary factor affecting the conformational change of POPC acyl chains. As previous publication reported that the acyl chains are packed close to each other and the dispersion interactions between the CH2 groups of adjacent chains are enhanced in gel state, thus the frequency of CH2 group is lower in the gel than in the fluid state.48 We can presume that the conformational change of POPC acyl chains result in the phenomena that the HC![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH motif is packed close to the other one of an adjacent POPC molecule and the dispersion interactions between the HC
CH motif is packed close to the other one of an adjacent POPC molecule and the dispersion interactions between the HC![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH motifs of adjacent chains are enhanced when amino acid methyl esters were mixed into POPC membrane. To obtain more detailed information about the phenomena that HC
CH motifs of adjacent chains are enhanced when amino acid methyl esters were mixed into POPC membrane. To obtain more detailed information about the phenomena that HC![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH motif interact with the other HC
CH motif interact with the other HC![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH motif directly, QM calculations were performed in truncated systems of 3-hexene and 3-hexene complex at the MP2/6-31G (d,p) level.
CH motif directly, QM calculations were performed in truncated systems of 3-hexene and 3-hexene complex at the MP2/6-31G (d,p) level.
In 3-hexene–3-hexene complex, the frequency of the C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C stretching mode bathochromically shifted by about 4.48 cm−1, and the
C stretching mode bathochromically shifted by about 4.48 cm−1, and the ![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH stretching band bathochromically shifted by about 1.59 cm−1, as listed in Table 4. The corresponding optimised geometries evidenced that the 3-hexene is preferentially located below the C
CH stretching band bathochromically shifted by about 1.59 cm−1, as listed in Table 4. The corresponding optimised geometries evidenced that the 3-hexene is preferentially located below the C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C bond of 3-hexene at the parallel offset π–π interaction mode. And the vertical distance of two 3-hexene is 3.47 Å. Fig. 4 shows that the amplitude of bathochromic shifts of the C
C bond of 3-hexene at the parallel offset π–π interaction mode. And the vertical distance of two 3-hexene is 3.47 Å. Fig. 4 shows that the amplitude of bathochromic shifts of the C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C stretching vibrational frequencies for 3-hexene–3-hexene complexes at the offset π–π interaction mode decreased with the distance of the C9 and C27 increased from 3.27 Å to 3.87 Å every 0.2 Å. The corresponding atomic coordinates found for the minimum energy conformers of the 3-hexene–3-hexene complexes with different distance are available in the Tables S2–S5 in the ESI.† The intensity of offset π–π interaction between the two HC
C stretching vibrational frequencies for 3-hexene–3-hexene complexes at the offset π–π interaction mode decreased with the distance of the C9 and C27 increased from 3.27 Å to 3.87 Å every 0.2 Å. The corresponding atomic coordinates found for the minimum energy conformers of the 3-hexene–3-hexene complexes with different distance are available in the Tables S2–S5 in the ESI.† The intensity of offset π–π interaction between the two HC![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH motifs is inversely proportional to the distance of the HC
CH motifs is inversely proportional to the distance of the HC![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH motifs. It demonstrates that the dispersion interactions are enhanced when the HC
CH motifs. It demonstrates that the dispersion interactions are enhanced when the HC![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH motif is packed close to the other one, and the corresponding C
CH motif is packed close to the other one, and the corresponding C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C and
C and ![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH stretching bands show red shifts.
CH stretching bands show red shifts.
![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C) and ν(
C) and ν(![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH) for 3-hexene and 3-hexene–3-hexene complex at the MP2/6-31G (d,p) level
CH) for 3-hexene and 3-hexene–3-hexene complex at the MP2/6-31G (d,p) level
| Sample | ν(C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C) C) |
ν(![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH) CH) |
ν(C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C) (scaled) C) (scaled) |
ν(![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH) (scaled) CH) (scaled) |
|---|---|---|---|---|
| 3-Hexene | 1748.01 | 3206.75 | 1630.89 | 2991.90 |
| 3-Hexene and 3-hexene | 1743.20 | 3205.05 | 1626.41 | 2990.31 |
 | ||
Fig. 4 Red shifts of the C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C stretching frequencies for 3-hexene–3-hexene complexes at the offset π–π interaction mode with different distance of the C9 and C27 (please refer to Fig. S2 in the ESI† for atom numberings) determined by QM calculations at the MP2/6-31G (d,p) level. C stretching frequencies for 3-hexene–3-hexene complexes at the offset π–π interaction mode with different distance of the C9 and C27 (please refer to Fig. S2 in the ESI† for atom numberings) determined by QM calculations at the MP2/6-31G (d,p) level. | ||
As dispersion interactions play important roles in numerous chemical and biochemical systems.49 These interaction can be involved in the molecular recognition50 and stability of proteins.51 The MP2/6-31G (d,p) level has been used to geometry optimization of saccharide–aromatic residue systems that contain CH–π and OH–π interactions52 and thermochemistry analysis of the π–π stacking interactions of various aromatic ring systems.53 Besides, the ωB97X-D/6-31G (d,p) level has been used to geometry optimization of asphaltene model compounds and analysis the contributions of hydrogen-bonding and π–π stacking interactions to the aggregation.54 So these methods are appropriate to calculate dispersion interactions, such as N/CH–π interactions, π–π stacking interactions, etc.
The amino acid methyl esters may be located at the surface of POPC bilayers, the side chain groups may not insert deeply into the hydrophobic core of POPC membrane. When the side chain groups of amino acid methyl esters insert into the head region of POPC bilayers serving as anchors, the conformation and packing of surrounding lipid acyl chains change, and the dispersion interactions between the HC![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH motifs of adjacent chains are enhanced. The aromatic amino acid methyl esters can alter the physicochemical properties of POPC bilayers such as molecular packing and fluidity, being relevant to the function of membrane proteins and permeability of lipid bilayers.
CH motifs of adjacent chains are enhanced. The aromatic amino acid methyl esters can alter the physicochemical properties of POPC bilayers such as molecular packing and fluidity, being relevant to the function of membrane proteins and permeability of lipid bilayers.
In accordance with previous polarised ATR-FTIR spectroscopy and QM calculation results, investigations reported above have demonstrated that the membrane order decrease and the conformation of lipid acyl chains change result in the significant red shifts of C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C and
C and ![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH stretching mode of POPC acyl chains when the aromatic amino acid methyl esters were mixed in. Besides, it also maybe speculated that the conformational change of POPC acyl chains result in the phenomena that the HC
CH stretching mode of POPC acyl chains when the aromatic amino acid methyl esters were mixed in. Besides, it also maybe speculated that the conformational change of POPC acyl chains result in the phenomena that the HC![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH motif is packed closed to the other one of an adjacent POPC molecule and the dispersion interactions between the HC
CH motif is packed closed to the other one of an adjacent POPC molecule and the dispersion interactions between the HC![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH motifs of adjacent chains are enhanced, thus the red shifts of the C
CH motifs of adjacent chains are enhanced, thus the red shifts of the C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C and
C and ![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH stretching mode exist. The frequencies of the
CH stretching mode exist. The frequencies of the ![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH and C
CH and C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C stretching bands of C
C stretching bands of C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C bond in POPC acyl chain can be seen as the useful intrinsic IR probe. The specific intensity, bandwidth or wavenumber changes of different IR vibration modes representing the characteristic functional group in biological systems can be used to monitor the simultaneously subtle changes in the structure and function of protein and model membrane. The findings of this study can shed valuable insight into how different hydrophobicity and size of amino acid methyl esters influence the packing of unsaturated lipid acyl chains and membrane structure. Besides, the intrinsic IR probe is useful to detect in various biological and chemistry systems.
C bond in POPC acyl chain can be seen as the useful intrinsic IR probe. The specific intensity, bandwidth or wavenumber changes of different IR vibration modes representing the characteristic functional group in biological systems can be used to monitor the simultaneously subtle changes in the structure and function of protein and model membrane. The findings of this study can shed valuable insight into how different hydrophobicity and size of amino acid methyl esters influence the packing of unsaturated lipid acyl chains and membrane structure. Besides, the intrinsic IR probe is useful to detect in various biological and chemistry systems.
4. Conclusions
FTIR spectroscopy combined with QM calculations was used to investigate the red shifts of the![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH and C
CH and C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C stretching bands of HC
C stretching bands of HC![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH motif in POPC acyl chains. Besides, the utility of using lipid containing HC
CH motif in POPC acyl chains. Besides, the utility of using lipid containing HC![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH motif to probe the specific interaction between unsaturated lipids and aromatic residues was demonstrated. ATR-FTIR spectroscopy combined with polarisation experiments showed that the HC
CH motif to probe the specific interaction between unsaturated lipids and aromatic residues was demonstrated. ATR-FTIR spectroscopy combined with polarisation experiments showed that the HC![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH motif of POPC acyl chains was influenced by aromatic amino acid methyl esters significantly. QM calculation results demonstrated that the red shifts of the
CH motif of POPC acyl chains was influenced by aromatic amino acid methyl esters significantly. QM calculation results demonstrated that the red shifts of the ![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH and C
CH and C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C stretching mode for POPC acyl chains are not through a direct interaction between the side chain groups and the HC
C stretching mode for POPC acyl chains are not through a direct interaction between the side chain groups and the HC![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH motif but are mainly because of the conformational change of POPC acyl chains, thus results in the phenomena that the HC
CH motif but are mainly because of the conformational change of POPC acyl chains, thus results in the phenomena that the HC![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH motif is packed closed to the other one of an adjacent acyl chain with the dispersion interactions enhancing. The difference of red shifts depends on the hydrophobicity and structural size of side chain groups of amino acid methyl esters. This study provides a valuable insight into the influence of the lipid acyl chains packing by small molecules interacting with membrane, the dynamics properties of biological membranes and raises possibilities for the binding of corresponding membrane protein to the membrane.
CH motif is packed closed to the other one of an adjacent acyl chain with the dispersion interactions enhancing. The difference of red shifts depends on the hydrophobicity and structural size of side chain groups of amino acid methyl esters. This study provides a valuable insight into the influence of the lipid acyl chains packing by small molecules interacting with membrane, the dynamics properties of biological membranes and raises possibilities for the binding of corresponding membrane protein to the membrane.
Abbreviations
| POPC | 2-Oleoyl-1-pamlitoyl-sn-glyecro-3-phosphocholine |
| ATR-FTIR | Attenuated total reflectance-Fourier transfer infrared spectroscopy |
| ZnSe | Zinc selenide |
| QM | Quantum mechanical |
| Trp-OMe | Tryptophan methyl ester |
| Phe-OMe | Phenylalanine methyl ester |
| Tyr-OMe | Tyrosine methyl ester |
| Leu-OMe | Leucine methyl ester |
| HEPES | 4-(2-Hydroxyethyl)-1-piperazineethanesulphonic acid |
Acknowledgements
We are grateful for the support for this work provided by the National Natural Science Foundation of China (30970657, 21475045), the Student Partners Program (09PJ1404300), and East China Normal University (79003A29, 79301207, 79301411 and 41500-515430-14100).References
- M. M. Kozlov, Nature, 2010, 463, 439–440 CrossRef CAS PubMed.
- R. Phillips, T. Ursell, P. Wiggins and P. Sens, Nature, 2009, 459, 379–385 CrossRef CAS PubMed.
- J. L. Kavanau, Nature, 1963, 198, 525–530 CrossRef CAS PubMed.
- M. Luckey, Membrane Structural Biology: With Biochemical and Biophysical Foundations, Cambridge University Press, 2014 Search PubMed.
- R. N. A. H. Lewis and R. N. McElhaney, Biochim. Biophys. Acta, 2013, 1828, 2347–2358 CrossRef CAS PubMed.
- M. Arczewska, D. M. Kamiński, E. Górecka, D. Pociecha, E. Rój, A. Sławińska-Brych and M. Gagoś, Biochim. Biophys. Acta, 2013, 1828, 213–222 CrossRef CAS PubMed.
- A. M. Whited and A. Johs, Chem. Phys. Lipids, 2015, 192, 51–59 CrossRef CAS PubMed.
- D. C. Mitchell, M. Straume and B. J. Litman, Biochemistry, 1992, 31, 662–670 CrossRef CAS PubMed.
- J. Šajbidor, Crit. Rev. Biotechnol., 1997, 17, 87–103 CrossRef PubMed.
- M. Caffrey and J. Hogan, Chem. Phys. Lipids, 1992, 61, 1–109 CrossRef CAS PubMed.
- B. Peter, A. A. Polyansky, S. Fanucchi and H. W. Dirr, Biochemistry, 2014, 53, 57–67 CrossRef CAS PubMed.
- L. Garneau, H. Klein, M. F. Lavoie, E. Brochiero, L. Parent and R. Sauve, J. Gen. Physiol., 2014, 143, 289–307 CrossRef CAS PubMed.
- J. Cheng, R. Goldstein, A. Gershenson, B. Stec and M. F. Roberts, J. Biol. Chem., 2013, 288, 14863–14873 CrossRef CAS PubMed.
- G. Wang, B. Mishra, R. F. Epand and R. M. Epand, Biochim. Biophys. Acta, 2014, 1838, 2160–2172 CrossRef CAS PubMed.
- K. S. Mineev, N. F. Khabibullina, E. N. Lyukmanova, D. A. Dolgikh, M. P. Kirpichnikov and A. S. Arseniev, Biochim. Biophys. Acta, 2011, 1808, 2081–2088 CrossRef CAS PubMed.
- E. V. Bocharov, M. L. Mayzel, P. E. Volynsky, K. S. Mineev, E. N. Tkach, Y. S. Ermolyuk, A. A. Schulga, R. G. Efremov and A. S. Arseniev, Biophys. J., 2010, 98, 881–889 CrossRef CAS PubMed.
- D. Williams, J. Vicogne, I. Zaitseva, S. McLaughlin and J. E. Pessin, Mol. Biol. Cell, 2009, 20, 4910–4919 CrossRef CAS PubMed.
- A. Merlino, G. Vitiello, M. Grimaldi, F. Sica, E. Busi, R. Basosi, A. M. D'Ursi, G. Fragneto, L. Paduano and G. D'Errico, J. Phys. Chem. B, 2012, 116, 401–412 CrossRef CAS PubMed.
- P. Wolfangel and K. Muller, J. Phys. Chem. B, 2003, 107, 9918–9928 CrossRef CAS.
- K. Cieslik-Boczula, B. Czarnik-Matusewicz, M. Perevozkina and M. Rospenk, RSC Adv., 2015, 5, 40455–40464 RSC.
- A. Barth, Biochim. Biophys. Acta, 2007, 1767, 1073–1101 CrossRef CAS PubMed.
- S. D. Moran and M. T. Zanni, J. Phys. Chem. Lett., 2014, 5, 1984–1993 CrossRef CAS PubMed.
- Y. Shai, Biochim. Biophys. Acta, 2013, 1828, 2306–2313 CrossRef CAS PubMed.
- S. Ye, K. T. Nguyen and Z. Chen, J. Phys. Chem. B, 2010, 114, 3334–3340 CrossRef CAS PubMed.
- L. Tamm and S. Tatulian, Q. Rev. Biophys., 1997, 30, 365–429 CrossRef CAS PubMed.
- S. Leekumjorn and A. K. Sum, J. Phys. Chem. B, 2007, 111, 6026–6033 CrossRef CAS PubMed.
- C. C. Valley, J. D. Perlmutter, A. R. Braun and J. N. Sachs, J. Membr. Biol., 2011, 244, 35–42 CrossRef CAS PubMed.
- M. J. Frisch, G. W. Trucks, H. B. Schlegel, G. E. Scuseria, M. A. Robb, J. R. Cheeseman, G. Scalmani, V. Barone, B. Mennucci, G. A. Petersson, H. Nakatsuji, M. Caricato, X. Li, H. P. Hratchian, A. F. Izmaylov, J. Bloino, G. Zheng, J. L. Sonnenberg, M. Hada, M. Ehara, K. Toyota, R. Fukuda, J. Hasegawa, M. Ishida, T. Nakajima, Y. Honda, O. Kitao, H. Nakai, T. Vreven, J. A. Montgomery Jr, J. E. Peralta, F. Ogliaro, M. Bearpark, J. J. Heyd, E. Brothers, K. N. Kudin, V. N. Staroverov, T. Keith, R. Kobayashi, J. Normand, K. Raghavachari, A. Rendell, J. C. Burant, S. S. Iyengar, J. Tomasi, M. Cossi, N. Rega, J. M. Millam, M. Klene, J. E. Knox, J. B. Cross, V. Bakken, C. Adamo, J. Jaramillo, R. Gomperts, R. E. Stratmann, O. Yazyev, A. J. Austin, R. Cammi, C. Pomelli, J. W. Ochterski, R. L. Martin, K. Morokuma, V. G. Zakrzewski, G. A. Voth, P. Salvador, J. J. Dannenberg, S. Dapprich, A. D. Daniels, O. Farkas, J. B. Foresman, J. V. Ortiz, J. Cioslowski and D. J. Fox, Gaussian 09, Revision B.01, Gaussian, Inc., Wallingford, CT, 2010 Search PubMed.
- J.-D. Chai and M. Head-Gordon, Phys. Chem. Chem. Phys., 2008, 10, 6615–6620 RSC.
- M. J. Frisch, M. Head-Gordon and J. A. Pople, Chem. Phys. Lett., 1990, 166, 275–280 CrossRef CAS.
- I. M. Alecu, J. Zheng, Y. Zhao and D. G. Truhlar, J. Chem. Theory Comput., 2010, 6, 2872–2887 CrossRef CAS PubMed.
- A. Blume, Curr. Opin. Colloid Interface Sci., 1996, 1, 64–77 CrossRef CAS.
- S. H. White and W. C. Wimley, Biochim. Biophys. Acta, 1998, 1376, 339–352 CrossRef CAS.
- V. Sivasakthi, A. Anbarasu and S. Ramaiah, Cell Biochem. Biophys., 2013, 67, 853–863 CrossRef CAS PubMed.
- A. Roy, P. Bouř and T. A. Keiderling, Chirality, 2009, 21, E163–E171 CrossRef CAS PubMed.
- W.-C. Chao, J.-Y. Shen, J.-F. Lu, J.-S. Wang, H.-C. Yang, K. Wee, L.-J. Lin, Y.-C. Kuo, C.-H. Yang, S.-H. Weng, H.-C. Huang, Y.-H. Chen and P.-T. Chou, J. Phys. Chem. B, 2014, 119, 2157–2167 CrossRef PubMed.
- K. L. Copeland, J. A. Anderson, A. R. Farley, J. R. Cox and G. S. Tschumper, J. Phys. Chem. B, 2008, 112, 14291–14295 CrossRef CAS PubMed.
- M. Levitt and M. F. Perutz, J. Mol. Biol., 1988, 201, 751–754 CrossRef CAS PubMed.
- T. Iwata, A. Watanabe, M. Iseki, M. Watanabe and H. Kandori, J. Phys. Chem. Lett., 2011, 2, 1015–1019 CrossRef CAS.
- S. Nicholas, Acta Crystallogr., Sect. C: Struct. Chem., 2015, 71, 211–215 CAS.
- J. R. Long, F. D. Mills, O. K. Ganesh, V. C. Antharam and R. S. Farver, Biochim. Biophys. Acta, 2010, 1798, 216–222 CrossRef CAS PubMed.
- A. V. Popova and D. K. Hincha, Chem. Phys. Lipids, 2004, 132, 171–184 CrossRef CAS PubMed.
- A. S. Rosa, A. C. Cutro, M. A. Frías and E. A. Disalvo, J. Phys. Chem. B, 2015, 119, 15844–15847 CrossRef CAS PubMed.
- D. Bilge, N. Kazanci and F. Severcan, J. Mol. Struct., 2013, 1040, 75–82 CrossRef CAS.
- Z. D. Schultz and I. W. Levin, Annu. Rev. Anal. Chem., 2011, 4, 343–366 CrossRef CAS PubMed.
- W.-N. Huang, Y.-H. Chen, C.-L. Chen and W. Wu, Langmuir, 2011, 27, 7034–7041 CrossRef CAS PubMed.
- D. G. Cameron, H. L. Casal and H. H. Mantsch, Biochemistry, 1980, 19, 3665–3672 CrossRef CAS PubMed.
- E. A. Disalvo, A. M. Bouchet and M. A. Frias, Biochim. Biophys. Acta, 2013, 1828, 1683–1689 CrossRef CAS PubMed.
- S. Ehrlich, J. Moellmann and S. Grimme, Acc. Chem. Res., 2013, 46, 916–926 CrossRef CAS PubMed.
- L. M. Salonen, M. Ellermann and F. Diederich, Angew. Chem., Int. Ed., 2011, 50, 4808–4842 CrossRef CAS PubMed.
- C. J. Pace, D. Kim and J. Gao, Chem.–Eur. J., 2012, 18, 5832–5836 CrossRef CAS PubMed.
- M. Kumari, R. B. Sunoj and P. V. Balaji, Org. Biomol. Chem., 2012, 10, 4186–4200 CAS.
- L. Moreira da Costa, S. R. Stoyanov, S. Gusarov, P. R. Seidl, M. C. J. Walkimar de and A. Kovalenko, J. Phys. Chem. A, 2014, 118, 896–908 CrossRef CAS PubMed.
- L. M. da Costa, S. R. Stoyanov, S. Gusarov, X. Tan, M. R. Gray, J. M. Stryker, R. Tykwinski, J. W. d. M. Carneiro, P. R. Seidl and A. Kovalenko, Energy Fuels, 2012, 26, 2727–2735 CrossRef CAS.
Footnote |
| † Electronic supplementary information (ESI) available. See DOI: 10.1039/c6ra05903a |
| This journal is © The Royal Society of Chemistry 2016 |