Molecular and biochemical characterization of a new thermostable bacterial laccase from Meiothermus ruber DSM 1279†
D. C. Kalyani,
L. Munk,
J. D. Mikkelsen and
A. S. Meyer*
Center for BioProcess Engineering, Dept. of Chemical and Biochemical Engineering, Technical University of Denmark, Building 229, DK-2800 Kgs. Lyngby, Denmark. E-mail: am@kt.dtu.DK
First published on 21st December 2015
Abstract
A new laccase gene (mrlac) from Meiothermus ruber DSM 1279 was successfully overexpressed to produce a laccase (Mrlac) in soluble form in Escherichia coli during simultaneous overexpression of a chaperone protein (GroEL/ES). Without the GroEL/ES protein, the Mrlac overexpressed in E. coli constituted a huge amount of the total cellular protein, but the enzyme was localized in the insoluble fraction with no activity in the soluble fraction. Co-expression of the Mrlac with the E. coli GroEL/ES drastically improved proper folding and expression of active Mrlac in the soluble fraction. Spectroscopic analysis of the purified enzyme by UV/visible and electron paramagnetic resonance spectroscopy confirmed that the Mrlac was a multicopper oxidase. The Mrlac had a molecular weight of ∼50 kDa and exhibited activity towards the canonical laccase substrates 2,2′-azino-bis(3-ethylbenzothiazoline-6-sulphonic acid) (ABTS), syringaldazine (SGZ), and 2,6-dimethoxyphenol (2,6-DMP). Kinetic constants Km and kcat were 27.3 μM and 325 min−1 on ABTS, 4.2 μM and 106 min−1 on SGZ, and 3.01 μM and 115 min−1 on 2,6-DMP, respectively. Maximal enzyme activity was achieved at 70 °C with ABTS as substrate. In addition, Mrlac exhibited a half-life for deactivation at 70 °C and 75 °C of about 120 min and 67 min, respectively, indicating that the Mrlac is intrinsically thermostable. Finally, Mrlac was efficient in catalyzing the removal of 2,4-dichlorophene (DCP) in aqueous solution, a trait which makes the enzyme potentially useful for environmentally friendly applications.
Introduction
Laccases (EC 1.10.3.2) belong to the multicopper oxidase (MCO) protein family and are produced by fungi, bacteria, plants, and insects. Laccases couple the one-electron oxidation of various phenolic substrates to the reduction of molecular oxygen to form water.1 Due to the use of oxygen as electron acceptor and no need for expensive co-factors, laccases find applications in several biotechnological processes including dye decoloration,2 bioremediation,3 and have also been suggested to be potentially useful in lignocellulosic biofuel processing4 and lignin modification.5To date, our understanding of laccases mainly originates from studies on fungal enzymes. However, identification and characterization of laccases from bacterial sources may provide for novel functionalities. Although the production yields and redox potential of bacterial laccases are lower than those of fungal laccases, the bacterial laccases have numerous advantageous properties that are more favorable for industrial applications compared to fungal laccases.6 Most of the reported bacterial laccases have thus been found to possess distinctive properties, such as being highly thermotolerant, maintaining high levels of activity in neutral to alkaline conditions,7 and having high halide tolerance.8 In contrast, the activity of fungal laccases decays rapidly at high temperatures and pH values.9 Moreover, bacterial laccases are suitable for overproduction in heterologous hosts like Escherichia coli, and their expression level, stability and catalytic properties are far easier to improve by directed evolution compared to their fungal counterparts.6,10 Thermostability is considered a key prerequisite for maintaining catalytic robustness of laccases or other enzymes in many industrial processes.11 The most well-known bacterial laccase is CotA (an endospore coat protein) from Bacillus subtilis which has high thermostability.12 Other thermostable bacterial laccases have been found in Thermus thermophilus,13 Thermobaculum terrenum,14 B. clausii, and B. coagulans.15
Chlorophenols have been widely used for a long time in wood preservation, anticorrosive rust production, fungicides, and pesticides.16 Chlorophenols are generally recognized as xenobiotic pollutants that enter the environment through various industrial operations such as pulp bleaching, pesticide production, chlorination of water, etc.17 and industrial waste water containing chlorophenols is harmful to human health as a consequence of its potential carcinogenic and mutagenic activities.18,19 Because of their carcinogenicity, toxicity and persistence, chlorophenols are listed among top priority control pollutants by the US EPA and the EU.20 Processes for removing chlorophenols usually involve physical, chemical, electrochemical and/or biological methods.21,22 Biological degradation of chlorophenols offers advantages including high specificity, low cost and a more environmentally friendly profile than its physical or chemical counterparts.17,23
In the present paper we report the identification, expression, purification, and biochemical characterization of a new multi copper oxidase from the thermophilic bacterium Meiothermus ruber. As the active holoenzyme production was difficult using E. coli (DE3), and most of the over-expressed recombinant enzyme aggregated in the inclusion body, we attempted to gain large yields of soluble and active enzyme through optimization of expression conditions. The expression yields in soluble active form were notably improved by chaperone (GroEL/GroES9 (plasmid pGro7)) assisted expression in microaerobic condition at 20 °C. The purified laccase was also tested for its ability to remove the chlorinated pesticide 2,4-dichlorophene (DCP).
Materials and methods
Materials
All reagents were of analytical grade and purchased from commercial sources. Laccase from Trametes versicolor (TvLac, EC 1.10.3.2), 2,2′-azino-bis(3-ethylbenzothiazoline-6-sulphonic acid) (ABTS), 2,6-dimethoxyphenol (2,6-DMP), syringaldazine (SGZ) and DCP were purchased from Sigma-Aldrich (St Louis, MO, USA).Reagents for the polymerase chain reaction (PCR), Ex-Taq DNA polymerase, genomic DNA extraction kit, and pGEM-T easy vector were purchased from Promega (Madison, USA). The pET22b(+) expression vector and plasmid isolation kit were purchased from Qiagen (Hilden, Germany). Oligonucleotide primers were obtained from Macrogen (Netherlands).
Strains and culture conditions
Meiothermus ruber DSM 1279 was obtained from DSMZ (Braunschweig, Germany). E. coli strain DH5α (F−ϕ80dlacZΔM15 Δ(lacZYA-argF) U169 recA1 endA1 hsdR17 (rk−, mk+) supE44 thi-1 gyrA relA1), E. coli BL21 (DE3) [F−ompT hsdSB (rB− mB−) gal dcm rne131 (DE3)], E. coli BL21 (DE3) pLysS [F−ompT gal hsdSB (rB− mB−) dcm gal λ(DE3), pLysS], Tuner™ (DE3) [F−ompT hsdSB (rB− mB−) gal dcm lacY1 (DE3)] were purchased from Novagen (Darmstadt, Germany). E. coli strain C41 (DE3) [F−ompT gal hsdSB (rB− mB−) dcm lon λ(DE3)] was obtained from Avidis (Saint-Beauzire, France) and chaperone competent cell pGro7/BL21 (DE3) was purchased from TaKaRa Bio, Inc. (Otsu, Japan). M. ruber DSM 1279 was cultivated overnight in potato P-YE thermus medium (potato starch 5 g l−1, peptone 5 g l−1, yeast extract 0.2 g l−1) at 60 °C and 180 rpm.Cloning and expression optimization of the M. ruber laccase (Mrlac)
Genomic DNA from M. ruber was extracted by using the Promega Wizard Genomic DNA purification kit. Amplification of the gene was performed by PCR with the primers 5′-catatgATGAAGCTAACCCGACGC-3′ and 5′-gaatccTTATTC-GGCTTCTACCTGC-3′ using genomic DNA of M. ruber as the template. NdeI and EcoRI endonuclease recognition sites are shown in lowercase. The amplified PCR product with flanking restriction sites was first cloned into pGEM-T easy vector and then transformed into E. coli DH5α. Plasmid DNA (with insert) and pET22b+ were digested with restriction enzymes NdeI and EcoRI to release the insert (mrlac) and to create a nick in the expression vector, respectively. The mrlac gene released from pGEM-T vector was ligated with pET22b+ vector to give pET22b-Mrlac, a plasmid that was under the control of the T7 promoter. The sequence of the mrlac insert in the resulting pET22b-Mrlac plasmid was verified by sequencing (Macrogen, Netherlands). The mrlac gene was cloned in pET22b+ and transformed into expression strains: E. coli BL21 (DE3), E. coli BL21 (DE3) pLysS, Tuner™ (DE3), E. coli C41 (DE3) and pGro7/BL21 (DE3). The overnight culture in Luria-Bertani (LB) medium was diluted to 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 100 with fresh media containing ampicillin (100 mg ml−1) and, when required, chloramphenicol and arabinose (34 mg ml−1 and 0.5 mg ml−1), respectively, and incubated at 30 °C with shaking (180 rpm). After cells grew to an OD600 of 0.5–0.6, the cells were induced with 0.1 mM isopropyl-β-D-thiogalactoside (IPTG) and supplemented with 0.2 mM CuSO4. The cells were then incubated at 25 °C in semi-microaerobic conditions (by turning off the shaker 4 h after induction). Cells were harvested after 20 h by centrifugation (8000 × g, 15 min, and 4 °C) and the pellets suspended in 20 mM potassium phosphate buffer, pH 7.6, with 1 mM protease inhibitor, phenylmethanesulfonyl fluoride (PMSF), and then sonicated on ice. Disrupted cells were removed by centrifugation (18
100 with fresh media containing ampicillin (100 mg ml−1) and, when required, chloramphenicol and arabinose (34 mg ml−1 and 0.5 mg ml−1), respectively, and incubated at 30 °C with shaking (180 rpm). After cells grew to an OD600 of 0.5–0.6, the cells were induced with 0.1 mM isopropyl-β-D-thiogalactoside (IPTG) and supplemented with 0.2 mM CuSO4. The cells were then incubated at 25 °C in semi-microaerobic conditions (by turning off the shaker 4 h after induction). Cells were harvested after 20 h by centrifugation (8000 × g, 15 min, and 4 °C) and the pellets suspended in 20 mM potassium phosphate buffer, pH 7.6, with 1 mM protease inhibitor, phenylmethanesulfonyl fluoride (PMSF), and then sonicated on ice. Disrupted cells were removed by centrifugation (18![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 × g, 15 min, and 4 °C) and the level of protein expression was checked on SDS-PAGE. The soluble and active expression was obtained only in chaperone competent cells pGro7/BL21 (DE3). Further, to obtain the best condition of expression with high amounts of soluble and active protein, expression conditions were optimized with regard to IPTG concentration (0.1–1 mM), CuSO4 (0.1–1 mM) and at microaerobic conditions in different temperatures (18, 20, 25 and 30 °C).
000 × g, 15 min, and 4 °C) and the level of protein expression was checked on SDS-PAGE. The soluble and active expression was obtained only in chaperone competent cells pGro7/BL21 (DE3). Further, to obtain the best condition of expression with high amounts of soluble and active protein, expression conditions were optimized with regard to IPTG concentration (0.1–1 mM), CuSO4 (0.1–1 mM) and at microaerobic conditions in different temperatures (18, 20, 25 and 30 °C).
Purification
After expression, cultures were harvested by centrifugation (8000 × g, 15 min, and 4 °C). The cell pellets were resuspended in potassium phosphate buffer (20 mM, pH 7.6) containing 0.1 mM PMSF. Cell disruption was performed by sonification on ice, and the cell debris was removed by centrifugation (8000 × g, 30 min, 4 °C). Afterwards, the supernatant was incubated for 30 min at 65 °C, and denatured proteins were removed by centrifugation (10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 × g, 15 min, 4 °C). Purification was performed on an Äkta explorer FPLC-system (GE Healthcare, München, Germany). Ten milliliters of diluted protein solution were loaded on a 5 ml pre-equilibrated Q-Sepharose (GE Healthcare, München, Germany) column. The column was washed with five column volumes of 20 mM potassium phosphate buffer, pH 7.6. Laccase was eluted in a stepwise gradient with 0–1 M NaCl in 20 mM potassium phosphate buffer, pH 7.6. Fractions containing laccase activity (as measured using the ABTS assay) were pooled and concentrated by ultrafiltration (membrane cutoff 30 kDa, Millipore, USA). Protein concentration was determined using the Bradford assay with bovine serum albumin used as a standard. Sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) was done in a Minigel-Twin (Biometra, Goetttingen, Germany) using a 10% polyacrylamide running gel.
000 × g, 15 min, 4 °C). Purification was performed on an Äkta explorer FPLC-system (GE Healthcare, München, Germany). Ten milliliters of diluted protein solution were loaded on a 5 ml pre-equilibrated Q-Sepharose (GE Healthcare, München, Germany) column. The column was washed with five column volumes of 20 mM potassium phosphate buffer, pH 7.6. Laccase was eluted in a stepwise gradient with 0–1 M NaCl in 20 mM potassium phosphate buffer, pH 7.6. Fractions containing laccase activity (as measured using the ABTS assay) were pooled and concentrated by ultrafiltration (membrane cutoff 30 kDa, Millipore, USA). Protein concentration was determined using the Bradford assay with bovine serum albumin used as a standard. Sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) was done in a Minigel-Twin (Biometra, Goetttingen, Germany) using a 10% polyacrylamide running gel.
UV-visible and EPR spectra of the Mrlac protein
The UV-visible absorption spectrum was obtained at room temperature in 50 mM citrate phosphate buffer (pH 7.5) using a Shimadzu UV 3100 spectrophotometer. EPR spectra were measured with a Bruker ESP-106 spectrometer. Purified enzyme was added to a 4 mm Wilmad EPR tube, and frozen in liquid nitrogen. The following parameters were applied: 9.65 GHz microwave frequency, 10 mW microwave power, 10 G modulation amplitude, 100 kHz modulation frequency, 328 ms time constant, and 82 ms conversion time.Biochemical characterization Mrlac
The activity of Mrlac was tested in 50 mM McIlvaine's buffer (pH 2.6–8) and 50 mM Tris–HCl (pH 8–9) at room temperature following the oxidation of the substrates 2,2′-azino-bis(3-ethylbenzothiazoline-6-sulphonic acid) (ABTS (ε = 36![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 M−1 cm−1 at 420 nm)), 2,6-dimethoxyphenol (2,6-DMP (ε = 27
000 M−1 cm−1 at 420 nm)), 2,6-dimethoxyphenol (2,6-DMP (ε = 27![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 500 M−1 cm−1 at 469 nm)), and syringaldazine (SGZ (ε = 65
500 M−1 cm−1 at 469 nm)), and syringaldazine (SGZ (ε = 65![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 M−1 cm−1 at 530 nm)). The rates of substrate oxidation were determined by measuring the absorbance increase at the respective wavelengths, and the molar extinction coefficients (ε) were obtained from the literature.24,25 The activity of purified laccase towards the ABTS substrate was examined at 20–80 °C and the optimal pH value. For testing thermal stability, Mrlac and Tvlac (benchmark) were incubated at 70 °C, 75 °C and 80 °C for 0 to 480 min. After incubation, samples were immediately used to measure residual activities with ABTS, as described above, and the half-time of activity loss was determined by non-linear regression using the equation of exponential decay (a(t) = a0
000 M−1 cm−1 at 530 nm)). The rates of substrate oxidation were determined by measuring the absorbance increase at the respective wavelengths, and the molar extinction coefficients (ε) were obtained from the literature.24,25 The activity of purified laccase towards the ABTS substrate was examined at 20–80 °C and the optimal pH value. For testing thermal stability, Mrlac and Tvlac (benchmark) were incubated at 70 °C, 75 °C and 80 °C for 0 to 480 min. After incubation, samples were immediately used to measure residual activities with ABTS, as described above, and the half-time of activity loss was determined by non-linear regression using the equation of exponential decay (a(t) = a0![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) exp(−t
exp(−t![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) ln
ln![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2/T1/2)) by OriginPro 8.5 (OriginLab Corporation, Northampton, MA, USA). Data were collected in triplicate.
2/T1/2)) by OriginPro 8.5 (OriginLab Corporation, Northampton, MA, USA). Data were collected in triplicate.
Kinetic constants of Mrlac were determined with the substrates 2,6-DMP, ABTS and SGZ. Kinetic constants were calculated by the Michaelis–Menten method using the GraphPad Prism 5 program.
Effects of metal ions on the activity of purified Mrlac
The effects of various metal ions, at a concentration of 2 mM, on purified Mrlac activity were evaluated by separately adding different metal ions such as Cu2+ (copper sulfate), Mg2+ (magnesium chloride), Zn2+ (zinc chloride), Mn2+ (manganese sulfate), Fe3+ (ferric chloride), Ba2+ (barium chloride), Ni2+ (nickel chloride). Laccase activity determined in the absence of any metal ion was measured as a control (100%). Activities were measured at optimum pH and room temperature.Homology modeling and validation
The 3D homology model of Mrlac was generated using the Build Homology Models (MODELER) module in Discovery Studio 4.0 (DS 4.0; Accelrys Software Inc., San Diego, CA). The crystal structure of Thermus thermophilus (PDB entry 2XU9) was used as a template.26 Comparative modeling was performed to generate the most probable structure of the query protein by alignment with the template sequences. Sequence identity between the target and the template was found to be 65% according to BLAST parameters. The fit of the models' sequence in their current 3D environment was evaluated by Profile-3D Score/Verify Protein as implemented in DS 3.1. The discrete optimized protein energy score in MODELER was also calculated to determine the quality of the protein structures. In order to confirm consistency of the model for docking studies, model validation was carried out using PROCHECK.The removal of 2,4-dichlorophene (DCP)
DCP stock solutions were dissolved in 10% acetonitrile. The assay was done with various concentrations of laccase (0.5–1.5 U ml−1) and DCP (0.3–0.6 mM) in 50 mM citrate phosphate buffer (pH 5.5) at 50 °C. The mixture was incubated for various time periods under continuous vibration (450 rpm). Controls for all experiments included boiled enzymes and no enzyme.The degradation of 2,4-DCP was monitored by colorimetric assay.27 The assay solution was composed of diluted sample (50.0 ml), buffer solution (5.0 ml, 100 mM aqueous ammonia), potassium ferricyanide reagent (1.0 ml, 80 mg ml−1 K3Fe(CN)6 in water), and 4-aminoantipyrene (4-AAP) solution (1.0 ml, 2% 4-AAP in water). The resulting compound was a quinone type dye with a characteristic absorption peak at 510 nm. Absorbance was used to calculate the concentrations of DCP according to a calibration curve. Data collection was controlled by the program Tecan i-control, version 1.5.14.0 (Tecan). The removal efficiency was calculated with the following formula:
Results and discussion
Identification, cloning and overexpression of the recombinant Mrlac
To identify amino acid sequences corresponding to MCOs with laccase activity, we performed a blast search in the ExPASy proteomic server. The search for putative MCOs within diverse bacterial genomes revealed a vast number of candidate genes, particularly in Bacilli, Streptomycetes, and Pseudomonads. We focused on MCOs with laccase activity from thermostable organisms. The open reading frame (ORF) of Meiothermus ruber (accession no. YP003506102) encodes a 499 amino acid protein with a calculated molecular mass 55.4 kDa and a pI of 8.8.Multiple sequence alignment, using ClustalW (http://www.ebi.ac.uk/clustalw/), showed that Mrlac contains four histidine-rich copper motifs which are characteristic of MCOs (Fig. 1). Further, a relatively high sequence identity to thermostable T. thermophilus laccase (65%) was found. The pET vector is known as one of the most powerful expression systems. The Mrlac gene was amplified from genomic DNA of M. ruber DSM 1279 and inserted into the pET22b+ expression vector under control of the T7-phage promoter. A high level of expression of Mrlac was achieved (Fig. S1a†), and distinct bands were detected with the expected molecular mass (∼50 kDa). However, almost the entire enzyme was collected into the insoluble fraction as an aggregate without activity in the soluble fraction. A common limitation of recombinant bacterial laccase production is the formation of insoluble protein aggregates known as inclusion bodies, and it is a well-known phenomenon in heterologous laccase production, particularly in E. coli.10 In order to resolve the problem of inclusion body formation, E. coli cells were co-transformed with the pET vector containing the Mrlac gene (T7 promoter, Amp resistance) and the pGro7 vector encoding for GroEL and GroES chaperones (araB promoter, Chl resistance). The use of chaperone co-expression to improve recombinant protein production has been reported in the literature.28–30 The co-expression of GroEL/ES prevented Mrlac from forming inclusion bodies in the cell as demonstrated here. The semi-microaerobic conditions also favored copper accumulation in the cells.31,32 In addition to adding a copper supplement to the medium we lowered the incubation temperature to 20 °C. The synergistic effect of co-expression of GroEL/ES and the semi-microaeration led to expression of an active, soluble enzyme. In this way, expression of active enzyme was significantly improved. The highest yields of active Mrlac enzyme (measured on ABTS) were obtained with the GroEL/ES co-expression after 20 h at 25 °C in LB medium supplemented with 0.2 mM CuSO4 and 0.7 mg ml−1 arabinose under static conditions (Fig. S1a and b†).
Purification and spectroscopic characterization of the secreted protein
After expression of Mrlac under the optimized conditions, cells were harvested and lysed by sonification. To simplify the subsequent purification steps, the supernatant was heated to 65 °C for 30 min. While most of the E. coli proteins precipitated, no loss in activity was observed for Mrlac.The protein was further purified using anion exchange chromatography (Q-Sepharose). SDS-PAGE analysis of the enzyme showed a band with a molecular weight of ∼50 kDa (Fig. S1c†). When isolated, Mrlac from M. ruber has the typical deep blue color generally observed for MCOs and the UV-visible (UV-vis) spectrum of the protein showed a consistent intense absorption band at ∼610 nm, originating from a Cys-S to T1 Cu(II)33 charge transfer transition (Fig. 2a).
Additionally in the UV-vis spectrum, a broad shoulder was observed at ∼330 nm, indicative of an antiferromagnetically coupled T3 binuclear Cu(II) pair. The EPR spectrum (Fig. 2b) of the purified enzyme, had contributions from two paramagnetic centers: (g∥ = 2.22/A∥ = 82 × 10−4 cm−1, and g∥ = 2.24/A∥ = 192 × 10−4 cm−1) T1 and T2 Cu(II)'s, respectively. These results are consistent with reported EPR spectrum for B. subtilis CotA34 and other LMCOs,14,31 suggesting a similar chemical structure around the copper sites.
Biochemical properties of Mrlac
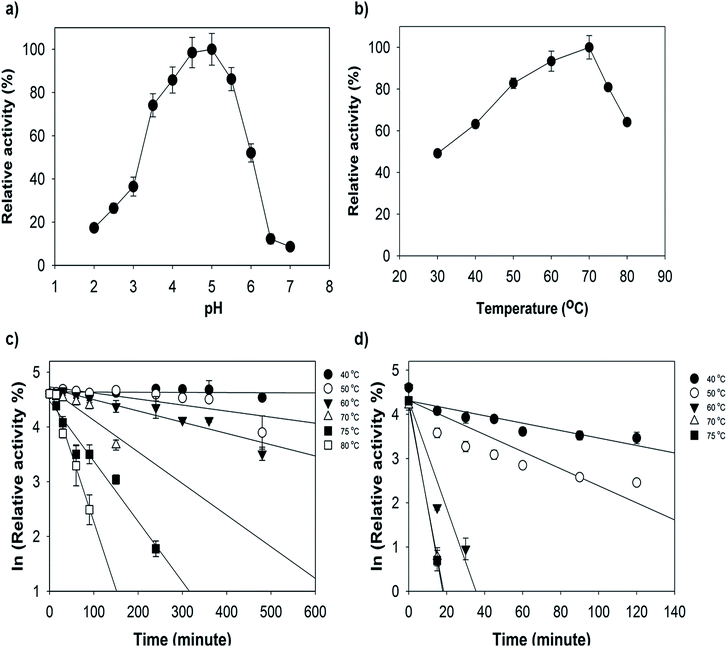
Enzyme activities towards nonphenolic (ABTS) and phenolic (2,6-DMP and SGZ) substrates were determined. The pH optima of the Mrlac activities were determined to be approximately pH 5 for ABTS (Fig. 3a), pH 8 for 2,6-DMP and pH 7.5 for SGZ (data not shown). The pH optima of Mrlac for the oxidation of ABTS, SGZ and 2,6-DMP are similar to those of other bacterial laccases.12,13,35 As shown in Table 1, Michaelis–Menten parameters were determined for ABTS, 2,6-DMP and SGZ at their optimal pH and 40 °C. Remarkably, Km values of Mrlac towards 2,6-DMP and SGZ were much lower than those reported for CotA from B. subtilis36,37 and other bacterial laccases.8,15,35,38,39| Substrate | εmax (M−1 cm−1) | pH | Km (μM) | kcat (min−1) | kcat/Km (s−1 μM−1) |
|---|---|---|---|---|---|
| ABTS | 36![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 000 |
5 | 27.3 | 325 | 0.19 |
| 2,6-DMP | 27![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 500 500 |
8.0 | 3.01 | 115 | 0.63 |
| SGZ | 65![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 000 |
7.5 | 4.2 | 106 | 0.42 |
Fig. S2† shows the predicated laccase-aided modification of non-phenolic (ABTS) and phenolic (2,6-DMP) substrate during oxidative catalysis. As shown in Fig. S2a,† in first step of oxidation, ABTS is freely oxidized by laccase to the cation radical ABTS+˙ (forms intense green-blue colour).40 The cation radical can be oxidized further to the dication (ABTS2+) (Christopher et al., 2014).41 However, 2,6-DMP undergoes a single electron oxidation to produce 2,6-dimethoxy-phenoxyl radical species (forms intense red-brown colour) (Fig. S2b(I)†), that resonate with corresponding para-radical species (Fig. S2b(II)†). It has been postulated phenolic substrates release a proton and electron when they are oxidized to radical products by the enzymatic action of a laccase.42 These radicals can be react non-enzymatically to other phenolic radical or molecular oxygen to form quinones (2,6-dimethoxy-p-benzoquinone). Further, oligomers of 2,6-DMP can form through two oxidative couplings namely C–C dimers or C–O dimers.42,43
To determine the temperature dependence of the enzyme activity, turnover studies were conducted with ABTS at ∼70 °C (Fig. 3b). Furthermore, thermostability was investigated by pre-incubation of the Mrlac and Tvlac at 70, 75, and 80 °C. As seen in Fig. 3c, Mrlac showed moderate stability at elevated temperatures, the half-life for deactivation was 120 min at 70 °C, 67.1 min at 75 °C and 28.1 min at 80 °C.
However, the fungal laccase (Tvlac) used in this study was not stable at elevated temperatures, the half-life for deactivation was 5.7 min at 60 °C, 2.7 min at 70 °C and 2 min at 75 °C (Fig. 4d). The optimal temperature of Mrlac at approximately 70 °C is identical to that observed for the CotA-laccase of B. subtilis,12 but lower than the 92 °C determined for the laccase of the thermophilic bacteria T. thermophilus.13
The thermostability experiment (Fig. 3c) showed that the new Mrlac is one of the most stable bacterial laccase with a half-life 120 min at 70 °C, in contrast to fungal laccase (Fig. 3d) and other reported bacterial laccases.35,44,45
The stability of Mrlac is an excellent feature, especially when compared to fungal laccases. However, the T. thermophilus laccase with a half-life of 868 min at 80 °C is the most thermostable laccase reported so far.13 The lower thermostability of the Mrlac in comparison to the T. thermophilus enzyme supports results from a previous analysis described by Miyazaki,13 in which parameters known to affect protein thermostability have been outlined. These include proline content (10% and 6.8% for T. thermophilus laccase and Mrlac, respectively) and the aliphatic index values, defined as the relative volume of the protein occupied by aliphatic chains of alanine, valine, isoleucine, and leucine (96.39% and 92.24% for T. thermophilus laccase and Mrlac, respectively).
Effect of metal ions on Mrlac activity
The enzyme activity was enhanced by metal ions such as Cu, Mn, Ba (each at 1 mM), while the metals like Ni, Mg, Fe and Zn reduced the activity considerably at a concentration of 1 mM (Fig. S3†).Previous research has demonstrated that Mn(II) oxidation in several bacteria such as Leptothrix species,46 Pseudomonas putida,47 Pedomicrobium species,48 and Bacillus species49,50 is an enzymatic process, and many MCOs are postulated to be directly involved. Furthermore, it was observed that Mn-oxidizing ability was lost upon the disruption of a gene whose sequence identifies it as a MCO.48 The Mn2+ oxidation driven by Mrlac may also have useful biotechnological applications in the fields of biomaterials, enzyme catalysts, bioremediation, metal recovery, and bioenergy.
Overall structure description of Mrlac
The amino acid sequence of Mrlac was aligned with the laccase of T. Thermophilus (PDB entry 2XU9) using the Align Multiple Sequences module of DS 4.0 (Fig. S5†). A homology model of Mrlac was generated based on the crystal structure of the laccase from T. Thermophilus (Fig. S5a†).The root mean square deviation (RMSD) between the model and template was calculated in order to determine the reliability of the model by superimposing the model on the template crystal structure.
The RMSD was estimated to be 0.394 Å on the basis of the C-alpha atoms (Fig. S5a†). A calculated Ramachandran plot suggested that 93.7%, 5.3%, 0.5%, and 0.5% of the residues in the derived model are in, most favored, additional allowed, generously allowed and disallowed regions, respectively (data not shown). Altogether, 99.5% of the residues were placed in the favored and the allowed categories combined.
A model structure with a high percentage of residues in the favored and the allowed categories combined are likely to represent a good protein fold. Thus PROCHECK validated the folding integrity of our model and indicated that the model structure derived from the 2XU9 template was of high quality in terms of protein fold. Laccases are a family of enzymes typically composed of three plastocyanin-like domains (Cu-oxidase super family domains). The sequence alignments indicated that Mrlac features four highly homologous regions, consisting of histidine-rich motifs that constitute the copper-binding sites.12 The four highly conserved regions include “HPxHLHG” and “HCHxxxH” motifs at domain 3 as well as “HxH” and “WYHxH” motifs at domain 1 (x: any amino acids). The three-dimensional structure of Mrlac built by homology modeling suggested that residues in the T1 copper ion site present a planar triangular coordination with two histidines (His427 and His485) and a cysteine (Cys480), while the axial position of the T1 copper consists of Met490 (Fig. S5b†). The T2 copper center is coordinated to His430 and His105, whereas the T3 copper centers are coordinated to three histidine residues each (His417, His432, and His479 for T3 (a); and His107, His145, and His481 for T3 (b)) (Fig. S4c–e†).
Degradation of DCP using Mrlac
The interest in laccases has increased in recent years because of its ability to oxidize a variety of xenobiotic compounds in the presence or absence of synthetic or natural redox mediators.51Unlike peroxidases, oxidation using laccases does not require hydrogen peroxide, which is harmful to the environment. Therefore, laccases are considered to be more promising for enzymatic removal of phenol-compounds from polluted wastewater.20 In the present work, the degradation of DCP by Mrlac was studied, and the kinetic data were fitted to an intermediate model (Fig. 4a). The kinetic curves followed a first-order reaction equation: C = C0![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) exp(−k1t), in which C0 is the initial concentration of the reactant, C is the concentration of reactant at time t, and k1 is the reaction constant of a first-order reaction. All the correlation coefficients (R2) were in the range 0.96–0.98. The kinetic reaction constant, k1, which is related to the degradation of DCP, was 0.0020, 0.0051 and 0.015 min−1, for laccase concentrations of 0.5, 1, and 1.5 U ml−1, respectively. The removal percentage of DCP reached ∼92% within 180 min with 1.5 U ml−1 laccase, so for further study the amount of laccase was set as 1.5 U ml−1. Fig. 4b shows the reaction of Mrlac with DCP concentrations in the range 0.4–0.6 mM. At 0.4 mM, DCP was completely removed from the solution over 240 min, while 60 and 38% of the 0.5 and 0.6 mM DCP were removed over 300 min, respectively. The HPLC analysis of sample collected at the beginning of reaction showed one major peak for 2,4-DCP at retention time 3.40 (Fig. S6a†). As, the reaction progressed the concentration of 2,4-DCP in reaction mixture decreased and emergence of additional peaks were observed due to degradation at retention time 1.80 min, 2.48 min (Fig. S6b†).
exp(−k1t), in which C0 is the initial concentration of the reactant, C is the concentration of reactant at time t, and k1 is the reaction constant of a first-order reaction. All the correlation coefficients (R2) were in the range 0.96–0.98. The kinetic reaction constant, k1, which is related to the degradation of DCP, was 0.0020, 0.0051 and 0.015 min−1, for laccase concentrations of 0.5, 1, and 1.5 U ml−1, respectively. The removal percentage of DCP reached ∼92% within 180 min with 1.5 U ml−1 laccase, so for further study the amount of laccase was set as 1.5 U ml−1. Fig. 4b shows the reaction of Mrlac with DCP concentrations in the range 0.4–0.6 mM. At 0.4 mM, DCP was completely removed from the solution over 240 min, while 60 and 38% of the 0.5 and 0.6 mM DCP were removed over 300 min, respectively. The HPLC analysis of sample collected at the beginning of reaction showed one major peak for 2,4-DCP at retention time 3.40 (Fig. S6a†). As, the reaction progressed the concentration of 2,4-DCP in reaction mixture decreased and emergence of additional peaks were observed due to degradation at retention time 1.80 min, 2.48 min (Fig. S6b†).
Consistent with the findings in other studies using laccase and chlorophenols,52,53 we found that M. ruber laccase could oxidize DCP. Although the highest laccase activity was obtained at 70 °C, chlorophene removal assays were carried out at 50 °C. It seems Mrlac is more stable at 50 °C and the temperature of industrial effluent is typically between 40 and 50 °C. As displayed in Fig. 4a, a high removal rate (0.015 min−1) of DCP was observed at 1.5 U ml−1 Mrlac with ∼92% of the total DCP removed.
Jeon et al.53 reported a maximum degradation efficiency of ∼73% at a pH 5, temperature 30 °C, time 24 h and an enzyme and mediator (cocktail ABTS and vanillin) concentration of 3 U ml−1 and 2 mM respectively. However, Bhattacharya and Banerjee18 reported ∼98% removal efficiency of 2,4-DCP at pH 6, temperature 40 °C within 9 h. In another study Kim and Nicell54 and Murugesan et al.55 used the synthetic mediators ABTS, 1-HBT (1-hydroxybenzotriazole) and TEMPO (2,2′,6,6′-tetramethoxy piperidine-1-oxyl), and the natural phenolic compound syringic acid for triclosan (5-chloro-2-(2,4-dichlorophenoxy)phenol) removal. However, despite their effective removal of triclosan, it was found that the oxidized ABTS radical were highly toxic in the Microtox toxicity test.54 Nevertheless, mediator usage for the degradation is being limited due to their side reaction (with substrate or product) which leads to reduced yield and productivity, high cost and create extra pollution.5,56 It is important to remark that most of the previous chlorophenol biodegradation studies were conducted over ∼9–24, while the Mrlac produced in the present study and under the selected conditions allowed a significant reduction in chlorophenol concentration (0.4 mM) in a shorter reaction time (4 h) and without any mediator.
Conclusions
We found that laccase from M. ruber (Mrlac) can be properly folded and synthesized in E. coli under microaerobic conditions during coexpression with a chaperone protein. After the expression and purification step, its biochemical and kinetic properties were studied. The spectrophotometric characterization (UV-visible and EPR) and sequence alignment of Mrlac showed that the enzyme has all the fingerprint spectral features that are characteristic for laccases like MCOs. The purified Mrlac laccase displayed the ability to oxidize chlorophenol in a reduced reaction time compared to previously discovered laccases without the addition of any meditator.Acknowledgements
This research was partially supported by the Danish National Advanced Technology Foundation via the B21st technology platform (Grant no. 001-2011-4). The authors would like to thank Dr Søren Brander for his help in EPR spectral analysis.References
- S. Jones and E. Solomon, Cell. Mol. Life Sci., 2015, 72, 869–883 CrossRef CAS.
- S. Rangabhashiyam, N. Anu and N. Selvaraju, Res. J. Chem. Environ., 2013, 17, 88–95 CAS.
- P. J. Strong and H. Claus, Crit. Rev. Environ. Sci. Technol., 2011, 41, 373–434 CrossRef.
- T. Kudanga and M. Le Roes-Hill, Appl. Microbiol. Biotechnol., 2014, 98, 6525–6542 CrossRef CAS PubMed.
- L. Munk, A. Sitarz, D. Kalyani, J. D. Mikkelsen and A. S. Meyer, Biotechnol. Adv., 2015, 33, 13–24 CrossRef CAS.
- V. Brissos, L. Pereira, F. D. Munteanu, A. Cavaco-Paulo and L. O. Martins, Biotechnol. J., 2009, 4, 558–563 CrossRef CAS PubMed.
- P. Sharma, R. Goel and N. Capalash, World J. Microbiol. Biotechnol., 2007, 23, 823–832 CrossRef CAS.
- S. Brander, J. D. Mikkelsen and K. P. Kepp, PLoS One, 2014, 9, e99402 Search PubMed.
- P. Baldrian, FEMS Microbiol. Rev., 2006, 30, 215–242 CrossRef CAS PubMed.
- Z. M. Fang, P. Zhou, F. Chang, Q. Yin, W. Fang, J. Yuan, X. C. Zhang and Y. Z. Xiao, PLoS One, 2014, 9(7), e102423 Search PubMed.
- Z. Y. Zhao, J. Liu, M. Hahn, S. Z. Qiao, A. P. J. Middelberg and L. Z. He, RSC Adv., 2013, 3, 22008–22013 RSC.
- L. O. Martins, C. M. Soares, M. M. Pereira, M. Teixeira, T. Costa, G. H. Jones and A. O. Henriques, J. Biol. Chem., 2002, 277, 18849–18859 CrossRef CAS PubMed.
- K. Miyazaki, Extremophiles, 2005, 9, 415–425 CrossRef CAS PubMed.
- S. Brander, J. D. Mikkelsen and K. P. Kepp, J. Mol. Catal. B: Enzym., 2015, 112, 59–65 CrossRef CAS.
- J. Ihssen, R. Reiss, R. Luchsinger, L. Thony-Meyer and M. Richter, Sci. Rep., 2015, 5, 10465 CrossRef CAS.
- L. Yin, Z. Shen, J. Niu, J. Chen and Y. Duan, Environ. Sci. Technol., 2010, 44, 9117–9122 CrossRef CAS PubMed.
- M. Z. Khan, P. K. Mondal and S. Sabir, J. Hazard. Mater., 2011, 190, 222–228 CrossRef CAS PubMed.
- S. S. Bhattacharya and R. Banerjee, Chemosphere, 2008, 73, 81–85 CrossRef CAS.
- S. Li, X. Ma, L. Liu and X. Cao, RSC Adv., 2015, 5, 1902–1909 RSC.
- J. Jia, S. Zhang, P. Wang and H. Wang, J. Hazard. Mater., 2012, 205–206, 150–155 CrossRef CAS PubMed.
- S. Sharma, M. Mukhopadhyay and Z. V. P. Murthy, Sep. Purif. Rev., 2013, 42, 263–295 CrossRef CAS.
- Z.-l. Li, J. Nan, J.-q. Yang, X. Jin, A. Katayama and A.-j. Wang, RSC Adv., 2015, 5, 89157–89163 RSC.
- Y. Wang, X. Chen, J. Liu, F. He and R. Wang, Environ. Sci. Pollut. Res. Int., 2013, 20, 6222–6231 CrossRef CAS PubMed.
- D. Litthauer, M. J. van Vuuren, A. van Tonder and F. W. Wolfaardt, Enzyme Microb. Technol., 2007, 40, 563–568 CrossRef CAS.
- D. Kalyani, S. S. Dhiman, H. Kim, M. Jeya, I. W. Kim and J. K. Lee, Process Biochem., 2012, 47, 671–678 CrossRef CAS.
- H. Serrano-Posada, B. Valderrama, V. Stojanoff and E. Rudino-Pinera, Acta Crystallogr., Sect. F: Struct. Biol. Cryst. Commun., 2011, 67, 1595–1598 CrossRef CAS PubMed.
- R. J. King, K. A. Short and R. J. Seidler, Appl. Environ. Microbiol., 1991, 57, 1790–1792 CAS.
- A. Kondo, J. Kohda, Y. Endo, T. Shiromizu, Y. Kurokawa, K. Nishihara, H. Yanagi, T. Yura and H. Fukuda, J. Biosci. Bioeng., 2000, 90, 600–606 CrossRef CAS PubMed.
- A. de Marco, Nat. Protoc., 2007, 2, 2632–2639 CrossRef CAS PubMed.
- M. Goyal and T. K. Chaudhuri, Int. J. Biochem. Cell Biol., 2015, 64, 277–286 CrossRef CAS PubMed.
- P. Durao, I. Bento, A. T. Fernandes, E. P. Melo, P. F. Lindley and L. O. Martins, J. Biol. Inorg. Chem., 2006, 11, 514–526 CrossRef CAS PubMed.
- M. Mohammadian, M. Fathi-Roudsari, N. Mollania, A. Badoei-Dalfard and K. Khajeh, J. Ind. Microbiol. Biotechnol., 2010, 37, 863–869 CrossRef CAS.
- A. E. Palmer, D. W. Randall, F. Xu and E. I. Solomon, J. Am. Chem. Soc., 1999, 121, 7138–7149 CrossRef CAS.
- E. Solomon, U. Sundaram and T. Machonkin, Chem. Rev., 1996, 96, 2563–2605 CrossRef CAS PubMed.
- K. Koschorreck, S. M. Richter, A. B. Ene, E. Roduner, R. D. Schmid and V. B. Urlacher, Appl. Microbiol. Biotechnol., 2008, 79, 217–224 CrossRef CAS PubMed.
- P. Durao, Z. J. Chen, C. S. Silva, C. M. Soares, M. M. Pereira, S. Todorovic, P. Hildebrandt, I. Bento, P. F. Lindley and L. O. Martins, Biochem. J., 2008, 412, 339–346 CrossRef CAS PubMed.
- N. Gupta and E. T. Farinas, Protein Eng., Des. Sel., 2010, 23, 679–682 CrossRef CAS PubMed.
- M. Gunne and V. B. Urlacher, PLoS One, 2012, 7, e52360 CAS.
- S. Callejon, R. Sendra, S. Ferrer and I. Pardo, Appl. Microbiol. Biotechnol., 2015 DOI:10.1007/s00253-015-7158-0.
- A. Marjasvaara, J. Janis and P. Vainiotalo, J. Mass Spectrom., 2008, 43, 470–477 CrossRef CAS PubMed.
- L. P. Christopher, B. Yao and Y. Ji, Frontiers in Energy Research, 2014, 2(12), 1–13 Search PubMed.
- J. P. Kallio, S. Auer, J. Janis, M. Andberg, K. Kruus, J. Rouvinen, A. Koivula and N. Hakulinen, J. Mol. Biol., 2009, 392, 895–909 CrossRef CAS PubMed.
- Y. Y. Wan, Y. M. Du and T. S. Miyakoshi, Sci. China, Ser. B: Chem., 2008, 51, 669–676 CrossRef CAS.
- G. Diamantidis, A. Effosse, P. Potier and R. Bally, Soil Biol. Biochem., 2000, 32, 919–927 CrossRef CAS.
- A. A. Telke, G. S. Ghodake, D. C. Kalyani, R. S. Dhanve and S. P. Govindwar, Bioresour. Technol., 2011, 102, 1752–1756 CrossRef CAS.
- P. L. A. M. Corstjens, J. P. M. de Vrind, T. Goosen and E. W. d. V. d. Jong, Geomicrobiol. J., 1997, 14, 91–108 CrossRef CAS.
- K. Geszvain, J. K. McCarthy and B. M. Tebo, Appl. Environ. Microbiol., 2013, 79, 357–366 CrossRef CAS PubMed.
- J. P. Ridge, M. Lin, E. I. Larsen, M. Fegan, A. G. McEwan and L. I. Sly, Environ. Microbiol., 2007, 9, 944–953 CrossRef CAS PubMed.
- C. N. Butterfield, A. V. Soldatova, S. W. Lee, T. G. Spiro and B. M. Tebo, Proc. Natl. Acad. Sci. U. S. A., 2013, 110, 11731–11735 CrossRef CAS PubMed.
- J. M. Su, P. Bao, T. L. Bai, L. Deng, F. Liu and J. He, PLoS One, 2013, 8(4), e60573 CAS.
- A. Canas and S. Camarero, Biotechnol. Adv., 2010, 28, 694–705 CrossRef CAS PubMed.
- M. A. Ullah, H. Kadhim, R. A. Rastall and C. S. Evans, Appl. Microbiol. Biotechnol., 2000, 54, 832–837 CrossRef CAS.
- J. R. Jeon, K. Murugesan, Y. M. Kim, E. J. Kim and Y. S. Chang, Appl. Microbiol. Biotechnol., 2008, 81, 783–790 CrossRef CAS.
- Y.-J. Kim and J. A. Nicell, J. Chem. Technol. Biotechnol., 2006, 81, 1344–1352 CrossRef CAS.
- K. Murugesan, Y. Y. Chang, Y. M. Kim, J. Jong-Rok, E. J. Kim and Y. S. Chang, Water Res., 2010, 44, 298–308 CrossRef CAS PubMed.
- A. Kunamneni, S. Camarero, C. Garcia-Burgos, F. J. Plou, A. Ballesteros and M. Alcalde, Microb. Cell Fact., 2008, 7, 32 CrossRef PubMed.
Footnote |
| † Electronic supplementary information (ESI) available. See DOI: 10.1039/c5ra24374b |
| This journal is © The Royal Society of Chemistry 2016 |