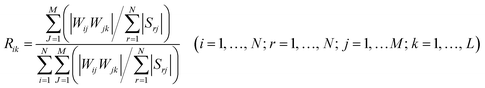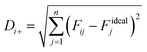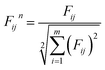The optimization of the fermentation process of wheat germ for flavonoids and two benzoquinones using EKF-ANN and NSGA-II
Ziyi Zheng,
Xiaona Guo,
Kexue Zhu*,
Wei Peng and
Huiming Zhou*
State Key Laboratory of Food Science and Technology, Collaborative Innovation Center for Food Safety and Quality Control, School of Food Science and Technology, Jiangnan University, 1800 Lihu Avenue, Wuxi 214122, Jiangsu Province, People’s Republic of China. E-mail: kxzhu@jiangnan.edu.cn; hmzhou@jiangnan.edu.cn; Fax: +86 510 85809610
First published on 23rd May 2016
Abstract
The fermentation process for maximizing the flavonoid and methoxy-ρ-benzoquinone (MBQ) + 2,6-dimethoxy-ρ-benzoquinone (DMBQ) content of wheat germ was modeled and optimized. First, agitation speed, initial pH, fermentation temperature and fermentation time were used to construct a 4-8-2 ANN model with back-propagation (BP) and extended Kalman filter (EKF) learning algorithms. The regression coefficients (R2) between experimental and predicted values indicated that EKF-ANN models had better accuracy. Second, a multi-objective optimization procedure using non-dominated sorting genetic algorithm II (NSGA-II) was performed to create non-dominated optimal solutions which gave an insight on the optimum fermentation parameters for maximum flavonoid and MBQ + DMBQ content. Third, an approach based on LINMAP was utilized for finding the final compromise solution from the obtained non-dominated optimal solutions. The optimization results show a 13.34% increase in flavonoid content and a 7.92% increase in the total content of MBQ and DMBQ simultaneously, compared with the highest values in original design.
1. Introduction
Even though wheat germ is the most nutritious part of the wheat kernel, it is mainly used as a kind of fodder for poultry.1 Recently, a number of studies have shown the potential of fermented wheat germ in medical applications. A continuous supplement of fermented wheat germ extract for more than six months has a positive effect towards progression-free survival in colorectal cancer patients.2 Another study suggested that extract of fermented wheat germ was able to act on lymphoid tumor cells by decreasing the expression of major histocompatibility complex class I proteins, thus promoting apoptosis of tumor cells.3 Moreover, in an in vitro study, extract of fermented wheat germ exerted significant antitumor activity in 32 human cancer cell lines, and showed synergistic drug interaction with 5-FU, oxaliplatin and irinotecan.4 Two remarkable antitumor components of fermented wheat germ are two methoxy-substituted benzoquinones, i.e. methoxy-ρ-benzoquinone (MBQ) and 2,6-dimethoxy-ρ-benzoquinone (DMBQ).4 The bioconversion pathways of MBQ and DMBQ have been elucidated. MBQ and DMBQ are derived from hydroquinones substituted by β-1,6-linked oligosaccharides in wheat germ, which can firstly be hydrolysed to their aglycones, i.e., 2,6-ρ-dimethoxy hydroquinone and methoxy-ρ-hydroquinone by β-glucosidase. Then the two hydroquinones are oxidized to homologous ρ-benzoquinones by peroxidase.5 Nevertheless, Lot6p (YLR011wp), a soluble quinone reductase from Saccharomyces cerevisiae, could react with the two benzoquinones by reduction to give back the corresponding hydroquinones,6 as shown in Fig. 1. Moreover, fermented wheat germ also contains significant levels of flavonoids which have been widely proved to reduce the risk of cancer.7,8 Numerous authors found that the conversion of glycosylated flavonoids into corresponding aglycones resulted in better biological activity during yeast or lactobacillus fermentation.9,10 Nevertheless, a research study reported that flavonol glycosides were reduced during fermentation due to oxidative degradation.11To our knowledge, hardly any literature is available based on flavonoids of fermented wheat germ. Regarding the two benzoquinones, a previous study showed that the total content of MBQ and DMBQ in fermented wheat germ increased by 470% compared with the total content in raw wheat germ.12 On the other hand, it can be concluded that factors in the fermentation process also have an influence on the activities of β-glucosidase and peroxidase, thus affecting the total content of MBQ and DMBQ in fermented wheat germ. Similarly, factors in the fermentation process affect the amount of flavonoids in fermented wheat germ. Nevertheless, there is not much information about the effects of the fermentation parameters on the production of MBQ, DMBQ and flavonoids. Statistical methods, such as the response surface method (RSM) have been widely used to optimize the fermentation process in other studies.13,14 However, the development of accurate models for the fermentation process is still the key difficulty due to the non-linear nature of biochemical interactions.15 Recently, artificial neural networks (ANN) have been proved to be feasible and effective for a wide range of applications, ranging from the power industry, highway transportation, metal processes, medicine etc., and are particularly more effective in modeling highly non-linear bioprocesses than response surface methodology.16
Among a large number of available training algorithms in ANN, the back propagation (BP) algorithm is the most commonly applied especially for the fermentation process.17–19 However, the BP algorithms have some disadvantages such as: (1) slow convergence speed; (2) falling easily into local optimal solutions; and (3) being sensitive to the noise of the training data set.20 Recently, the Kalman filter and Kalman filter’s variant algorithm have been developed.21–23 The Kalman filter and Kalman filter’s variant algorithm may be effective for noisy behavioral problems because of their inherent ability to suppress noise.20
After the model is established, decision making techniques are needed to obtain the final optimal solution in multi-objective optimization. In general, optimization of multiple objectives aims at capturing the multiple and conflicting solutions, implying that it is difficult to work out a solution that optimizes all objectives at the same time.24 The genetic algorithm (GA) is a stochastic global search algorithm based on the laws of biological evolution and has been widely used for multi-objective optimization.25,26 The combined ANN-GA has been proved to be effective in modeling and optimizing various bioprocesses.27,28 This algorithm is able to solve smooth or non-smooth optimizations with or without bound and linear constraints, and it does not require functions to be differentiable or continuous. There are many modalities of the GA such as the niched Pareto genetic algorithm (NPGA),29 the non-dominated sorting genetic algorithm II (NSGA-II),30 the strength Pareto evolutionary algorithm (SPEA), and the rank-density based genetic algorithm (RDGA). Among them, the non-dominated sorting genetic algorithm II, which is a further improvement of NSGA, is one of the most powerful multi-objective optimization algorithms and much work about this algorithm has been done.31–34
To the authors’ knowledge, the fermentation process of wheat germ has never been modeled using a neural network. In the current study, artificial neural network models were developed with two suitable training algorithms for predicting the amounts of flavonoids and methoxy-ρ-benzoquinone + 2,6-dimethoxy-ρ-benzoquinone, as two objectives. After obtaining a more accurate model, optimal parameters were worked out by applying ANN combined with NSGA-II, thus the production of MBQ + DMBQ and flavonoids could be improved simultaneously under the guidance of the optimized parameters.
2. Materials and methods
2.1. Chemical characterization of wheat germ
The raw wheat germ was obtained from Yihai Kerry Food Industry Co, LTD (Kunshan, China). It was stored at −18 °C in polyethylene pouches prior to use. Proteins were determined using the Kjeldahl method. Moisture, fat, fibre, ash and total carbohydrates were determined according to the Approved Methods of the American Association of Cereal Chemists (AACC, 2003).2.2. Strain and cultivation
The strain S. cerevisiae (RC212, Huankai Biological Technology Co., Ltd, Guang Zhou, China) was cultivated on a YPD (1% yeast extract, 1% peptone, 2% glucose, 2% agar) medium in a 500 ml Erlenmeyer flask under aerobic conditions with chloramphenicol (0.01%) to avoid bacterial growth. The Erlenmeyer flask was incubated in a controlled incubator shaker at 30 °C. Yeast culture growth was measured with plate counting. An inoculum of ca. 2 × 107 cfu per ml was inoculated in a 500 ml Erlenmeyer flask with 200 ml of tap water and 10 g of wheat germ. After fermentation, each sample was freeze-dried and then kept in the refrigerator.2.3. Determination of flavonoids
The extraction of flavonoids was carried out using a method previously reported with modification.35 Briefly, 0.5 g samples were treated with 50 ml of chilled acidified methanol (methanol![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 M HCl, 90
1 M HCl, 90![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 10, v/v). The extract was stirred for 15 min using a magnetic stirrer and then centrifuged at 2500g for 10 min. The filtered supernatant was concentrated by evaporation below 50 °C and reconstituted to a volume of 10 ml with chilled acidified methanol (methanol
10, v/v). The extract was stirred for 15 min using a magnetic stirrer and then centrifuged at 2500g for 10 min. The filtered supernatant was concentrated by evaporation below 50 °C and reconstituted to a volume of 10 ml with chilled acidified methanol (methanol![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 M HCl, 90
1 M HCl, 90![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 10, v/v).
10, v/v).
The total flavonoid amount in the lyophilized fermentation powder samples was determined using a method described previously with modifications.36 An aliquot (0.5 ml) of the above extract was added to ethanol/ultrapure water/ethanolamine (80/15/5, v/v/v) with 3 ml of diphenyl borinic acid ethanolamine ester (DPBA) and vortex-mixed for 5 min. The absorbance was measured at 410 nm using a microplate reader (Bio Tek Instruments, Inc., Winooski, Vermont, USA) against a blank solution containing 70% (v/v) methanol. The measurements were performed in triplicate. Quantification was performed using a standard curve of quantitative catechin solutions and expressed as mg of catechin equivalents per 100 g of sample.
2.4. Determination of methoxy-ρ-benzoquinone and 2,6-dimethoxy-ρ-benzoquinone
Ten grams of lyophilized sample were dissolved in 250 ml of double distilled water and extracted three times by shaking with 100 ml of CHCl3. CHCl3 layers were collected and washed three times with distilled water, then dried over anhydrous Na2SO4. The filtrate was evaporated to a dry condition using a vacuum evaporator at 30 °C. The dry material was dissolved into the mobile phase and then filtered through a 0.22 μm PTFE filter aid. 20 μl of filtrate was injected into the HPLC column and measured by HPLC, using a previous method with modifications.12 The HPLC system was equipped with a C-18 (5 μm, 250 × 4.6 mm) column (Macherey-Nagel, Germany) and a UV 900 detector operating at 275 nm. The mobile phase was a 20% acetonitrile![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 80% water (v/v) mixture. The flow rate was 0.7 ml min−1 and the column oven was set to 30 °C. MBQ and DMBQ (Sigma) were dissolved in the mobile phase as references. The identification of the peaks was confirmed by retention time. The MBQ and DMBQ concentrations were extrapolated from pure standards. All quantifications were determined in triplicate. Results are expressed as mg g−1 of dry matter (d.m.).
80% water (v/v) mixture. The flow rate was 0.7 ml min−1 and the column oven was set to 30 °C. MBQ and DMBQ (Sigma) were dissolved in the mobile phase as references. The identification of the peaks was confirmed by retention time. The MBQ and DMBQ concentrations were extrapolated from pure standards. All quantifications were determined in triplicate. Results are expressed as mg g−1 of dry matter (d.m.).
2.5. Modeling of the artificial neural network
A total of 81 experiments were used in the current study. Four independent variables viz., agitation speed (40–200 rpm), initial pH (4–8), fermentation temperature (24–40 °C), and fermentation time (5–45 h), were used to construct a 4-8-2 ANN topology to validate the relationship between the four parameters, the flavonoid and MBQ + DMBQ, as shown in Fig. 2. The pH was adjusted by adding dilute H2SO4 (NaOH) to the tap water, before adding S. cerevisiae inoculum and wheat germ. The Erlenmeyer flasks containing the fermentation broth were then subjected to incubation (ZQTY-50, Zhichu Instrument CO., Ltd, Shanghai, China). The incubator was adjusted to the required agitation speed and fermentation temperature.A neural net which is a connected parallel architecture, contains an input layer of neurons (independent variables), one or more hidden layers and an output layer of neurons (dependent variables),37 as shown in Fig. 2. The output function can be represented according to the following expression:
| f2(x, whidden, woutput) = woutputf1(whiddenx + biashidden) + biasoutput |
2.6. Sensitivity analysis
To determine the significance of the independent variables (agitation speed, initial pH, fermentation temperature, and fermentation time) to the two objectives (flavonoid and MBQ + DMBQ), sensitivity analysis was implemented using Garson’s modified algorithm, which has been used in fermentology studies.39 The product sum of connected weights obtained from the trained ANN model was used to determine the significance of the independent variables. The relative importance (Rik) of the input variables is defined by:where Rik is the relative importance of input variable i to output k; N is the number of neurons in the input layer; M is the number of neurons in the hidden layer and K is the number of neurons in the output layer; Wij is the connected weight between input node i and hidden node j; Wjk is the weight between hidden node j and output node k; and Srj represents the sum of WijWjk in neuron j of the hidden layer.
2.7. Multi-objective optimization using NSGA-II
When an optimization problem involves more than one objective, the task of finding optimum solutions is known as multi-objective optimization (MOO). In a problem with two objective functions f1(x) and f2(x), solution xi is a Pareto optimal solution only if there is no other solution xj satisfying f1(xj) ≥ f1(xi) and f2(xj) > f2(xi). The solutions xi in this set are called the Pareto set. The objective function values for a given Pareto optimal set are called the Pareto frontier.25,40In this study, optimization is a procedure of finding and comparing feasible solutions until no better solution can be found. For this purpose, non-dominated sorting genetic algorithm II (NSGA-II) was utilized, which has been successfully applied in multi-objective optimization in numerous fields especially complex bio-processes.25,40 The principle of the non-dominated sorting genetic algorithm II (NSGA-II) is described as follows: a randomly-generated initial parent population (Pi) is treated by the fast non-dominated sorting approach to obtain the offspring population Oi. Then the offspring population Oi is combined with the parent population (Pi) to obtain a new population Ri (Ri = Pi + Oi). After Ri is sorted by the fast non-dominated approach and the individual crowded parent comparison is computed, a new parent population Pi+1 is obtained by eliminating the unqualified individual. With circling and eliminating as above, a group of optimum solutions, which are the Pareto optimal set are gained.31 A flow chart of the proposed ANN-NSGA-II is given in Fig. 3.
3. Results and discussion
3.1. Evaluation of BP-ANN and EKF-ANN
The datasets from experiments were divided randomly into two groups: training datasets and test datasets. The test datasets were not used in training the ANN model, but in evaluating the prediction performance of the trained model.27 Matlab 7.0 was used to train the BP-ANN and EKF-ANN models. The mean square error (MSE) between the predicted values and observed values was set at 0.005, and the ANN model training was stopped when the training curve converged to 0.005. The predicted values of the BP-ANN and EKF-ANN models were compared with the original experimental values of test datasets in Fig. 4. As can be seen in Fig. 4, both models were able to predict the flavonoid and MBQ + DMBQ content accurately. Nevertheless, the results indicated that the EKF-ANN model run on the test datasets generated a slightly better correlation (R2 = 0.983 for the flavonoid and 0.974 for MBQ + DMBQ) between experimental values and predicted values of output than the BP-ANN model (R2 = 0.976 for the flavonoid and 0.97 for MBQ + DMBQ). In regression, the coefficient of determination (R2) is a degree of fitting of the regression line approximating the experimental values. Thus, the higher correlation value (R2) confirmed the EKF-ANN model’s ability to predict more accurately than the BP-ANN model.In Fig. 5, the production of the flavonoid and MBQ + DMBQ is plotted versus four fermentation parameters. Two other fermentation parameters in each figure are set at median values. As can be seen in Fig. 5a, increasing the agitation speed from 40 rpm to 200 rpm decreased the flavonoid content. Similarly, increasing the fermentation temperature had a negative effect on the flavonoid content of the fermented wheat germ (Fig. 5b). It can also be noticed that the production of the flavonoid decreased at first and then increased with increasing initial pH, and the production of the flavonoid reached its maximum value of 3.6 mg g−1 in the group with pH = 4, under an agitation speed of 40 rpm (Fig. 5a). From Fig. 5b, it can be observed that increasing the fermentation time (ranging from 5 h to 45 h) firstly increased and then decreased the production of the flavonoid. The bioconversion of the flavonoid was mainly related to S. cerevisiae’s metabolism. On the one hand, S. cerevisiae, which possessed glycosidase activity, was capable of catalyzing the deglycosylation of the flavonoid glycosides, and the aglycones have been proved to be absorbed in humans more easily than the glycosides.41 On the other hand, S. cerevisiae was capable of deconjugating flavonoid aglycones.11,42 Consequently, the increased metabolism of S. cerevisiae may result in the reduction of flavonoids in fermentation. For example, increasing the agitation speed improved the metabolism of S. cerevisiae, thus leading to a decline in the flavonoid content. Similarly, as shown in Fig. 5a and b, over a pH range of 4.0 to 8.0, the minimum flavonoid production was found in the group with pH = 6.0, indicating that 6.0 may be the optimum pH for S. cerevisiae to be present. It could also be concluded that the optimum growth temperature for S. cerevisiae to be present may be 36 °C. Finally, over a range of 5–25 h, the increased flavonoid production may be due to substances being consumed during fermentation. After 25 h, the cleavage of the flavonoid caused by S. cerevisiae led to the decrease in the flavonoid content.
The values of MBQ + DMBQ production predicted by the EKF-ANN model concerning the effect of four fermentation parameters are shown in Fig. 5c and d. As can be seen in Fig. 5c, the production of MBQ + DMBQ increased with increasing agitation speed. This phenomenon might be related to the increasing wheat germ cells’ and S. cerevisiae cells’ permeability with increasing agitation speed, thus releasing more intracellular peroxidase, β-glucosidase and hydroquinone glucosides into the fermentation broth. These hydroquinone glucosides were hydrolysed to the corresponding aglycones by β-glucosidase, and the generated hydroquinones were oxidized to the homologous ρ-benzoquinones by peroxidase.5 The increased β-glucosidase, peroxidase and hydroquinone glucosides enhanced the bioconversion production of MBQ and DMBQ. This feature also provided a direction for increasing the bioconversion production of MBQ and DMBQ: the permeability of wheat germ and S. cerevisiae should be enhanced during fermentation by appropriate treatments, such as ultrasound. From Fig. 5c and d, it can also be observed that the production of MBQ + DMBQ increased at first and then decreased with increasing fermentation temperature or initial pH. This feature indicated that the appropriate fermentation temperature and initial pH might enhance β-glucosidase and peroxidase activities, thus leading to an increased production of MBQ + DMBQ. Finally, Fig. 5d shows a positive correlation between the production of MBQ + DMBQ and the fermentation time (5–40 h). A possible reason was that within 40 h, the peroxidase activity was higher than the activity of Lot6p, which was a oxidoreductase catalyzing the two-electron reduction.43 After 40 h, with increased quinones, S. cerevisiae may leak more Lot6p, which has been shown to defend S. cerevisiae against the cytotoxic effect of quinones by two-electron reduction via a ‘ping-pong bi bi’ mechanism.44 Moreover, the decrease of peroxidase activity after 40 h may also lead to the reduction of MBQ + DMBQ.
3.2. Contribution of each fermentation variable
In the EKF algorithm applied in the training ANN, the weight and bias values (between the input layer and the hidden layer, and those between the hidden layer and the output layer) were randomly initialized and continually adjusted to minimize the MSE.27 The final weight values were able to be utilized to evaluate the contribution of each fermentation parameter to the production of the flavonoid and MBQ + DMBQ.39 By calculating the product sum of the input-hidden and hidden-output layers, it was found that the fermentation time contributed more than the other parameters for the flavonoid, followed by initial pH, temperature and agitation speed (Fig. 6). For the MBQ + DMBQ content, fermentation time impacted most significantly compared to the other parameters, and the order of importance of the other fermentation parameters can be ranked as agitation speed > fermentation temperature > initial pH. | ||
| Fig. 6 Contribution of each fermentation parameter to the production of the flavonoid and MBQ + DMBQ. | ||
3.3. Bi-objective optimization
As mentioned in section 3.2, fermentation time had the greatest influence on the production of both the flavonoid and MBQ + DMBQ. However, there was a distinct negative correlation between the fermentation time and the production of the flavonoid, while fermentation time had a positive effect on the production of MBQ + DMBQ. A similar paradoxical effect could be observed for the agitation speed.Therefore, after succeeding to develop the EKF-ANN model, an effort was made to optimize the fermentation parameters (i.e., agitation speed, initial pH, fermentation temperature and fermentation time) for maximum flavonoid production and maximum MBQ + DMBQ production as a bi-objective problem. In the current optimization work, NSGA-II was applied together with the validated EKF-ANN model as the fitness function. Individuals (first parent population) of higher fitness values were selected with higher probabilities, thus becoming the offspring population through fast non-dominated sorting, calculating the crowding distance and genetic operation (tournament selection, crossover and mutation). The parent population and offspring population were combined, and the combined population was used to generate the next parent population based on fast non-dominated sorting and crowding distance calculations. Circling as above continued until the number of iterations achieved a maximum.
The Pareto-optimal frontiers (best rank) for the optimum flavonoid and MBQ + DMBQ production evaluated over 500 generations with the random seed set at 0.6 are depicted in Fig. 7. Further, results for 600 generations (not shown) were almost identical to those for 500 generations. Another test was made with a random seed of 0.7 which affected the initial population of chromosomes. Although the two random seeds were different, the test showed that the same Pareto-optimal frontiers were generated for different values of random seed. Fig. 7 also clearly shows the conflict between the two objectives, i.e. the production of the flavonoid and MBQ + DMBQ. Any change which increased (decreased) the production of the flavonoid, resulted in the decline (improvement) of the production of MBQ + DMBQ. Similarly, any change which increased (decreased) the production of MBQ + DMBQ, resulted in the decline (improvement) of the production of the flavonoid. This feature once again revealed the necessity for bi-objective optimization in the parameter design of fermentation. At design point A in Fig. 7, the maximum MBQ + DMBQ production was achieved while the production of the flavonoid was at a minimum. This indicates that point A was an optimal solution when the production of MBQ + DMBQ was considered as the single objective. At point B, on the other hand, the maximum flavonoid production was achieved while the minimum MBQ + DMBQ production occurs, indicating that point B was the optimum solution when the production of the flavonoid was regarded as the single objective. The optimum solutions of the two objectives for 10 typical points from the set of non-dominated solutions (the Pareto-optimal set), and the actual data from the designed parameters are listed in Table 1.
| Fermentation parameters | Objectives | |||||
|---|---|---|---|---|---|---|
| Speed (rpm) | pH | Temperature (°C) | Time (h) | Flavonoid (mg g−1) | MBQ + DMBQ (mg g−1) | |
| P1 | 148 | 6.02 | 31.9 | 40.5 | 1.62 | 2.81 |
| P2 | 163 | 5.81 | 32.4 | 39.3 | 1.85 | 2.76 |
| P3 | 122 | 7.02 | 30.6 | 41.1 | 2.12 | 2.69 |
| P4 | 97 | 6.21 | 28.1 | 37.5 | 2.48 | 2.59 |
| P5 | 78 | 5.22 | 27.2 | 39.1 | 2.93 | 2.48 |
| P6 | 77 | 6.98 | 34.2 | 33.8 | 3.29 | 2.36 |
| P7 | 88 | 6.23 | 33.2 | 29.5 | 3.50 | 2.27 |
| P8 | 90 | 5.97 | 32.1 | 28.2 | 3.61 | 2.22 |
| P9 | 42 | 5.12 | 32.3 | 38.3 | 3.77 | 2.12 |
| P10 | 62 | 7.33 | 26.3 | 41.5 | 3.91 | 1.97 |
| A | 142 | 6.09 | 31.6 | 39.8 | 1.46 | 2.85 |
| B | 20 | 4.0 | 24.0 | 24.9 | 4.20 | 0.98 |
| F | 89 | 6.14 | 32.6 | 28.9 | 3.55 | 2.25 |
Each point shown in Fig. 7 on the Pareto frontier was potentially the final optimum design. Nevertheless, it was obvious that it was impossible to maximize the two objectives simultaneously. Consequently, the selection of a optimum point from the Pareto frontier entailed a process of decision making. Various decision making methods have previously been explained thoroughly.45 Among these methods, LINMAP (Linear Programming Techniques for Multidimensional Analysis of Preference) is a popular technique which calculates the Euclidean distance between each point on the Pareto frontier and the ideal point, shown as
Before the LINMAP method can work out the point (ifinal) which has the minimum distance from Fidealj, each objective is Euclidean nondimensionalized using the following equation:
After calculating the Euclidean distance between each point on the Pareto frontier and the ideal point, the equilibrium point was obtained. This non-dominated solution could be considered as the final solution with 3.55 mg g−1 of the flavonoid and 2.25 mg g−1 of MBQ + DMBQ. The design parameters of the final solution (F) are listed in Table 1.
4. Conclusion
In the current paper, the bi-objective fermentation optimization of wheat germ has been carried out. The fermentation parameters (input nodes) were agitation speed, initial pH, fermentation temperature and fermentation time, while the flavonoid production and MBQ + DMBQ production were considered as the bi-objectives (output nodes). A set of Pareto optimal frontier points were worked out using NSGA-II. The bi-objective optimization led to the discovery of trade-off points among the objective functions. Finally, a final set of design points from the non-dominated solutions were obtained by utilizing decision making methods viz. LINMAP. The final results show a 13.34% increase in the flavonoid content and a 7.92% increase in the total content of MBQ and DMBQ simultaneously, compared with the highest values in original design. The present study also demonstrated the potential of EKF-ANN for modeling the process of biochemical fermentation owing to its higher accuracy than conventional BP-ANN modeling.Acknowledgements
This work was co-financed by the National Natural Science Foundation of China (Grant No. 31101384), Qing Lan Project, the National Key Technology R&D Program (2013AA102201), China Postdoctoral Science Foundation (Grant no. 2014M560396), the Key Technologies R&D Program of Jiangsu (BE2014325), Jiangsu Planned Projects for Postdoctoral Research Funds (Grant no. 1402072C), and the Jiangsu province ‘Collaborative Innovation Center for Modern Grain Circulation and Safety’ industry development program.Notes and references
- A. Brandolini, P. Castoldi, L. Plizzari and A. Hidalgo, J. Cereal Sci., 2013, 58, 123–131 CrossRef CAS.
- F. Jakab, Y. Shoenfeld, Á. Balogh, M. Nichelatti, A. Hoffmann, Z. Kahán, K. Lapis, Á. Mayer, P. Sápy, F. Szentpétery, A. Telekes, L. Thurzó, A. Vágvölgyi and M. Hidvégi, Br. J. Cancer, 2003, 89, 465–469 CrossRef CAS PubMed.
- F. B. Roberta, H. Mate, Y. Shoenfeld, I. Gabriela, D. Dmytro, T. F. Rita, V. Csaba, T. Andras, R. Akos and M. Eva, Int. J. Oncol., 2002, 23, 563–570 Search PubMed.
- T. Mueller, K. Jordan and W. Voigt, J. Exp. Clin. Cancer Res., 2011, 42, 42 CrossRef PubMed.
- J. M. Zapata, A. A. Caldéron, R. Muñoz and A. Ros Barceló, Biochimie, 1992, 74, 143–148 CrossRef CAS PubMed.
- S. Soliner, S. Deller, P. Macheroux and B. A. Palfey, Biochemistry, 2009, 48, 8636–8643 CrossRef PubMed.
- S. V. Tokalov, A. M. Abramyuk and N. D. Abolmaali, Nutr. Cancer, 2010, 62, 795–801 CrossRef CAS PubMed.
- S. Ronnekleiv-Kelly, M. Nukaya, P. Carney and G. D. Kennedy, Gastroenterology, 2014, 146, S-1023 CrossRef.
- S. Schmidt, S. Rainieri, S. Witte, U. Matern and S. Martens, Appl. Environ. Microbiol., 2011, 77, 1751–1757 CrossRef CAS PubMed.
- S. J. Hur, S. Y. Lee, Y. C. Kim, I. Choi and G. B. Kim, Food Chem., 2014, 160, 346–356 CrossRef CAS PubMed.
- Y. Kim, K. L. Goodner, J.-D. Park, J. Choi and S. T. Talcott, Food Chem., 2011, 129, 1331–1342 CrossRef CAS.
- C. Rizzello, T. Mueller, R. Coda, F. Reipsch, L. Nionelli, J. Curiel and M. Gobbetti, Microb. Cell Fact., 2013, 12, 1–9 CrossRef PubMed.
- K.-W. Jung, D.-H. Kim, H.-W. Kim and H.-S. Shin, Int. J. Hydrogen Energy, 2011, 36, 9626–9631 CrossRef CAS.
- R. Islam, R. Sparling, N. Cicek and D. B. Levin, Int. J. Mol. Sci., 2015, 16, 3116–3132 CrossRef CAS PubMed.
- A. Singh, A. Majumder and A. Goyal, Bioresour. Technol., 2008, 99, 8201–8206 CrossRef CAS PubMed.
- L. V. A. Reddy, Y.-J. Wee, J.-S. Yun and H.-W. Ryu, Bioresour. Technol., 2008, 99, 2242–2249 CrossRef CAS PubMed.
- E. Betiku and A. E. Taiwo, Renewable Energy, 2015, 74, 87–94 CrossRef CAS.
- H. Jiang, G. Liu, X. Xiao, C. Mei, Y. Ding and S. Yu, Microchem. J., 2012, 102, 68–74 CrossRef CAS.
- J. Sharon Mano Pappu, G. K. Vijayakumar and V. Ramamurthy, Bioresour. Technol., 2013, 130, 224–230 CrossRef CAS PubMed.
- H. Yang, J. Li and F. Ding, Neurocomputing, 2007, 70, 625–632 CrossRef.
- D. Andre, A. Nuhic, T. Soczka-Guth and D. U. Sauer, Eng. Appl. Artif. Intell., 2013, 26, 951–961 CrossRef.
- R. Bisoi and P. K. Dash, Applied Soft Computing, 2014, 19, 41–56 CrossRef.
- K. Chen and J. Yu, Appl. Energy, 2014, 113, 690–705 CrossRef.
- E. Asadi, M. G. d. Silva, C. H. Antunes, L. Dias and L. Glicksman, Energ. Build., 2014, 81, 444–456 CrossRef.
- R. Soleimani, N. A. Shoushtari, B. Mirza and A. Salahi, Chem. Eng. Res. Des., 2013, 91, 883–903 CrossRef CAS.
- A. Jamali, P. Ahmadi and M. N. Mohd Jaafar, Appl. Therm. Eng., 2014, 64, 293–306 CrossRef CAS.
- J. Peng, F. Meng and Y. Ai, Bioresour. Technol., 2013, 138, 345–352 CrossRef CAS PubMed.
- T. Sathish and R. S. Prakasham, J. Chem. Technol. Biotechnol., 2010, 85, 50–58 CrossRef CAS.
- C. Zhang, Y. Chen, M. Shi and G. P. Peterson, Appl. Therm. Eng., 2009, 29, 3340–3345 CrossRef.
- Z. Chen, X. Yuan, B. Ji, P. Wang and H. Tian, Energy Convers. Manage., 2014, 84, 390–404 CrossRef.
- J. Zhang, H. Zhu, C. Yang, Y. Li and H. Wei, Energy Convers. Manage., 2011, 52, 538–546 CrossRef.
- F. Hajabdollahi, Z. Hajabdollahi and H. Hajabdollahi, Applied Soft Computing, 2012, 12, 3648–3655 CrossRef.
- M. M. Etghani, M. H. Shojaeefard, A. Khalkhali and M. Akbari, Appl. Therm. Eng., 2013, 59, 309–315 CrossRef CAS.
- J. Yazdi and S. A. A. Salehi Neyshabouri, Environ. Model. Software, 2014, 60, 31–44 CrossRef.
- H. Ti, Q. Li, R. Zhang, M. Zhang, Y. Deng, Z. Wei, J. Chi and Y. Zhang, Food Chem., 2014, 159, 166–174 CrossRef CAS PubMed.
- L. N. Pietrzak and F. W. Collins, J. Cereal Sci., 1996, 23, 85–91 CrossRef CAS.
- Y. Mustafa, G. Jaid, A. Alwared and M. Ebrahim, Environ. Sci. Pollut. Res., 2014, 21, 7530–7537 CrossRef CAS PubMed.
- H. R. Maier and G. C. Dandy, Math. Comput. Model., 2001, 33, 669–682 CrossRef.
- L. López-Rosales, J. J. Gallardo-Rodríguez, A. Sánchez-Mirón, A. Contreras-Gómez, F. García-Camacho and E. Molina-Grima, Bioresour. Technol., 2013, 146, 682–688 CrossRef PubMed.
- F. C. Lee, G. P. Rangaiah and A. K. Ray, Biotechnol. Bioeng., 2007, 98, 586–598 CrossRef CAS PubMed.
- R. Di Cagno, F. Mazzacane, C. G. Rizzello, O. Vincentini, M. Silano, G. Giuliani, M. De Angelis and M. Gobbetti, J. Agric. Food Chem., 2010, 58, 10338–10346 CrossRef CAS PubMed.
- A. M. Aura, K. A. O’Leary, G. Williamson, M. Ojala, M. Bailey, R. Puupponen-Pimiä, A. M. Nuutila, K. M. Oksman-Caldentey and K. Poutanen, J. Agric. Food Chem., 2002, 50, 1725–1730 CrossRef CAS PubMed.
- S. Sollner, R. Nebauer, H. Ehammer, A. Prem, S. Deller, B. A. Palfey, G. Daum and P. Macheroux, FEBS J., 2007, 274, 1328–1339 CrossRef CAS PubMed.
- S. Sollner, M. Durchschlag, K.-U. Fröhlich and P. Macheroux, FEMS Yeast Res., 2009, 9, 885–891 CrossRef CAS PubMed.
- G. Liu, Renewable Sustainable Energy Rev., 2014, 31, 611–621 CrossRef.
| This journal is © The Royal Society of Chemistry 2016 |