Dual-responsive star-shaped polypeptides for drug delivery†
Wenlong Wang‡
a,
Liang Zhang‡a,
Mengtao Liua,
Yuan Le*a,
Shanshan Lva,
Jiexin Wanga and
Jian-Feng Chenab
aState Key Laboratory of Organic-Inorganic Composites, Beijing University of Chemical Technology, Beijing 100029, P. R. China. E-mail: leyuan@mail.buct.edu.cn
bResearch Center of the Ministry of Education for High Gravity Engineering and Technology, Beijing University of Chemical Technology, Beijing 100029, PR China
First published on 2nd December 2015
Abstract
Core cross-linked star-shaped polypeptides based on poly(L-glutamic acid)-poly(L-phenylalanine-co-L-cystine) copolymer have been successfully synthesized and thoroughly characterized. The star polypeptides can self-assemble to form 50 nm micelles in aqueous medium, which respond rapidly to both pH change within the physiologically relevant pH range and a reduction environment mimicking the intracellular space. Water-soluble doxorubicin hydrochloride and hydrophobic resveratrol are loaded into the star polypeptides micelles through electrostatic and hydrophobic interactions respectively. The drug loading content can be controlled by tuning the composition of the star polypeptides. The in vitro release studies indicate dual sensitivity enabled rapid drug release at pH 5.5 and 10 mM dithiothreitol (DTT), mimicking the intracellular environment. Furthermore, the star polypeptides are biocompatible and interact well with cells in vitro. Confocal fluorescence microscopy and flow cytometry assays show these star polypeptides can be quickly internalized and effectively deliver the drugs into HeLa cells to inhibit cell growth.
1. Introduction
Biodegradable polymeric micelles formed by amphiphilic block copolymers have been receiving a growing interest in the field of drug delivery,1–3 owing to their excellent performance in enhancing drug solubility, prolonging drug circulation time and minimizing side effects.4 Various intelligent polymers in response to internal or external stimuli, such as temperature,5 pH,6,7 light,8 ultrasound,9 and redox potential,10 have been used extensively by drug delivery systems. Among the stimulus-responsive polymers, pH and redox-responsive polymers appear to be the most attractive candidates for targeted and controlled drug delivery because of the significant differences in the glutathione (GSH) concentration and pH between different physiological tissues.11,12 The physiological pH of blood and normal tissues is 7.4, but the extracellular pH in some tumor tissues is 6.8, and the intracellular endo/lysosomal pH ranges from 4.0 to 5.5.13,14 Meanwhile, the GSH concentration is 2–20 μM in blood and normal tissue, but it is around 10 mM in tumor cells.15 Thus, these dual-stimuli responsive polymers have unprecedented control over drug release and delivery to increase efficacy in vitro and in vivo.16–18Although much research has been carried out on polymeric micelles for stimuli-responsive drug delivery systems, their clinical applications are facing tremendous challenges including poor in vivo stability, low drug loading levels and slow drug release.19,20 In comparison with conventional linear polymers, star polymers with three-dimensional architecture exhibit unique properties such as enhanced stability,21 high encapsulation capabilities,22 as well as a large number of internal and peripheral functionalities.23,24 They are expected for potential clinic application. Polypeptides, which are poly(amino acid)s linked by peptide bonds, are unique biodegradable and biocompatible polymers with structures mimicking natural proteins.25 Meanwhile, polypeptides can undergo transitions in response to external stimuli,26,27 and their side chains could be incorporated with various functional moieties for designing intelligent polymer systems.28–30 Therefore, it is interesting to explore the potential of stimuli-responsive star polypeptides for improving drug delivery.
To our knowledge, star polypeptides as dual pH and redox-responsive micelles have not been reported yet. Moreover, the reported responsive polymers carry drugs just through physical interaction or via chemical conjugation despite that both hydrophilic and hydrophobic drugs can be entrapped into the polymeric micelles.31–33 In addition, the process of encapsulation, tunable parameters, and biocompatibility are not well characterized for the application to drug delivery. There is great demand to design smart systems with enhanced functionality and precise molecular control.
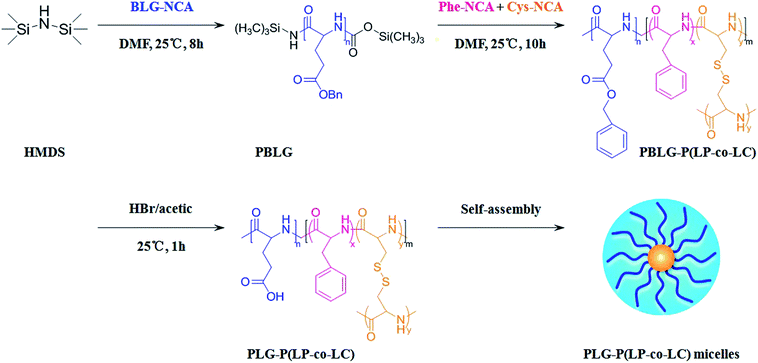
In this work, we designed a novel core cross-linked star polypeptide based on poly(L-glutamic acid)-poly(L-phenylalanine-co-L-cystine) (PLG-P(LP-co-LC)) for dual responsive drug delivery, as shown in Scheme 1. By taking advantage of the star polypeptide properties, PLG-P(LP-co-LC) copolymers are expected to undergo spontaneous self-assembling to micelles in aqueous solutions resulting in a PLG outer corona and hydrophobic PLP/PLC inner core. Doxorubicin hydrochloride (DOX) was selected as a hydrophilic drug and resveratrol (RES) was used as a hydrophobic drug. RES (trans-3,5,4′-trihydroxy-stilbene), a natural polyphenol compound found in grapes and berries,34,35 is well known for its activities of antioxidant, cardioprotection, anti-inflammatory36 as well as anti-cancer properties.37,38 The anionic PLG provides strong electrostatic interaction with cationic DOX, and the hydrophobic PLP serves as a reservoir for RES. The physicochemical properties, drug loading and release performances of the star polypeptides have been investigated. Furthermore, cell cytotoxicity and internalization behavior of the drug loaded star polypeptides have also been explored to evaluate the potential for in vivo drug delivery.
2. Experimental
2.1 Materials
L-Glutamic acid, L-phenylalanine, L-cystine, dithiothreitol (DTT), doxorubicin hydrochloride (DOX) and resveratrol (RES) were purchased from Aladdin Industrial Corporation (Shanghai, China). Benzyl-L-glutamate (BLG) and N-benzyloxycarbonyl-L-cystine (Z-L-cystine) were synthesized according to the literature procedure.39,40 Hexamethyldisilazane (HMDS) (99.9%), anhydrous N,N-dimethylformamide (DMF), 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT), and 4′,6-diamidino-2-phenylindole dihydrochloride (DAPI) were purchased from Sigma-Aldrich (St Louis, MO, USA) and used as received. Tetrahydrofuran (THF), dichloromethane (DCM) and 1,4-dioxane were obtained from Sinopharm Chemical Reagent (Beijing, China) and distilled prior to use. All the other reagents and solvents were analytical grade and used without further purification.2.2 Characterizations
NMR spectroscopy was performed on a Bruker AV III 400 NMR spectrometer using the deuterated solvent as reference and a sample concentration of ca. 20 mg mL−1. FT-IR spectra were recorded on a Bruker VERTEX 70 instrument using the potassium bromide (KBr) method. The carbon, hydrogen, nitrogen and sulfur element contents of the star polypeptides were estimated by Vaio EL cube elemental analysis. Scanning electron microscopy (SEM) measurements were performed on a JEOL JSM-7800F at an accelerating voltage of 10 kV. A drop of the micelles solution (0.05 g L−1) was deposited onto a piece of cover glass and allowed to dry in air at 25 °C before measurements. Dynamic laser scattering (DLS) measurements were performed on a Malvern ZS90 with a vertically polarized He–Ne laser (633 nm) at an angle of 173° and a temperature of 25 ± 0.1 °C. The initial PBLG or PBLG-P(LP-co-LC) concentrations of 10 mg mL−1 in DMF were used then serial dilutions were performed until stable spectra were obtained. All sample solutions were filtered using 0.45 μm filters. Molecular weight and molecular weight distributions were determined using an Agilent 2600 Series gel penetration chromatography (GPC) instrument at 25 °C using DMF with 0.5 M LiBr as the eluent at a flow rate of 1 mL min−1. The standard curve was determined using a series of narrow distribution polystyrene standard samples.2.3 Synthesis of BLG-NCA, Phe-NCA and Cys-NCA
The amino acids were converted to their NCA derivatives using either triphosgene or thionyl chloride, respectively. Benzyl-L-glutamate (1.8 g, 7.6 mmol) was dissolved in 30 mL THF under nitrogen. Triphosgene (0.9 g, 2.9 mmol) was added and the mixture was stirred at 40 °C until the cloudy solution turned clear. The solvent was removed in vacuo and the resulting residue was purified by recrystallization with ethyl acetate and n-hexane to afford γ-benzyl-L-glutamate-N-carboxyanhydride (BLG-NCA) (yield: 95.1%). 1H NMR (400 MHz, CDCl3, δH, ppm): 2.21 (m, 2H, –CH2–), 2.60 (dd, 2H, –CH2COO), 4.45 (t, 1H, –CHN–), 5.15 (s, 2H, –CH2O), 7.35 (m, 5H, ArH–), 6.28 (s, 1H, –NH–). L-Phenylalanine N-carboxyanhydride (Phe-NCA) was synthesized following the same procedure as for the synthesis of BLG-NCA (yield: 91.5%). 1H NMR (400 MHz, CDCl3, δH, ppm): 3.15 (m, 2H, –CH2–), 4.53 (dd, 2H, –CHN–), 5.78 (s, 1H, –NH–), 7.30 (m, 5H, ArH–).Z-L-Cystine (2.0 g, 3.9 mmol) was dissolved in 20 mL 1,4-dioxane. Thionyl chloride (1.2 mL, 16.5 mmol) was added and the mixture was reacted at 55 °C for 3 h. After removal of the solvent in vacuo, the resulting residue was precipitated in DCM to obtain L-cystine-N-carboxyanhydride (Cys-NCA) (yield: 92.3%). 1H NMR (400 MHz, DMSO-d6, δH, ppm) 3.20 (m, 4H, –SCH2CH–), 4.77 (dd, 2H, –CHN–), 9.25 (s, 2H, –NH–).
2.4 Synthesis of star polypeptide PLG-P(LP-co-LC)
The star polypeptide PLG-P(LP-co-LC) was synthesized through a one-step ring-opening polymerization of BLG-NCA, Cys-NCA and Phe-NCA in DMF using HMDS as the initiator. In brief, BLG-NCA (1.0 g, 3.3 mmol) and HMDS (17 μL, 82 μmol) were dissolved in anhydrous DMF (10 mL) in a glove box. The reaction mixture was stirred at room temperature until the NCA’s anhydride peak at 1786 cm−1 disappeared, as determined by FT-IR. The concentration of poly(γ-benzyl-L-glutamate) (PBLG) was diluted to 1 mg mL−1 using DMF (containing 0.1 M LiBr) for measuring the molecular weight of PBLG. Then, Phe-NCA (0.4 g, 2.3 mmol) and Cys-NCA (0.3 g, 1.0 mmol) were added to the remaining PBLG solution, and the reaction was maintained for another 10 h. The polymer solution was precipitated into an excess amount of methanol to afford the resulting star polypeptide PBLG-P(LP-co-LC) (yield: 86.5%). 1H NMR (400 MHz, TFA-d6, δH, ppm): 2.21 (d, 2H, –CHCH2–), 2.48 (s, 2H, –CH2COOH), 4.80 (s, 2H, CH2C6H5), 5.33 (s, H, –CHN), 7.78 (m, 5H, ArH–).Subsequently, star polypeptide PBLG-P(LP-co-LC) (2.0 g) was dissolved in TFA (10 mL) and 4 mL HBr (33% in acetic acid) was added. After stirring for 1 h at room temperature, the mixture was precipitated into an excess amount of ice diethyl ether. After drying under a vacuum, the crude product was purified by dialysis against deionized water (MWCO = 3500 Da), and lyophilized to afford water soluble star polypeptide PLG-P(LP-co-LC), yielding a white solid (yield: 87.2%). 1H NMR (400 MHz, D2O, δH, ppm): 1.92 (d, 2H, –CHCH2–), 2.23 (s, 2H, –CH2COOH), 4.26 (s, H, –CHN–), 7.26 (m, 5H, ArH–).
2.5 Stimuli-responsive behavior of star polypeptides
The blank micelles were prepared by a dialysis method. The star polypeptide PLG-P(LP-co-LC) was dissolved in dimethyl sulfoxide (DMSO), and the solution was added to deionized water under stirring for 24 h. To remove the solvents, the solution was dialyzed against deionized water for 72 h.The pH-responsiveness of PLG-P(LP-co-LC) was examined by analyzing the size and zeta potential of the micelles solution at different pH values using DLS. Furthermore, the disassembly of micelles in response to redox stimuli was examined by DLS and 1H NMR spectroscopy. DTT was dissolved in degassed micelles solution (10 mM DTT) under nitrogen. The obtained solution was stirred at 37 °C, and the size of PLG-P(LP-co-LC) was determined by DLS at different time intervals. After 24 h, the resulting solution was dialyzed and lyophilized for 1H NMR analysis.
2.6 Preparation of drug loaded micelles
The DOX-loaded micelles were prepared according to the following method. In brief, PLG-P(LP-co-LC) (50 mg) was dissolved in distilled water and DOX (10 mg) dissolved in DMSO was added dropwise. After stirring overnight in the dark, the organic solvent and free DOX were removed by dialysis using a dialysis bag (MWCO = 3500 Da) against deionized water for 24 h, and then freeze-dried to obtain the DOX-loaded micelles.The RES-loaded micelles were prepared by a dialysis method. Briefly, 50 mg of PLG-P(LP-co-LC) and 5 mg of RES were dissolved in DMSO, and the solution was added dropwise to the deionized water under stirring. The mixture was maintained at room temperature under stirring for 12 h. The solution was dialyzed against excess deionized water with a dialysis bag (MWCO = 3500 Da) for 24 h, and then filtered through a 0.45 μm pore-sized microporous membrane. The RES-loaded micelles were obtained after lyophilization.
DOX loaded micelles were determined by UV absorption at 480 nm and RES loaded micelles were detected at 306 nm. The drug loading content (DLC, wt%) and drug loading efficiency (DLE, wt%) were calculated according to the following formulas
2.7 In vitro drug release
The release profiles of the drugs from the micelles were investigated in PBS at different pH values (7.4, 6.8 and 5.5) or different concentrations of DTT (0 mM, 5 mM and 10 mM) by the dialysis method. The weighed freeze-dried drug loaded micelles were suspended in 5 mL of the release medium and transferred into a dialysis bag (MWCO = 3500 Da). The release experiment was initiated by placing the dialysis bag into release medium at 37 °C with continuous shaking at 100 rpm. At predetermined intervals, 5 mL of the incubated solution was withdrawn and replaced with equal volume of fresh release medium. The amounts of DOX and RES were determined using a UV-Vis spectrometer (SHIMADZU UV 2600) at 480 nm and 306 nm, respectively. All measurements were conducted in triplicate.2.8 Cell cultures
HeLa cells were cultured in Dulbecco’s modified Eagle’s medium (DMEM) containing 10% fetal bovine serum (FBS), supplemented with 50 U mL−1 penicillin and 50 U mL−1 streptomycin, and incubated at 37 °C in 5% CO2 atmosphere.2.9 Cell viability assays
The in vitro cytotoxicities of blank micelles, free drugs, and drug loaded micelles were assessed with MTT assays. HeLa cells were seeded in 96-well plates at ∼4000 cells per well in 100 μL of DMEM containing 10% fetal bovine serum, supplemented with 50 U mL−1 penicillin and 50 U mL−1 streptomycin, and incubated at 37 °C in 5% CO2 atmosphere for 24 h. The culture mediums were replaced with 200 μL fresh mediums containing different concentrations of star polypeptides (0–500 μg mL−1). The absorbency of the solution was measured on a Multiskan MK3 microplate reader at 490 nm. The cell viability (%) was calculated by (Asample/Acontrol) × 100, where Asample and Acontrol denote the absorbencies of the sample and control wells, respectively. Data are presented as means ± standard deviation (n = 3).2.10 Intracellular drug release
The cellular uptake and intracellular release behaviors of DOX loaded micelles were determined by confocal laser scanning microscopy (CLSM) of the HeLa cells. The cells were seeded on coverslips in 6-well plates with a density of ∼40![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 cells per well in 2 mL of DMEM and cultured for 24 h, and the cells were incubated with DOX loaded micelles at a concentration of 20 μg mL−1 in complete DMEM. After 4 h, 12 h and 24 h incubation, the culture medium was removed and cells were washed three times with PBS. Then, the cells were fixed with 4% paraformaldehyde at room temperature, and the cell nuclei were stained with 4′,6-diamidino-2-phenylindole (DAPI). The treated cells were visualized under a laser scanning confocal microscope (Leica TCS SP5).
000 cells per well in 2 mL of DMEM and cultured for 24 h, and the cells were incubated with DOX loaded micelles at a concentration of 20 μg mL−1 in complete DMEM. After 4 h, 12 h and 24 h incubation, the culture medium was removed and cells were washed three times with PBS. Then, the cells were fixed with 4% paraformaldehyde at room temperature, and the cell nuclei were stained with 4′,6-diamidino-2-phenylindole (DAPI). The treated cells were visualized under a laser scanning confocal microscope (Leica TCS SP5).
The cellular uptake of micelles was also analyzed quantitatively using flow cytometry. HeLa cells were seeded at a density of 2 × 104 cells per well in 6-well plates and incubated for 24 h. After incubating with DOX loaded micelles (at a concentration of 20 μg mL−1) for 4, 12 and 24 h, the cells were washed three times with PBS, harvested and subsequently resuspended in 0.5 mL PBS for flow cytometry analysis (FC500, Beckman Coulter, USA). The instrument was calibrated with non-treated cells (negative control) to identify viable cells, and the cells were determined from a fluorescence scan performed with 1 × 104 cells using the FL1-H channel.
2.11 Statistical analysis
All the values were presented as mean ± standard deviation (SD) of at least three independent measurements. Statistical significance was tested by one-way ANOVA followed by a Student’s t-test for multiple comparison tests. Differences characterized by *P < 0.05 were considered statistically significant. All statistical analysis was performed using SPSS, version 22.3. Results and discussion
3.1 Synthesis and characterization of star polypeptide PLG-P(LP-co-LC)
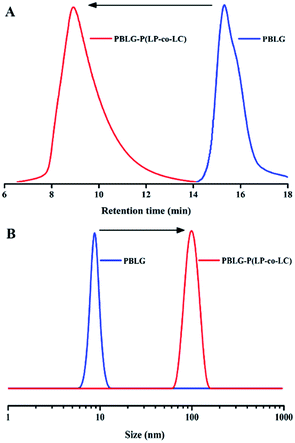
The star polypeptide PLG-P(LP-co-LC) was synthesized through a core cross-linked strategy using sequential ring opening polymerization of BLG-NCA, Phe-NCA and Cys-NCA using HMDS as the initiator, followed by deprotecting the groups in HBr/acetic acid. The structures of the NCA monomers were identified by FT-IR and 1H NMR. Fig. 1S† shows the FT-IR spectra of the NCA monomers with various amino acids. The peaks centered at 1750 and 1850 cm−1 were assigned to the characteristic peak of carbonyl from the anhydride. As shown in Fig. 2SA,† the resonances at 7.35 ppm (peak a) and 6.28 ppm (peak d) were ascribed to the Ar of BLG and NH of NCA. In Fig. 2SB,† the peaks at 4.53 ppm (peak c) and 5.78 ppm (peak b) were assigned to the CH and NH of anhydride. Peak a indicated the presence of benzene in Phe-NCA. Fig. 2SC† shows that the peaks at 9.25 ppm and 3.20 ppm were ascribed to NH and CH2 of Cys-NCA. The 1H NMR and FT-IR spectroscopy confirmed the successful synthesis of the NCA monomers.Then, the polymerization initiated by HMDS was monitored via the FT-IR intensity of the NCA’s anhydride peak at 1787 cm−1;41,42 the disappearance of the NCA’s anhydride peak confirmed the complete consumption of the monomers. Upon addition of Phe-NCA and cross-linker Cys-NCA, the weight average molar mass (Mw) of the polypeptide increased from 7.5 kDa to 932.4 kDa (Fig. 1A), and the size measured by DLS significantly increased from 8.5 nm to 101.3 nm (Fig. 1B). The 1H NMR spectrum of PBLG-P(LP-co-LC) (Fig. 2B) showed the characteristic signals (peak a) of the BLG units, but the signals of the cross-linked core component were not visible due to the reduced segmental mobility.43,44 Furthermore, PLG-P(LP-co-LC) was prepared by removing the benzyloxycarbonyl groups of PBLG-P(LP-co-LC). The disappearance of the ester C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O stretching peak at 1725 cm−1 (Fig. 2A) and the benzyl group signal at 7.36 ppm (Fig. 2C) indicated the complete removing of the protecting group in the polypeptide. The final molar composition ratio of the monomeric repeating units in PLG, PLP and PLC blocks was 40
O stretching peak at 1725 cm−1 (Fig. 2A) and the benzyl group signal at 7.36 ppm (Fig. 2C) indicated the complete removing of the protecting group in the polypeptide. The final molar composition ratio of the monomeric repeating units in PLG, PLP and PLC blocks was 40![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 24
24![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 10 (calculated by elemental analysis).
10 (calculated by elemental analysis).
 | ||
| Fig. 2 FT-IR spectra (A) of star polypeptides PLG-P(LP-co-LC) (a) and PBLG-P(LP-co-LC) (b); 1H NMR of PBLG-P(LP-co-LC) (B, TFA-d6) and PLG-P(LP-co-LC) (C, D2O). | ||
3.2 Stimuli-responsive behavior of star polypeptides
The pH and redox-responsive behaviors of the star polypeptide PLG-P(LP-co-LC) were investigated by DLS and 1H NMR, as shown in Fig. 3 and 4. The zeta potentials of PLG-P(LP-co-LC) significantly increased as the pH decreased, and the carboxyl groups in PLG were in a fully deionized state below pH 5.0, resulting in significant agglomeration.45 Similar pH-dependent agglomeration was also proved by the size (Fig. 3), which dramatically increased to 1306 nm with the decrease of pH from 8 to 3. It is well known that the disulfide linkages are stable under normal physiological conditions but respond to reductive conditions (e.g. GSH, DTT) via reversible cleavage into free thiols.46 The star polypeptide PLG-P(LP-co-LC) could preserve its core–shell structure without DTT. In the presence of DTT (10 mM), a significant change in size was observed, wherein the size gradually increased from 80 to 140 nm in 5 h and over 290 nm in 24 h (Fig. 4A). These results suggest that the disulfide linkages of PLG-P(LP-co-LC) were cleaved by 10 mM DTT so that the core structure of the star polypeptide micelles dissociated which led to the size increase. Meanwhile, the characteristic signals corresponding to the core segments were also observed in the 1H NMR spectrum after cleavage, including signals at 7.2 ppm (C6H5 in PLP) and 2.9–3.2 ppm (–CH2– in PLC) (Fig. 4B). | ||
| Fig. 4 Size (A) and 1H NMR spectrum (B, D2O) of star polypeptide PLG-P(LP-co-LC) response to 10 mM DTT. | ||
3.3 Characterization of drug loaded micelles
The star polypeptide PLG-P(LP-co-LC) with hydrophilic PLG arms and hydrophobic PLP core spontaneously formed core–shell micelles in aqueous solutions. The morphologies of the micelles were studied by SEM (Fig. 5A), which showed that micelles were spherical shaped with an average diameter of around 50 nm. In contrast, the size measured by DLS was 80 nm (Fig. 5D). The smaller values from SEM observations should be due to the dehydration of the micelles during the SEM sample preparation process. Furthermore, DOX and RES were loaded into the micelles by the nanoprecipitation technique. Compared to blank micelles, the size of drug loaded micelles increased and maintained a narrow distribution as shown in Fig. 5D, which confirmed drugs being effectively entrapped into micelles. Meanwhile, the morphologies of the drug loaded micelles were measured by SEM measurements. SEM images revealed that all micelles possessed uniformly spherical morphologies.Then, drug loading capacity (DLC) and efficiency (DLE) were measured and are shown in Table 1. The DLC and DLE values of DOX were higher than those of RES owing to different interactions between the drugs and micelles. After DOX loading, the zeta potentials of micelles changed from −50.1 mV to −19.3 mV due to the carboxyl groups being neutralized, suggesting that the positively charged DOX was captured by the carboxyl groups of PLG through electrostatic attraction.47,48 However, RES was loaded into micelles through aromatic stacking interactions with PLP, and the zeta potentials of RES-loaded micelles displayed a slight change. In order to prove the hypothesis, we adjusted compositions of the polypeptides and investigated the DLC and DLE values of both drugs (Table 1). With the ratio of hydrophilic PLG decreasing, the DLC of DOX reduced from 15.8% to 7.8% and DLE decreased to 65.3%, which was attributed to the weakened electrostatic attraction between DOX and PLG. In contrast, with the ratio of PLP increasing, hydrophobic interaction between RES and the core was enhanced, leading to an increase in the DLC and DLE of RES to 10.1% and 50.5%, respectively.49
| Star polypeptides | Feed ratioa | Resultant ratiob | DLC (%) | DLE (%) | ||
|---|---|---|---|---|---|---|
| DOX | RES | DOX | RES | |||
| a Feed molar ratio of BLG-NCA/Phe-NCA.b Resultant molar ratio of PLG/PLP calculated by elemental analysis. | ||||||
| PLG-P(LP-co-LC)-1 | 1.0/0.7 | 1.0/0.6 | 15.8 | 5.8 | 89.7 | 28.5 |
| PLG-P(LP-co-LC)-2 | 1.0/1.0 | 1.0/0.8 | 13.2 | 6.9 | 81.6 | 34.3 |
| PLG-P(LP-co-LC)-3 | 1.0/1.2 | 1.0/1.0 | 7.8 | 10.1 | 65.3 | 50.5 |
3.4 Release behavior of drug loaded micelles
The DOX and RES release behaviors of the micelles were assessed using a dialysis method at 37 °C in phosphate buffered saline (PBS) at different pH values (7.4, 6.8 and 5.5) and different concentrations of DTT (0 mM, 5 mM and 10 mM). As shown in Fig. 6, the release of encapsulated drugs through a change in pH has been exploited. The release of DOX was greatly influenced by the environmental acidity, about 22% and 71% of DOX were released at pH 7.4 and pH 5.5 after 72 h (Fig. 6A). However, the release of RES followed an opposite trend (Fig. 6B), with only 20% of RES released at pH 5.5 after 72 h. This pH dependent release behavior was attributed to the deionized carboxyl groups, which resulted in weakened electrostatic interaction and strengthened hydrophobic interaction.Meanwhile, the drug release behaviors under reductive conditions were also investigated. RES release was accelerated in PBS with 10 mM DTT (Fig. 6D), about 80% of RES was released in 72 h due to the swelling of the micelles caused by the cleavage of the disulfide bonds. In comparison, the stable electrostatic interaction between the drug and the micelles resulted in only a slight increase in DOX release (Fig. 6C). Furthermore, both drugs rapidly released under simulated endosomal condition (PBS buffer at pH 5.5 with 10 mM DTT), especially in the first 10 h. Within 72 h, over 85% DOX and 72% RES were released. These results indicated that the pH and redox-sensitive stars could effectively release loaded drugs in endosomal environments.
3.5 In vitro cytotoxicity studies
The cytotoxicity of the star polypeptide PLG-P(LP-co-LC) was evaluated using the MTT assay. HeLa cell lines were utilized. As shown in Fig. 7, the viabilities of HeLa were ∼100% at 30 μg mL−1 for 24 h, gradually decreasing with increased concentration of polymer and incubated time. However, the viabilities of HeLa were above 80% at all test concentrations up to 500 μg mL−1, revealing remarkable safety and biocompatibility of the star PLG-P(LP-co-LC).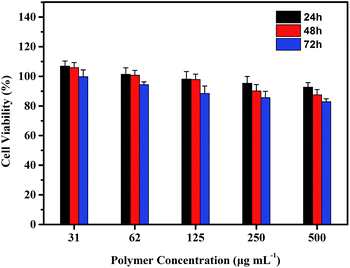 | ||
| Fig. 7 Cell viabilities of HeLa incubated with PLG-P(LP-co-LC) for 24 h, 48 h and 72 h (n = 3, mean ± SD). | ||
To verify the effect of the drug-loaded system, the proliferation inhibition effects of free drugs and drug loaded micelles against HeLa cells were tested using the MTT assay. The cell viability histograms are shown in Fig. 8. After 24 h, 48 h, and 72 h incubation, all free drugs and drug loaded micelles showed dose and time-dependent cell proliferation inhibition behavior. The viabilities of cells incubated with free DOX and DOX-loaded micelles had similar changes in trends. With the increase in drug concentration from 0.6 to 10 μg mL−1, cell viabilities decreased from 80% to 30% for 24 h. With the extension of cultivation time, the drugs were taken more into cells and thus the decrease of cell viabilities results. For RES, cell toxicity was lower than DOX. The IC50 values of free drugs and drug-loaded micelles are summarized in Table 2. The IC50 values of DOX loaded micelles were 2.05, 1.26 and 0.84 μg mL−1 for 24, 48 and 72 h, respectively, which were similar to that of free DOX. The IC50 values of RES loaded micelles were less than that of free RES.
 | ||
| Fig. 8 Cell viabilities of HeLa cells incubated with free DOX (A), free RES (B), DOX-loaded micelles (C), RES-loaded micelles (D) for 24 h, 48 h and 72 h (n = 3, mean ± SD) (*p < 0.05, **p < 0.01). | ||
| Free DOX | Free RES | DOX loaded micelles | RES loaded micelles | |
|---|---|---|---|---|
| 24 h | 2.05 | 33.34 | 2.20 | 22.81 |
| 48 h | 1.26 | 27.31 | 1.32 | 17.13 |
| 72 h | 0.84 | 20.08 | 0.89 | 11.40 |
3.6 Cellular uptake behavior of DOX loaded micelles
The cellular uptake and intracellular release behavior of drug loaded micelles in HeLa cells were investigated by CLSM and flow cytometry. The cellular nuclei were stained with DAPI (blue). DOX (red) was loaded in PLG-P(LP-co-LC) for subcellular observation. As shown in Fig. 9A, DOX fluorescence could be observed in the cell cytoplasm and distributed widely in the cytoplasm after 4 h incubation, indicating that the drug loaded micelles could be successfully internalized by cancer cells via endocytosis. When the incubation period increased to 12 h and 24 h, the cell uptake of drug loaded micelles was enhanced and the fluorescence became even stronger. Meanwhile, the DOX was mainly located in cytoplasm and partly entered into nuclei for DOX loaded treated cells (Fig. 9B and C), suggesting that the DOX had been released and escaped from the endo/lysosomes to the nuclei.The cellular uptake of DOX loaded micelles was further quantitatively evaluated by flow cytometry, and HeLa cells were penetrated with the same processes as revealed by fluorescence microscopy.
As displayed in Fig. 10, the histograms of the pretreated cells incubated with drug loaded micelles shift clearly to the direction of high fluorescence intensity compared with the control cells. Moreover, the mean fluorescence intensity (MFI) of HeLa cells incubated with DOX-loaded micelles for 4 h was 10.8, which increased to 21.6 after 24 h incubation, indicating a time-dependent characteristic. These results demonstrated that the stars can efficiently deliver and release drug into tumor cells. The reason might be attributed to different cell uptake pathways of free drugs and drug loaded micelles, and the controlled release manner of drug-loaded micelles.50
 | ||
| Fig. 10 Flow cytometric profiles and MFI of HeLa cells incubated with DOX loaded micelles for 4 h, 12 h and 24 h. | ||
4. Conclusions
In conclusion, we have successfully synthesized dual responsive core cross-linked star polypeptides based on PLG-P(LP-co-LC) via ring opening polymerization. The micelles self-assembled from the star polypeptide PLG-P(LP-co-LC) have pH and redox sensitivities. DOX and RES were separately loaded into the star polypeptides, and the drug loading capacity can be tuned by adjusting the hydrophilic/hydrophobic ratios of the polypeptides. The release of the drugs was accelerated at low pH or high DTT concentration. CLSM images and flow cytometry analysis indicated a fast internalization of the drug loaded micelles via endocytosis. The cytotoxicity studies showed that the star polypeptides were non-toxic, and drug loaded micelles exhibited high tumor accumulation and superior antitumor efficiency. It is evident that these pH and redox-responsive biocompatible micelles have tremendous potential for targeted and controlled intracellular drug delivery.Acknowledgements
This work was supported by National Key Basic Research Program of China (No. 2015CB932100), National “863” Program of China (No. 2013AA032201), Program for New Century Excellent Talents in University of China (NCET-12-0760).Notes and references
- C. Deng, Y. Jiang, R. Cheng, F. Meng and Z. Zhong, Nano Today, 2012, 7, 467–480 CrossRef.
- K. Kataoka, A. Harada and Y. Nagasaki, Adv. Drug Delivery Rev., 2012, 64, 37–48 CrossRef.
- Z. Ge and S. Liu, Chem. Soc. Rev., 2013, 42, 7289–7325 RSC.
- Y. Bae and K. Kataoka, Adv. Drug Delivery Rev., 2009, 61, 768–784 CrossRef CAS PubMed.
- B. Chang, X. Sha, J. Guo, Y. Jiao, C. Wang and W. Yang, J. Mater. Chem., 2011, 21, 9239–9247 RSC.
- S. P. Nunes, A. R. Behzad, B. Hooghan, R. Sougrat, M. Karunakaran, N. Pradeep, U. Vainio and K. V. Peinemann, ACS Nano, 2011, 5, 3516–3522 CrossRef CAS PubMed.
- J. Su, F. Chen, V. L. Cryns and P. B. Messersmith, J. Am. Chem. Soc., 2011, 133, 11850–11853 CrossRef CAS PubMed.
- D. He, X. He, K. Wang, J. Cao and Y. Zhao, Langmuir, 2012, 28, 4003–4008 CrossRef CAS PubMed.
- H. J. Kim, H. Matsuda, H. Zhou and I. Honma, Adv. Mater., 2006, 18, 3083–3088 CrossRef CAS.
- X. Wang, X. Cai, J. Hu, N. Shao, F. Wang, Q. Zhang, J. Xiao and Y. Cheng, J. Am. Chem. Soc., 2013, 135, 9805–9810 CrossRef CAS PubMed.
- D. L. Liu, X. Chang and C. M. Dong, Chem. Commun., 2013, 49, 1229–1231 RSC.
- F. Shi, J. Ding, C. Xiao, X. Zhuang, C. He, L. Chen and X. Chen, J. Mater. Chem., 2012, 22, 14168–14179 RSC.
- T. Zhou, C. Xiao, J. Fan, S. Chen, J. Shen, W. Wu and S. Zhou, Acta Biomater., 2013, 9, 4546–4557 CrossRef CAS PubMed.
- H. Deng, Y. Zhang, X. Wang, Y. Cao, J. Liu, J. Liu, L. Deng and A. Dong, Acta Biomater., 2015, 11, 126–136 CrossRef CAS PubMed.
- D. J. Phillips, J. P. Patterson, R. K. O’Reilly and M. I. Gibson, Polym. Chem., 2014, 5, 126–131 RSC.
- S. Yoon, W. J. Kim and H. S. Yoo, Small, 2013, 9, 284–293 CrossRef CAS PubMed.
- K. Liang, G. K. Such, Z. Zhu, Y. Yan, H. Lomas and F. Caruso, Adv. Mater., 2011, 23, H273–H277 CrossRef CAS PubMed.
- Y. J. Pan, Y. Y. Chen, D. R. Wang, C. Wei, J. Guo, D. R. Lu, C. C. Chu and C. C. Wang, Biomaterials, 2012, 33, 6570–6579 CrossRef CAS PubMed.
- S. C. Owen, D. P. Y. Chan and M. S. Shoichet, Nano Today, 2012, 7, 53–65 CrossRef.
- F. Meng, R. Cheng, C. Deng and Z. Zhong, Mater. Today, 2012, 15, 436–442 CrossRef CAS.
- G. Kreutzer, C. Ternat, T. Q. Nguyen, C. J. G. Plummer, J. A. E. Manson, V. Castelletto, I. W. Hamley, F. Sun, S. S. Sheiko, A. Herrmann, L. Ouali, H. Sommer, W. Fieber, M. I. Velazco and H. A. Klok, Macromolecules, 2006, 39, 4507–4516 CrossRef.
- C. Kojima, K. Kono, K. Maruyama and T. Takagishi, Bioconjugate Chem., 2000, 11, 910–917 CrossRef PubMed.
- J. T. Wiltshire and G. G. Qiao, Macromolecules, 2008, 41, 623–631 CrossRef CAS.
- B. Helms, S. J. Guillaudeu, Y. Xie, M. McMurdo, C. J. Hawker and J. M. J. Frechet, Angew. Chem., Int. Ed., 2005, 44, 6384–6387 CrossRef CAS PubMed.
- T. J. Deming, Prog. Polym. Sci., 2007, 32, 858–875 CrossRef CAS.
- S. Han, Y. Liu, X. Nie, Q. Xu, F. Jiao, W. Li, Y. Zhao, Y. Wu and C. Chen, Small, 2012, 8, 1596–1606 CrossRef CAS PubMed.
- H. Wu, L. Zhu and V. P. Torchilin, Biomaterials, 2013, 34, 1213–1222 CrossRef PubMed.
- V. K. Kotharangannagari, A. Sanchez-Ferrer, J. Ruokolainen and R. Mezzenga, Macromolecules, 2011, 44, 4569–4573 CrossRef CAS.
- Y. Xiao, H. Hong, A. Javadi, J. W. Engle, W. Xu, Y. Yang, Y. Zhang, T. E. Barnhart, W. Cai and S. Gong, Biomaterials, 2012, 33, 3071–3082 CrossRef CAS PubMed.
- S. J. Lee, K. H. Min, H. J. Lee, A. N. Koo, H. P. Rim, B. J. Jeon, S. Y. Jeong, J. S. Heo and S. C. Lee, Biomacromolecules, 2011, 12, 1224–1233 CrossRef CAS PubMed.
- W. Chen, P. Zhong, F. Meng, R. Cheng, C. Deng and Z. Zhong, J. Controlled Release, 2013, 169, 171–179 CrossRef CAS PubMed.
- S. Aryal, J. Hu and L. Zhang, ACS Nano, 2010, 4, 251–258 CrossRef CAS PubMed.
- J. Dai, S. Lin, D. Cheng, S. Zou and X. Shuai, Angew. Chem., Int. Ed., 2011, 50, 9404–9408 CrossRef CAS PubMed.
- M. Lagouge, C. Argmann, G. Zachary, H. Meziane, C. Lerin, F. Daussin, N. Messadeq, J. Milne, P. Lambert, P. Elliott, B. Geny, M. Laakso, P. Puigserver and J. Auwerx, Cell, 2006, 127, 1109–1122 CrossRef CAS PubMed.
- J. Zhang, Q. Mi and M. Shen, Food Chem., 2012, 131, 879–884 CrossRef CAS.
- T. Shen, X. N. Wang and H. X. Lou, Nat. Prod. Rep., 2009, 26, 916–935 RSC.
- A. Mattarei, M. Azzolini, M. Carraro, N. Sassi, M. Zoratti, C. Paradisi and L. Biasutto, Mol. Pharmaceutics, 2013, 10, 2781–2792 CrossRef CAS PubMed.
- Q. S. Tang, G. W. Li, X. N. Wei, J. Zhang, J. F. Chiu, D. Hasenmayer, D. Z. Zhang and H. Zhang, Cancer Lett., 2013, 336, 325–337 CrossRef CAS PubMed.
- E. R. Blout and R. H. Karlson, J. Am. Chem. Soc., 1956, 78, 941–946 CrossRef CAS.
- C. Gravel, D. Lapierre, J. Labelle and J. W. Keillor, Can. J. Chem., 2007, 85, 164–174 CrossRef CAS.
- H. Lu and J. Cheng, J. Am. Chem. Soc., 2007, 129, 14114–14115 CrossRef CAS PubMed.
- H. Lu and J. Cheng, J. Am. Chem. Soc., 2008, 130, 12562–12563 CrossRef CAS PubMed.
- M. H. Xiong, J. Wu, Y. C. Wang, L. S. Li, X. B. Liu, G. Z. Zhang, L. F. Yan and J. Wang, Macromolecules, 2009, 42, 893–896 CrossRef CAS.
- J. T. Wiltshire and G. G. Qiao, Macromolecules, 2006, 39, 4282–4285 CrossRef CAS.
- S. Takae, K. Miyata, M. Oba, T. Ishii, N. Nishiyama, K. Itaka, Y. Yamasaki, H. Koyama and K. Kataoka, J. Am. Chem. Soc., 2008, 130, 6001–6009 CrossRef CAS PubMed.
- Y. Tian, L. Bromberg, S. N. Lin, T. A. Hatton and K. C. Tam, J. Controlled Release, 2007, 121, 137–145 CrossRef CAS PubMed.
- M. K. Yu, Y. Y. Jeong, J. Park, S. Park, J. W. Kim, J. J. Min, K. Kim and S. Jon, Angew. Chem., Int. Ed., 2008, 47, 5362–5365 CrossRef CAS PubMed.
- M. Li, W. Song, Z. Tang, S. Lv, L. Lin, H. Sun, Q. Li, Y. Yang, H. Hong and X. Chen, ACS Appl. Mater. Interfaces, 2013, 5, 1781–1792 CAS.
- Z. L. Tyrrell, Y. Shen and M. Radosz, Prog. Polym. Sci., 2010, 35, 1128–1143 CrossRef CAS.
- M. Prabaharan, J. J. Grailer, S. Pilla, D. A. Steeber and S. Gong, Biomaterials, 2009, 30, 5757–5766 CrossRef CAS PubMed.
Footnotes |
| † Electronic supplementary information (ESI) available. See DOI: 10.1039/c5ra20972b |
| ‡ These authors contributed equally to this work. |
| This journal is © The Royal Society of Chemistry 2016 |