A new approach for pyrene bioremediation using bacteria immobilized in layer-by-layer assembled microcapsules: dynamics of soil bacterial community†
Abstract
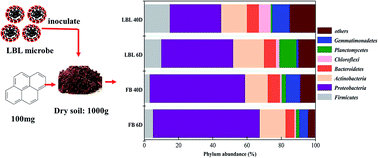
This study reports on the enhanced bioremediation of pyrene (PYR)-contaminated soil resulting from organisms immobilized in layer-by-layer (LBL) assembled microcapsules. The characterization by microscopy indicated that the shape of the microcapsule was like a flake with a diameter of 3–4 μm and that bacteria were encapsulated in the microcapsules. Soil remediation experiments revealed that PYR with an initial concentration of 100 mg kg−1 in dry soil could be 81% removed by an immobilized consortium (107 CFU g−1 in dry soil) in 40 days, while only 42% was removed by the free bacteria. Moreover, the LBL-immobilized cells could cause a significant increase in the biodiversity of the bacterial community, soil enzyme activity and the number of PYR-degrading bacteria in the soil, successfully accounting for accelerated PYR removal. Illumina MiSeq sequencing results showed that Proteobacteria and Actinobacteria were observed as the predominant groups during bioremediation in the LBL groups. The active uncultured bacteria belonged to Xanthomonadaceae, Planococcaceae, Pseudomonas, Mycobacterium, Sphingomonadaceae, Acinetobacter, Flavobacterium, Comamonadaceae, Bacillus, Sphingobacterium, Enterobacteriaceae, and Streptomyces, the latter two classes having rarely been associated with PAH-degrading activity. The results indicated that the LBL microcapsule treatment might be a potential bacteria immobilization option for soil bioremediation.

 Please wait while we load your content...
Please wait while we load your content...