Identification of potential quinoxalinone-based aldose reductase inhibitors by 3D-QSAR, molecular docking and molecular dynamics
Dan Zhoua,
Jianbo Chenb and
Yi Xu*a
aSchool of Chemical and Environmental Engineering, Shanghai Institute of Technology, 100 Haiquan Road, Shanghai 201418, China. E-mail: xuyi@sit.edu.cn; Fax: +86 21 60873567; Tel: +86 21 60873162
bCollege of Life and Environmental Sciences, Shanghai Normal University, Shanghai 200234, China
First published on 12th May 2016
Abstract
The aldose reductase ALR2 (EC1.1.1.21) has been confirmed to be a target for the treatment of chronic diabetic complications, and ALR2 inhibitors (commonly known as ARIs) are the most effective drugs for this condition. In this work, new potential ARIs were designed using a combination of molecular modeling techniques. First, three-dimensional quantitative structure–activity relationship (3D-QSAR) models were built by applying comparative molecular field analysis (CoMFA), comparative molecular similarity indices analysis (CoMSIA), and topomer CoMFA. The best CoMFA model yielded a q2 = 0.894 and r2 = 0.994, the best CoMSIA model yielded a q2 = 0.907 and r2 = 0.995, and the topomer CoMFA analysis yielded a q2 = 0.752 and r2 = 0.985, which all exhibited satisfactory correlations and predictive abilities. The results also indicated the steric, electrostatic, H-donor and hydrophobic fields to play key roles in the models. Molecular docking was then performed to explore the conformations of the inhibitors and the key amino acid residues at the docking pocket, and the binding between the inhibitors and the receptor protein. Molecular dynamics (MD) simulations further validated the reliability of the docking results. By analyzing the above results, six new potential inhibitor compounds were identified and predicted to have good biological activity. The information acquired in this study should provide valuable guidance for the development of potent and effective ALR2 inhibitors.
1. Introduction
Diabetes Mellitus (DM) is a type of complex and chronic metabolic disease characterized by hyperglycemia. The chronic complications of diabetes include neuropathy, nephropathy, cataracts, retinopathy, accelerated atherosclerosis and increased cardiovascular risk, which are threats to human health and may be lethal.1–4 With the improvement of living standards, the occurrence-probability of diabetes is becoming increasingly higher. The number of global diabetic patients is at least 120 million. In Europe and other developed countries, the incidence of diabetes is only second to cardiovascular disease; mortality due to diabetes ranks third in developed countries, behind cardiovascular disease and cancer. China has about 20 million to 30 million diabetic patients, more than any other country in the world. Prevention and treatment of diabetes has been a common and urgent problem to be solved in the world medical community.5Aldose reductase (ALR2, EC 1.1.1.21) is a member of the aldo-keto reductase superfamily,6,7 and catalyzes the reduction of glucose to sorbitol in the presence of the cofactor NADPH/NADP during the rate-determining step of the polyol metabolic pathway. Under normal conditions, glucose is mainly converted to glucose-6-phosphate by hexokinase and then enters into the glycolytic pathway. However, to the diabetic patients, the affinity between ALR2 and glucose will be improved and the polyol metabolic pathway is activated. This results in the glucose metabolism occurring in the polyol pathway instead of in the glycolytic pathway (Fig. 1). Along with issues regarding metabolism, such patients show increased levels of sorbitol, an imbalance between NADPH/NADP and NAD/NADH cofactors, and increased oxidative stress—all thought to be major causes of cellular damage that in turn cause diabetic complications. Recently, additional studies have provided evidence for playing a critical role in the development of long-term diabetic complications. In the past few decades, many ALR2 inhibitors (commonly known as ARIs) have been explored, but only one of them, epalrestat, is available for therapy. Most of the promising ARIs cannot be used clinically, mainly due to pharmacokinetic drawbacks, adverse side effects, or low in vivo efficacy.8–10 Therefore, new drugs to be applied for chronic metabolic disease are urgently needed. Recently, a series of new ARIs have been synthesized based on quinoxaline. These ARIs have been shown not only to be able to inhibit aldose reductase but also to directly suppress the oxidative stress that is closely related to diabetic complications. The traditional ARIs can only indirectly inhibit oxidative stress and the effect is not obvious.11 To develop more new and effective ARIs, an effective tool for drug design, the quantitative structure–activity relationship (QSAR) method, had already been applied in this area. Ghasemi et al.12 and Aalizadeh et al.13 have established good models to determine the correlation between inhibitors and receptors and have predicted new active inhibitors by this method.
In this work, 3D-QSAR methods, i.e., CoMFA,14 CoMSIA15 and the high-efficiency topomer CoMFA, were applied to gain insights into the key structural factors affecting the inhibitory activity of a new type of ARI. The developed models not only can be used to predict the activity of newly designed inhibitors, but also to provide beneficial information to modify structures for designing new inhibitors with desired inhibitory activity. In addition, molecular docking was carried out to study the binding modes of inhibitors at the active site of the ALR2. A molecular dynamics (MD) simulation was performed to confirm the reliability of the docking results. Based on the results of the calculation, we designed six new compounds predicted to have good biological activity and can act as a reference for synthesis.
2. Results and discussion
2.1 Computational approach
3D-QSAR studies and molecular docking were performed using the molecular modeling package SYBYL-X2.0 (Tripos, Inc., USA) running on a Windows 7 workstation. Three-dimensional structures of all the inhibitors in the training and testing sets were constructed by using the Sketch Molecule module. Energy minimization was performed by applying the Powell gradient algorithm with the Tripos force field and Gasteiger-Hückel charge. The maximum number of iterations for the minimization was set to 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000. The minimization was terminated when the energy gradient convergence criterion reached 0.005 kcal (mol−1 Å−1). MD simulations were carried out using the Sander module of the Amber12 software package. Xmgrace software was used to show figures of RMSD results, running on a Linux system.
000. The minimization was terminated when the energy gradient convergence criterion reached 0.005 kcal (mol−1 Å−1). MD simulations were carried out using the Sander module of the Amber12 software package. Xmgrace software was used to show figures of RMSD results, running on a Linux system.
2.2 Data sets
The 35 compounds involved in this study were reported recently by Qin.11 Their IC50 values were found by them to be in the range of 0.032–5.981 μM. The corresponding pIC50 values of the inhibitors can be derived from their IC50 values according to the formula pIC50 = −lg![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) IC50, whose values are proportional to their biological activities. The samples were randomly divided into a training set of 25 inhibitors for model generation and a test set of 10 inhibitors for model validation. A structurally and functionally diverse set of inhibitors were selected. The structures of these inhibitors and their pIC50 values against ARL2 are shown in Table 1. Inhibitor number 19 in the table showed the highest activity and was chosen as the template molecule for superpositions. A superposition of the training set molecules is shown in Fig. 2a and the atoms of the training molecules were fitted to each other to generate the superposition which is shown in red in Fig. 2b. Each of the training set structures was broken into two sets of fragments, R1 (red) and R2 (blue) which were used for generating topomer CoMFA, as shown in Fig. 2c.
IC50, whose values are proportional to their biological activities. The samples were randomly divided into a training set of 25 inhibitors for model generation and a test set of 10 inhibitors for model validation. A structurally and functionally diverse set of inhibitors were selected. The structures of these inhibitors and their pIC50 values against ARL2 are shown in Table 1. Inhibitor number 19 in the table showed the highest activity and was chosen as the template molecule for superpositions. A superposition of the training set molecules is shown in Fig. 2a and the atoms of the training molecules were fitted to each other to generate the superposition which is shown in red in Fig. 2b. Each of the training set structures was broken into two sets of fragments, R1 (red) and R2 (blue) which were used for generating topomer CoMFA, as shown in Fig. 2c.
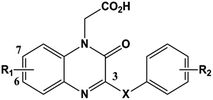
| No. | R1 | R2 | X | Actual pIC50 | Predicted CoMFA | Residual ΔpIC50 | Predicted CoMSIA | Residual ΔpIC50 |
|---|---|---|---|---|---|---|---|---|
| a Training = training set inhibitors.b Test = test set inhibitors. | ||||||||
| Traininga | ||||||||
| 1 | H | 4-OH | CH2–CH2 | 6.098 | 6.101 | −0.003 | 6.110 | 0.009 |
| 2 | 7-F | 4-OH | CH2–CH2 | 6.186 | 6.177 | 0.009 | 6.169 | −0.008 |
| 3 | H | H | O | 6.330 | 6.331 | −0.001 | 6.333 | 0.002 |
| 4 | H | H | S | 6.376 | 6.429 | −0.053 | 6.396 | −0.033 |
| 5 | H | 4-Br | S | 6.529 | 6.535 | −0.006 | 6.569 | 0.034 |
| 6 | 7-F | 4-Br | S | 6.719 | 6.758 | −0.039 | 6.767 | 0.009 |
| 7 | 6-Br | 4-Br | S | 6.496 | 6.417 | 0.079 | 6.437 | 0.020 |
| 8 | H | 4-Cl | S | 6.564 | 6.529 | 0.035 | 6.534 | 0.005 |
| 9 | 7-Cl | 4-Cl | S | 6.801 | 6.813 | −0.012 | 6.790 | −0.023 |
| 10 | H | H | CH2 | 5.954 | 5.954 | 0 | 5.953 | −0.001 |
| 11 | H | H | CH2–CH2 | 6.844 | 6.888 | −0.044 | 6.863 | −0.025 |
| 12 | H | H | — | 5.223 | 5.307 | −0.084 | 5.240 | −0.067 |
| 13 | H | 4-F | — | 5.471 | 5.461 | 0.01 | 5.508 | 0.047 |
| 14 | 7-Cl | 4-F | — | 5.819 | 5.799 | 0.02 | 5.774 | −0.025 |
| 15 | 4-Fluoroenyl | 4-F | — | 5.671 | 5.670 | 0.001 | 5.683 | 0.013 |
| 16 | H | 4-OH | — | 5.586 | 5.563 | 0.023 | 5.570 | 0.007 |
| 17 | 6-F | 2,4-(OH)2 | — | 7.200 | 7.191 | 0.009 | 7.180 | −0.011 |
| 18 | 6-Cl | 2,4-(OH)2 | — | 7.022 | 7.062 | −0.04 | 7.111 | 0.049 |
| 19 | 7-F | 2,4-(OH)2 | — | 7.495 | 7.438 | 0.057 | 7.424 | −0.014 |
| 20 | H | 3-Indole | — | 6.194 | 6.127 | 0.067 | 6.156 | 0.029 |
| 21 | 7-Br | 3-Indole | — | 6.434 | 6.473 | −0.039 | 6.478 | 0.005 |
| 22 | 7-Cl | 2-Benzothiophene | — | 6.623 | 6.628 | −0.005 | 6.596 | −0.032 |
| 23 | H | 4-OH | CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH CH |
6.740 | 6.682 | 0.058 | 6.696 | 0.014 |
| 24 | 7-F | 4-OH | CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH CH |
6.815 | 6.901 | −0.086 | 6.893 | −0.008 |
| 25 | H | 3-OCH3, 4-OH | CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH CH |
6.378 | 6.332 | 0.046 | 6.367 | 0.035 |
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) |
||||||||
| Testb | ||||||||
| 26 | 7-Cl | 4-Br | S | 6.487 | 6.816 | 0.329 | 6.822 | 0.335 |
| 27 | 7-Br | 4-Br | S | 6.330 | 6.841 | 0.511 | 6.885 | 0.555 |
| 28 | 7-F | 4-Cl | S | 7.252 | 6.754 | −0.498 | 6.733 | −0.519 |
| 29 | 7-Br | 4-Cl | S | 6.403 | 6.838 | 0.435 | 6.853 | 0.45 |
| 30 | H | H | CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH CH |
6.086 | 6.177 | 0.091 | 6.339 | 0.253 |
| 31 | 7-F | 4-F | — | 6.058 | 5.731 | −0.327 | 5.715 | −0.343 |
| 32 | H | 2,4-(OH)2 | — | 6.401 | 7.171 | 0.77 | 7.219 | 0.818 |
| 33 | 6-Br | 2,4-(OH)2 | — | 6.857 | 7.034 | 0.177 | 7.062 | 0.205 |
| 34 | 7-Cl | 2,4-(OH)2 | — | 7.161 | 7.498 | 0.337 | 7.485 | 0.324 |
| 35 | 7-Br | 2,4-(OH)2 | — | 7.041 | 7.510 | 0.469 | 7.552 | 0.511 |
2.3 Statistical results for models
A partial least squares (PLS) analysis,16 an extension of multiple regression analysis, was carried out to derive 3D-QSAR models. CoMFA, CoMSIA and topomer CoMFA models were developed and the final models were selected according to the statistical parameters. Here, we calculated various parameters to find the best models. The statistical results of the CoMFA, CoMSIA and topomer CoMFA models are summarized in Table 2. The internal validations of LOO cross-validated q2 and non-cross-validated coefficient r2 are commonly applied as the criteria of robustness and predictive ability of a QSAR model. The values considered to be statistically satisfactory for a QSAR model are q2 > 0.5 (0.2 for topomer CoMFA) and r2 > 0.831.17 As shown in Table 2, the predictions by the CoMFA and topomer CoMFA models appeared to be good, having LOO cross-validated coefficient q2 values of 0.894 and 0.752, the optimal number of components (ONC) (i.e., 9 and 7), and the correlation coefficient r2 values of 0.994 and 0.985, respectively. The standard error (S = 0.055) and F-statistic value (F = 260.638) of CoMFA further validated the developed model. The contributions of steric and electrostatic field were calculated to be 54.7 and 45.3%, respectively. For the CoMSIA analysis, an excellent r2 value of 0.995 for the prediction was obtained for this model with a q2 value of 0.907, and ONC of 11. SEE and F-statistic values were calculated to be 0.055 and 218.293, respectively. Five descriptor fields (steric, electrostatic, hydrophobic, hydrogen bond-donor, and hydrogen bond acceptor) were considered. Their contributions were 6.9%, 25.4%, 22.2%, 35.6% and 9.9%, respectively. The electrostatic, hydrophobic and hydrogen bond donor fields were found to make the significant contributions and to play significant roles in the prediction of inhibitory activity. Among them, the hydrogen bond donor field was shown to be the most important in the optimal CoMSIA model. The predicted values were found to be generally in good agreement with the experimental data, as shown in Table 1. The actual and predicted pIC50 value of the training and test set molecules are illustrated in Fig. 3.| CoMFA | CoMSIA | Topomer CoMFA | |
|---|---|---|---|
| q2 | 0.894 | 0.907 | 0.752 |
| ONC | 9.000 | 11.00 | 7 |
| r2 | 0.994 | 0.995 | 0.985 |
| SEE | 0.055 | 0.055 | — |
| F | 260.638 | 218.293 | — |
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) |
|||
| Fraction | |||
| Steric | 0.547 | 0.069 | — |
| Electrostatic | 0.453 | 0.254 | — |
| Hydrophobic | — | 0.222 | — |
| H-donor | — | 0.356 | — |
| H-acceptor | — | 0.099 | — |
 | ||
| Fig. 3 Scatter plots of the predicted versus actual pIC50 values for all the molecules based on the CoMFA (a) and CoMSIA models (b). | ||
2.4 Contour maps of models
The contour maps produced by CoMFA, CoMSIA and topomer CoMFA were analyzed to visualize the field effects on the target inhibitors in 3D space. For the convenience of the analysis, the most active molecule, i.e., number 19 in Table 1, was superimposed onto contour maps to explain the key structural features required for inhibitory activity. Such contour maps are important for designing new drugs because the regions shown in 3D space are where modifications of the molecular fields strongly correlate with variations in biological activity.The steric fields of CoMFA, CoMSIA and topomer CoMFA are shown in Fig. 4, where the green (favorable) and yellow (unfavorable) contours represent 80% and 20% levels of contribution, respectively. Fig. 4 shows that the steric contour map of CoMFA agreed with that of CoMSIA, which depicts a large green contour covering the hydroxyl group links to C2 and C4 of R2, suggesting the importance of the presence of a bulky group in this region for biological activity. The contour maps of topomer CoMFA (Fig. 4c and d) around R1 and R2 were found to be identical with those of the CoMFA and CoMSIA models, indicative of the consistency of the results. These results were found to be is in agreement with the experimental data. For example, inhibitor 19 (pIC50 = 7.495) was observed to have a large group at this position, whereas inhibitor 31 (pIC50 = 6.058) was seen to have a smaller group at this position. Similarly, the group in 34 was observed to be larger than that in 14, and the group in 32 was larger than that in 12. These results all indicate the importance of a bulky group at this position. The two yellow contours around C5 and C6 of R2 indicated that the compound with bulky substitution possessed poor biological activity.
 | ||
| Fig. 4 Contour maps of steric properties. Green contours indicate regions where bulky groups increased the activity, and yellow contours indicate regions where bulky groups decreased the activity. | ||
The electrostatic field contour maps of CoMFA, CoMSIA and topomer CoMFA are shown in Fig. 5. The electrostatic field is indicated by blue and red contours, where the blue regions denote the electropositive groups found to be favorable to the activity, and the red regions indicate the electronegative groups found to be favorable to the activity. As shown in Fig. 5a, the two medium sized red contours at C6 and C7 of R1 indicate the importance of electronegative atoms in improving the biological activity. This trend is reflected by the trends in the pIC50 values, with that of 17 (pIC50 = 7.200) > 18 (pIC50 = 7.022) > 33 (pIC50 = 6.857) and that of 19 (pIC50 = 7.495) > 34 (pIC50 = 7.161) > 35 (pIC50 = 7.041), (electronegativity: F > Cl > Br). Similarly, a medium-sized region of red contour appears in C4 of R2. This trend is also reflected in the pIC50 values of 16 (pIC50 = 5.586) and 12 (pIC50 = 5.223) in accordance with the result of the H atom replaced by hydroxyl in C4 of R2 (electronegativity: O > H ). The big blue contour observed near C5 and C6 of R2 suggested this position to not be suitable for substitution with the electronegative atoms or groups. The lower biological activity of 21 than that of 35 resulted from the influence of the pyrrole group, the same as for the relationship between 20 and 32. Topomer CoMFA yielded a similar result to those of the CoMFA and CoMSIA models, and the analysis of the topomer CoMFA result is therefore not further described.
In the hydrophobic fields, regions that favor the binding of hydrophobic and hydrophilic moieties are shown in yellow and white, respectively. In Fig. 6a, one large yellow (hydrophobic-favorable) contour was observed to cover the C7 of R1 and a small white (hydrophilic-favorable) contour to cover the C6 of R1, suggesting that the introduction of hydrophobic substituents into the C7 of R1 and hydrophilic substituents into the C6 of R1 would be beneficial for inhibitory activity. This prediction is in agreement with the experimental data: inhibitor 31 (7-F) > 13 (H); 19 (7-F) > 32 (H); 2 (7-F) > 1 (H); 24 (7-F) > 23 (H); 6 (7-F) > 5 (H) and 28 (7-F) > 8 (H). Besides, a big yellow contour was observed to be far from C2 of R2, indicating little effect of the hydrophobic field on inhibitory activity. The white contour on C4 of R2 indicated the preference for hydrophilic substituents, which is reflected in 12 (H) and 16 (–OH).
As shown in Fig. 6b, the hydrogen-bond donor field was found to be the most important contributor in the optimal CoMSIA model, playing a more important role in bioactivity than the other fields. The purple contours represent the positions where the hydrogen-bond donor disfavors biological activity, and the cyan contours indicate that the presence of donor groups in this region should produce better biological activity. Three purple contours were observed (Fig. 6b): one of them at a single-bonded O atom of a carboxyl, another in front of C4, and the third one far from C3 and C4 of R1. In Fig. 6c, the hydrogen-bond acceptor field is represented by magenta and red contours, with the magenta contours denoting regions where the hydrogen-bond acceptor group would be beneficial to the bioactivity, and the red contours representing the regions where the hydrogen-bond acceptor group would decrease the bioactivity. A small peak of magenta was observed to cover C4 of R2 and a large one was found around the C linked to the C of the carboxyl. Furthermore, a large extent of red contour inlayed in the double-bonded O atom of the carboxyl of inhibitor 16 was found to have a higher biological activity than 13, which confirmed the above analysis.
2.5 Docking analysis
Complementary information from docking studies is vital for the design of new molecules. Docking studies not only highlight the correspondence between the QSAR and binding conformations but also provide the crucial interactions with the active site for enhanced activity.18 Herein, the most active inhibitor 19 and least active inhibitor 12 toward ARL2 (PDB ID: 1Z3N) whose co-crystallized ligand is similar to current ARIs, were selected for more detailed analysis.Fig. 7 and 8 show the interacting mode of inhibitor 19 and 12 in the binding site of ALR2. As shown in Fig. 7, the quinoxalinone core of inhibitor 19 matched very well with the hydrophobic pocket (Fig. 7a) lined by Trp (20, 79, 111, 219) Val (47, 297), Phe (121, 122), Pro218, Asn 160 and other residues (Fig. 7b). Five hydrogen bonds formed between inhibitor 19 and residues Trp20, Tyr48, Gln49, His110 and Cys298 of ALR2. The corresponding hydrogen-bond distances were measured to be 1.92 Å (Trp20–H⋯HO–), 1.94 Å (Tyr48–OH⋯O–), 2.05 Å (Gln49–O⋯HO–), 2.02 Å (His110–N⋯HO–) and 2.53 Å (Cys298–NH⋯F–) (Fig. 7c). Inhibitor 19 was observed to be tightly bound to the active site of ALR2 by these H-bonds, with the carboxylate group of inhibitor 19 well inserted, using hydrogen bonds, into the anion-binding site of the side chains of Tyr48 and His110. Besides, many other types of interactions were also obtained. The electrostatic, van der Waals and π–π interactions are displayed in Fig. 7c by pink, green and orange, respectively.
Fig. 8 shows the docking result of inhibitor 12 with ALR2. This ligand was docked in the binding pocket with only two H-bonds, one formed between –NH of Trp20 and –O of the inhibitor with a distance of 2.681 Å, and the other one (1.97 Å) between the H atom linked to the N atom of the indole ring of the Trp111 side chain with an O atom in the quinoxalinone core. Compared with those of 19, hydrogen-bonding interactions in 12 were weakened, on account of it lacking an F atom in C7 of R1 and two hydroxyls in C2 of R2. This difference indicated the formation of hydrogen bonds to be necessary for the activity of ARIs. This result could explain the CoMSIA model of the 3D-QSAR, which predicted the hydrogen-bond donor field to provide the most important contributions in the model. All of the inhibitors with F atoms in C7 of R1 were found to have relatively high pIC50 values for ALR2, and these high values were probably due to the affinity of F atom to hydrogen bonds. Besides, C4 of R2 was also found to prefer hydrogen donors, and this preference is consistent with the analysis of the hydrogen-bond acceptor field contour maps in Fig. 6c. The electrostatic interactions of inhibitor 19 were observed to be much more evenly distributed than were those of inhibitor 12, which may be the reason for different pIC50 values between inhibitors 19 and 12, given the importance of the electrostatic field.
2.6 MD simulations
The structures of inhibitors 19 and 12 docked to ALR2 were used as the initial structures for MD simulations. The general AMBER force field (gaff)19 was used for the inhibitors and the ff99SB force field was used for the protein.20 The system was neutralized with Na+ counter ions. The whole system was computationally immersed in a rectangular box of TIP3P water molecules, which extended 10 Å from solute atoms in all three dimensions.A two-stage minimization strategy was used to relax each system: first, the protein was fixed, and the water molecules and ligand were minimized by carrying out 500 cycles of steepest descent and 500 cycles of conjugate gradient minimization; second, the whole system was minimized by performing 1000 cycles of steepest descent and 4000 cycles of conjugate gradient minimization. Then, each system was gradually heated in the NVT ensemble from 0 to 300 K over a period of 20 ps. Finally, 20 ns NPT MD simulations with a target temperature of 300 K and a target pressure of 1 atm were performed. The SHAKE procedure was employed to constrain all bonds involving hydrogen atoms, and the time step was set to 2 fs.21 Coordinate trajectories were saved every 20 ps throughout each of the MD runs. The MM optimization and MD simulations were accomplished by using the Sander program in AMBER12. The binding free energy ΔGbind was estimated according to eqn (1) instead of (2), considering the expensive computational cost and low prediction accuracy.22–34
| ΔGbind = ΔEMM + ΔGGB + ΔGSA | (1) |
| ΔEbind = ΔGcomplex − Gprotein − Gligand = ΔH + ΔGsolv − TΔS = ΔEMM + ΔGGB + ΔGSA − TΔS | (2) |
In these equations, ΔEMM is the gas-phase interaction energy between protein and ligand, which consists of the electrostatic (ΔEele) and van der Waals (ΔEvdw) terms; and ΔGGB and ΔGSA represent the polar and non-polar terms of desolvation free energy, respectively. The electrostatic solvation energy (ΔGGB) was calculated by using the GB model with the parameters developed by Onufriev et al. (igb = 2).35 The exterior dielectric constant was set to 80, and the solute dielectric constant was set to 1. The contribution of nonpolar residues to desolvation (ΔGSA) was estimated by calculating the solvent-accessible surface area (SASA) using the LCPO method.36 All energy components were calculated using 100 snapshots extracted from 19 to 20 ns. In order to analyze the interactions between each residue in ALR2 and inhibitors 19 and 12, MM/GBSA free energy decomposition in the MM_PBSA program in AMBER12 was employed.37–43 The residue-inhibitor interaction involved four components: the van der Waals contribution (ΔGvdw), electrostatic contribution (ΔGele), the polar part of desolvation (ΔGGB) and the nonpolar part of desolvation (ΔGSA). The exterior dielectric constant was set to 80, and the solute dielectric constant value was set to 1. The electrostatic desolvation energy (ΔGGB) was estimated by using the GB model based on the parameters developed. The nonpolar contribution of desolvation (ΔGSA) was determined by calculating the SASA using the ICOSA technique.40
As shown in Fig. 9a and b, the complexes 19-ALR2 and 12-ALR2 reached their equilibria after 11 ns and 5 ns, respectively. For each case the superimposition of the average structure from the last 2 ns with the initial structure is shown in Fig. 10. The RMSD values between each of the two complexes and their initial structures were found to be relatively small, within 2.5 Å and 1.5 Å, respectively, suggesting that the structure of ALR2 would only change slightly upon being bound by either of these inhibitors. The result of the MD simulations indicates that two whole systems of the complex are stabilized and that the docking results are reliable. The information from the last 2 ns of the 20 ns run was extracted to figure out the binding free energy. Then the energy was decomposed to inspect the contributor to the energy. Based on the MD simulations, the binding free energies for the inhibitors were calculated and are listed in Table 3. The binding free energy ΔGbind values of the compound 19-ALR2 and 12-ALR2 complexes were calculated to be −24.32 kcal mol−1 and −16.39 kcal mol−1, respectively. The gas-phase interaction energy (ΔEMM) was considered to have provided the largest contributions among the four components.
 | ||
| Fig. 9 The root-mean-square deviation (RMSD) of the docked complex 19 (a) and 12 (b) versus MD simulation structures. | ||
3. Molecular design of new ARIs
The detailed contour map analysis of CoMFA, CoMSIA and topomer CoMFA models allowed us to identify the structural requirements for the observed inhibitory activity. Based on the QSAR results, inhibitor 19, which was indicated to have the highest activity, was taken as a template to design new compounds. For example, since a green contour covering C2 and C4 of R2 indicated the importance of the presence of a bulky group in this region for biological activity, a bulky hydroxyl was introduced to this position, resulting in designed compounds 19(a–f). A red contour near C3 of R2 showed the importance of the electronegative atom at this position, and thus compounds 19(a–f) with one F, Cl and Br substituted at C3 of R2 were designed. A cyan contour around C2 of R2 indicated that the presence of donor groups in this region should produce better biological activity, so a guanidyl group was introduced to this position in 19(a–f). To predict their biological activities, the CoMFA and CoMSIA models were applied to these new molecules and the corresponding results are listed in Table 4. The results showed that the six predicted values of COMFA and CoMSIA are all higher than before, indicative of their good biological activity. Therefore it is predicted that these compounds perhaps should be regarded as good candidates for experimental synthesis.| NO | R1 | R2 | X | Predicted CoMFA | Predicted CoMSIA |
|---|---|---|---|---|---|
| 19 | 7-F | 2,4-(OH)2 | — | 7.484 | 7.582 |
| 19a | 7-F | 2-Guanidyl; 3-Br; 4-OH | — | 7.633 | 7.832 |
| 19b | 7-F | 2-Guanidyl; 3-Cl; 4-OH | — | 7.641 | 7.820 |
| 19c | 7-F | 2-Guanidyl; 3-F; 4-OH | — | 7.658 | 7.828 |
| 19d | 7-F | 2-Guanidyl; 3-Br; 4-NH2 | — | 7.557 | 8.381 |
| 19e | 7-F | 2-Guanidyl; 3-Cl, 4-NH2 | — | 7.572 | 8.366 |
| 19f | 7-F | 2-Guanidyl; 3-F, 4-NH2 | — | 7.592 | 8.371 |
4. Conclusion
In the present work, 35 potent quinoxalinone-based aldose reductase inhibitors were analyzed to develop QSAR models. The results of CoMFA, CoSIA and Topomer CoMFA models were convincing and comparable. The models were significantly favored by internal and external predictions as well as visualization of contour maps. Contour maps obtained from CoMFA, CoMSIA and topomer CoMFA analyses provided crucial clues that were used for the successful design of potent aldose reductase inhibitors with high predicted values. Molecular docking analyses of the representative inhibitors were performed to determine the binding modes of the inhibitors at the active sites of the ALR2 protein. Some key residues such as Asn 160 and the hydrophobic residues Trp (20, 79, 111, 219), Val (47, 297), Phe (121, 122), and Pro218, as well as hydrogen bonds between the inhibitors and Trp20, Tyr48, Gln49, His110 and Cys298 of the active site were observed. The molecules that interact with residues Trp20, Tyr48, Gln49, His110 and Cys298 may generate binding affinities that are less susceptible to point mutations. This study could serve as a basis for the development of aldose reductase inhibitors. MD simulations of two ALR2-inhibitor complexes further proved the reliability of docking results and gave their binding free energy values. From an energy decomposition analysis, the gas-phase interaction energy (ΔEMM) was considered to have provided the largest contribution among the four components. Six new compounds were designed, based on an analysis of the models, and these compounds may be considered for experimental synthesis. It was expected that the results from molecular modeling can provide valuable guidance for the development of potent and effective ALR2 inhibitors.Acknowledgements
This work was supported by National Natural Science Foundation (No. 21303105), Shanghai Committee of Science and Technology (No. 13430503400), the Science Foundation of Shanghai Institute of Technology (No. YJ2010-04) and the Scientific Research Foundation for the Returned Overseas Chinese Scholars, State Education Ministry (No. ZX2012-05).Notes and references
- American Diabetes Association, Diabetes Care, 2013, vol. 36, pp. 67–74 Search PubMed.
- P. Alexiou, K. Pegklidou, M. Chatzopoulou, I. Nicolaou and V. J. Demopoulos, Curr. Med. Chem., 2009, 16, 734–752 CrossRef PubMed.
- C. Yabenishimura, Pharmacol. Rev., 1998, 50, 21–33 Search PubMed.
- S. S. Chung, E. C. Ho, K. S. Lam and S. K. Chung, J. Am. Soc. Nephrol., 2003, 10, 1375–1387 Search PubMed.
- M. Cobuz and C. Cobuz, Rom. J. Diabetes Nutr. Metab. Dis., 2012, 19, 301–309 Search PubMed.
- Y. L. Kao, K. Donaghue, A. Chan, J. Knight and M. Silink, Diabetes Res. Clin. Pract., 1999, 46, 155–160 CrossRef PubMed.
- D. K. Moczulski, W. Burak, A. Doria, M. Zychma and E. Zukow-ska-Szczechowska, et al., Diabetologia, 1999, 42, 94–97 CrossRef PubMed.
- R. Kikkawa, I. Hatanaka, H. Yasuda, N. Kobayashiand and Y. Shigeta, et al., Diabetologia, 1983, 24, 290–292 CrossRef PubMed.
- M. Ramirez and N. Borja, Pharmacotherapy, 2008, 28, 646–655 CrossRef PubMed.
- S. Ao, Y. Shingu, C. Kikuchi and Y. Takano, et al., Metabolism, 1991, 40, 77–87 CrossRef PubMed.
- X. Qin, X. Hao, H. Han and S. Zhu, et al., J. Med. Chem., 2015, 58, 1254–1267 CrossRef PubMed.
- J. B. Ghasemi, E. Nazarshodeh and H. Abedi, J. Iran. Chem. Soc., 2015, 12, 1–11 CrossRef.
- R. Aalizadeh, E. Pourbasheer and M. R. Ganjali, Mol. Diversity, 2015, 19, 1–16 CrossRef PubMed.
- R. D. Cramer, D. E. Patterson and J. D. Bunce, J. Am. Chem. Soc., 1988, 110, 5959–5967 CrossRef PubMed.
- G. Klebe, U. Abraham and T. Mietzner, J. Med. Chem., 1994, 37, 4130–4146 CrossRef PubMed.
- L. Ståhle and S. Wold, J. Chemom., 1987, 1, 185–196 CrossRef.
- G. Alexander and T. Alexander, J. Mol. Graphics Modell., 2002, 20, 269–276 CrossRef.
- A. Politi, S. Durdagi, P. Moutevelis-Minakakis and G. Kokotos, et al., Eur. J. Med. Chem., 2009, 44, 3703–3711 CrossRef PubMed.
- J. Wang, R. M. Wolf, J. W. Caldwell and P. A. Kollman, et al., J. Comput. Chem., 2004, 25, 1157–1174 CrossRef PubMed.
- V. Hornak, R. Abel, A. Okur and B. Strockbine, et al., Proteins: Struct., Funct., Bioinf., 2006, 65, 712–725 CrossRef PubMed.
- J. P. Ryckaert, G. Ciccotti and H. J. C. Berendsen, Semin. Oncol., 1977, 23, 292–301 Search PubMed.
- J. Wang, T. Hou and X. Xu, Curr. Comput.-Aided Drug Des., 2006, 2, 287–306 CrossRef.
- T. Hou and R. Yu, J. Med. Chem., 2007, 50, 1177–1188 CrossRef PubMed.
- H. Liu, X. Yao, C. Wang and J. Han, Mol. Pharm., 2010, 7, 894–904 CrossRef PubMed.
- J. Zhang, T. Hou, W. Wang and J. S. Liu, Proc. Natl. Acad. Sci. U. S. A., 2010, 107, 1321–1326 CrossRef PubMed.
- T. Hou, J. Wang, Y. Li and W. Wang, J. Comput. Chem., 2011, 32, 866–877 CrossRef PubMed.
- T. Hou, J. Wang, Y. Li and W. Wang, J. Chem. Inf. Model., 2011, 51, 69–82 CrossRef PubMed.
- W. Xue, D. Pan, Y. Yang and H. Liu, et al., Antiviral Res., 2012, 93, 126–137 CrossRef PubMed.
- N. Homeyer and H. Gohlke, Mol. Inf., 2012, 31, 114–122 CrossRef.
- P. A. Kollman, I. Massova, C. Reyes and B. Kuhn, et al., Acc. Chem. Res., 2000, 33, 889–897 CrossRef PubMed.
- S. Huo, J. Wang, P. Cieplak and P. A. Kollman, et al., J. Med. Chem., 2002, 45, 1412–1419 CrossRef PubMed.
- B. Kuhn, P. Gerber, T. Schulz-Gasch and M. Stahl, J. Med. Chem., 2005, 48, 4040–4048 CrossRef PubMed.
- B. Kuhn and P. A. Kollman, J. Med. Chem., 2000, 43, 3786–3791 CrossRef PubMed.
- S. Huo, I. Massova and P. A. Kollman, J. Comput. Chem., 2002, 23, 15–27 CrossRef PubMed.
- A. Onufriev, D. A. Bashford and D. A. Case, Mon. Not. R. Astron. Soc., 2004, 55, 383–394 Search PubMed.
- J. Weiser, P. S. Shenkin and W. C. Still, J. Comput. Chem., 1999, 20, 217–230 CrossRef.
- T. Hou, Z. Xu, W. Zhang and W. A. McLaughlin, et al., Mol. Cell. Proteomics, 2009, 8, 639–649 Search PubMed.
- T. Hou, W. Zhang, D. Case and W. Wang, J. Mol. Biol., 2008, 376, 1201–1214 CrossRef PubMed.
- H. Gohlke, C. Kiel and D. A. Case, J. Mol. Biol., 2003, 330, 891–913 CrossRef CAS PubMed.
- T. Hou, N. Li, Y. Li and W. Wang, J. Proteome Res., 2012, 11, 2982–2995 CrossRef CAS PubMed.
- T. Hou, W. A. McLaughlin and W. Wang, Proteins: Struct., Funct., Bioinf., 2008, 71, 1163–1174 CrossRef CAS PubMed.
- T. Hou, W. Zhang, J. Wang and W. Wang, Proteins: Struct., Funct., Bioinf., 2009, 74, 837–846 CrossRef CAS PubMed.
- T. Hou, Y. Li and W. Wang, Bioinformatics, 2011, 27, 1814–1821 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2016 |