Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Mixed-monolayer glyconanoparticles for the detection of cholera toxin by surface enhanced Raman spectroscopy†
Jonathan
Simpson
a,
Derek
Craig
b,
Karen
Faulds
a and
Duncan
Graham
*a
aCentre for Molecular Nanometrology, Department of Pure and Applied Chemistry, University of Strathclyde (Technology and Innovation Centre), 99 George Street, Glasgow, G1 1RD, UK. E-mail: duncan.graham@strath.ac.uk
bOptical Manipulation Group, SUPA, University of St. Andrews, St. Andrews, KY16 9SS, UK
First published on 20th November 2015
Abstract
We have produced silver glyconanoparticles for the sensitive (56 ng mL−1), low volume and rapid detection of cholera toxin B-subunit (CTB) in synthetic freshwater (simulating the ion compositions of natural waters in which CTB could be found). This is achieved by monitoring the changes in surface enhanced Raman scattering (SERS) intensity of a Raman reporter bound to the glyconanoparticle surface. The particles selectively aggregate upon interaction with CTB, causing an increase in the measured SERS signal. The particles are designed to mimic the interactions involving the cell surface GM1 ganglioside and CTB. This is achieved by using a combination of polyethylene glycol linkers terminated with either galactose or sialic acid.
Conceptual insightsCholera affects 3–5 million people each year, resulting in 100![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000–120 000–120![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 reported deaths (excluding disasters). Effective water treatment relies on the ability to detect pathogens rapidly and sensitively, allowing a contaminated body of water to be isolated from local consumption promptly. Our newly created nanoparticle sensor can detect the cholera biomarker, cholera toxin B subunit (CTB), sensitively, rapidly and in low volume samples of synthetic freshwater. We demonstrate the first use of mixed carbohydrate monolayers for the rational design of carbohydrate-coated nanoparticles (glyconanoparticles) which effectively mimic the host's intestinal cell surface glycan (GM1). This subsequently allows selective interaction with the target protein, CTB. This research illustrates the first use of glyconanoparticles for the detection of CTB by surface enhanced Raman spectroscopy (SERS). The work presents significant developments in the understanding of multivalent carbohydrate–protein interactions and in the use of SERS-active glyconanoparticles for the detection of a bacterial toxin in samples which mimic natural bodies of water. 000 reported deaths (excluding disasters). Effective water treatment relies on the ability to detect pathogens rapidly and sensitively, allowing a contaminated body of water to be isolated from local consumption promptly. Our newly created nanoparticle sensor can detect the cholera biomarker, cholera toxin B subunit (CTB), sensitively, rapidly and in low volume samples of synthetic freshwater. We demonstrate the first use of mixed carbohydrate monolayers for the rational design of carbohydrate-coated nanoparticles (glyconanoparticles) which effectively mimic the host's intestinal cell surface glycan (GM1). This subsequently allows selective interaction with the target protein, CTB. This research illustrates the first use of glyconanoparticles for the detection of CTB by surface enhanced Raman spectroscopy (SERS). The work presents significant developments in the understanding of multivalent carbohydrate–protein interactions and in the use of SERS-active glyconanoparticles for the detection of a bacterial toxin in samples which mimic natural bodies of water.
|
Nanoparticles have many potential applications in healthcare, including improved drug delivery and biodetection.1–3 Selective and sensitive bio-detectors can be produced by functionalising metallic nanoparticles with biomolecules such as DNA or antibodies.4 Gold and silver nanoparticles functionalised in this way, bind complementary target biomolecules and assemble to form aggregates which is indicated by the appearance of plasmonic-coupling bands in the extinction spectrum.5–8 In addition to biomolecules, Raman active molecules can be coated onto the particles giving rise to surface enhanced Raman scattering.9,10 Target-mediated aggregation of the nanoparticles creates interparticle hot spots of electromagnetic energy which further enhances the Raman signals of the surface bound Raman reporter.11 SERS sensitivity rivals that of UV-visible extinction and fluorescence spectroscopy.12,13 Additionally, the molecularly specific Raman signals can allow for multiplexed detection by using multiple Raman reporters which is important for the analysis of multi-analyte samples.14–16 SERS nanoparticle biosensing has been applied to the detection of biomolecules (including DNA and antigens) along with the imaging and mapping of cells and their associated substructures.13,15,17 In addition to oligonucleotides and proteins, nanoparticles can be functionalised with carbohydrates and associated species for use in glyconanotechnology. This includes enzyme inhibition, oncology and the detection of lectins (carbohydrate binding proteins) or toxins related to disease.18–21 Carbohydrates, present extensively on cell surfaces, provide multifunctional binding sites involved in crucial endogenous processes as well as in pathogen establishment.22–24
Vibrio cholerae bacteria express lectins as AB toxins on the pili which are key to bacterial establishment and subsequent infection.24 The B subunit (CTB) is multifunctional, binding simultaneously to both the galactose and sialic acid moieties of the ganglioside, GM1, expressed on the surface of intestinal cells.24 Unlike the interaction between DNA base pairs or a peptide and protein, those between toxins and their related gangliosides rely on multiple unique species to stabilise the binding pocket simultaneously.19 The individually weak carbohydrate–protein interactions are strengthened by this multivalency.25,26
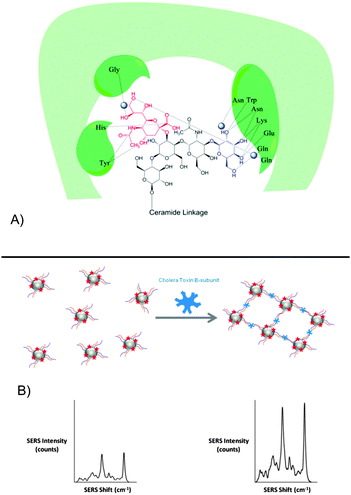
The mixed-carbohydrate particles described herein, mimic these extensive and complex interactions and have enabled the generation of glyconanoparticles which detect the CTB bacterial target both selectively and sensitively and in samples which simulate the environment in which the toxin could be found, namely synthetic freshwater samples which contain the ionic components of natural bodies of water. The approach for generating CTB-reactive particles involved deconstruction of the GM1 ganglioside to identify the major components in CTB binding. Galactose is a major component of the binding between the toxin and GM1 (see Fig. 1A), however, alone this interaction is weak (Gal–CTB Kd = 52 mM).26,27 By simultaneously binding sialic acid in a symbiotic interaction, as is the case for GM1, the binding pocket is significantly stabilised (GM1–CTB Kd = 0.3 nM).24
Previous research on CTB-reactive glyconanoparticles includes the development of lactose-coated gold nanoparticles for the detection of CTB at 54 nM (3 μg ml−1).18 Our novel, mixed-monolayer galacto-sialonanoparticles (GSNPs) allow for detection of CTB with greater sensitivity than previously achieved because of the GM1-mimicking surface and by using SERS for detection.
50 nm colloidal silver nanoparticles, prepared using ethylenediaminetetraacetic acid (EDTA) reduction, were used to produce the biosensor. The Raman molecule used in analysis was the benzotriazole dye molecule 4-((1H-benzo[d][1,2,3]triazol-6-yl)diazenyl)-3,5-dimethoxyphenol (RB1) previously prepared by our group.28 The carbohydrates were subsequently tethered to the nanoparticle surface via thiolated polyethylene glycol (PEG) linkers. PEG provides steric protection that helps discourage aggregation by increasing the minimum interparticle distance in the colloid. Additionally, non-specific binding of biological or chemical species present in a sample is reduced. PEGylated galactose and sialic acid were produced by amide coupling (see ESI,† Section 6 and Fig. S2 for details) and used to functionalise the RB1-coated silver nanoparticles with GM1 mimics.
The SERS-active GSNPs were characterised by UV-visible extinction spectroscopy, dynamic light scattering and gel electrophoresis (see ESI,† Table S4 and Fig. S3). The surface coverage-activity relationship was evaluated by functionalising the particles with galactose and sialic acid linkers of different lengths and altering the ratio between the linkers. While the PEG coating, together with size of CTB, leaves space between the particles upon CTB binding, plasmonic coupling has been shown to occur up to a distance of 70 nm.29,30 It was therefore proposed that upon aggregation of GSNPs with CTB, an increased SERS response would be measured. The GSNPs were incubated with 80 nM CTB for 5 minutes and the resulting aggregation measured by monitoring changes in SERS intensity of the RB1 peak at 1364 cm−1 (see Fig. 1B).
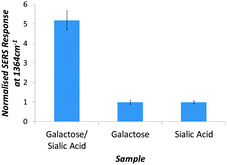
The importance of both sialic acid and galactose in binding GM1 is demonstrated in Fig. 2. The greatest SERS intensity change for the glyconanoparticles is noted with the mixed galactose/sialic acid (15![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1) monolayer. There is little interaction and hence low SERS increase when using sialic acid or galactose as the sole surface coating.
1) monolayer. There is little interaction and hence low SERS increase when using sialic acid or galactose as the sole surface coating.
Our investigations reveal a maximum normalised SERS enhancement of 5.2 at a PEGylated carbohydrate ratio (galactose to sialic acid) of 15![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 (see Table 1), where the two carbohydrate linker types are of different lengths (PEG12Gal and PEG18Sia). The extension of the sialic acid from the nanoparticle surface with respect to the galactose encourages maximal binding and hence aggregation. This arises from the minimal steric interference from surrounding galactose molecules with sialic acid. The reduced amount of sialic acid on the surface with respect to the galactose indicates the importance of isolated galactose/sialic acid–CTB interactions. The interaction, aggregation and hence SERS enhancement is reduced when the sialic acid is extended from the nanoparticle surface at an equivalent length to the galactose. While nanoparticles provide a multivalent platform that can strengthen individually weak carbohydrate–lectin/toxin interactions, they can also hinder binding if the surface coating is too crowded. Binding is discouraged as the spacing between repeating galactose and sialic acid pairs may not match the toxin shape or space between the toxin binding units. Additionally, closely-packed carbohydrates may hydrogen bond, reducing protein-binding. This provides an explanation for the reduced SERS enhancement in both the case of the PEG12Gal/PEG12Sia and PEG18Gal/PEG18Sia. Additionally in the case of the later, the increase in length of both PEG chains increases the distance between aggregated nanoparticles. The magnitude of SERS enhancement is distance dependent and increasing the interparticle distance with the longer PEG chains generates weaker hot spots, resulting in a lower enhancement compared with those particles coated with the shorter PEG12Gal.
1 (see Table 1), where the two carbohydrate linker types are of different lengths (PEG12Gal and PEG18Sia). The extension of the sialic acid from the nanoparticle surface with respect to the galactose encourages maximal binding and hence aggregation. This arises from the minimal steric interference from surrounding galactose molecules with sialic acid. The reduced amount of sialic acid on the surface with respect to the galactose indicates the importance of isolated galactose/sialic acid–CTB interactions. The interaction, aggregation and hence SERS enhancement is reduced when the sialic acid is extended from the nanoparticle surface at an equivalent length to the galactose. While nanoparticles provide a multivalent platform that can strengthen individually weak carbohydrate–lectin/toxin interactions, they can also hinder binding if the surface coating is too crowded. Binding is discouraged as the spacing between repeating galactose and sialic acid pairs may not match the toxin shape or space between the toxin binding units. Additionally, closely-packed carbohydrates may hydrogen bond, reducing protein-binding. This provides an explanation for the reduced SERS enhancement in both the case of the PEG12Gal/PEG12Sia and PEG18Gal/PEG18Sia. Additionally in the case of the later, the increase in length of both PEG chains increases the distance between aggregated nanoparticles. The magnitude of SERS enhancement is distance dependent and increasing the interparticle distance with the longer PEG chains generates weaker hot spots, resulting in a lower enhancement compared with those particles coated with the shorter PEG12Gal.
| Coverage type | Normalised SERS enhancement at 1364 cm−1 | |||
|---|---|---|---|---|
1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1a 1a |
3![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1a 1a |
15![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1a 1a |
30![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1a 1a |
|
| a The enhancement is a ratio of the RB1 signal at 1364 cm−1 prior to aggregation with 80 nM CTB to the same signal measured 5 minutes after CTB addition in HEPES buffer, pH 7.4. The laser excitation wavelength used is 514.5 nm with a 1 second acquisition, 3 accumulations and 3 replicates measured. | ||||
| PEG12Gal/PEG12Sia | 1.2 (±0.16) | 2.0 (±0.20) | 3.5 (±0.19) | 3.0 (±0.21) |
| PEG12Gal/PEG18Sia | 0.9 (±0.10) | 1.7 (±0.22) | 5.2 (±0.49) | 3.3 (±0.51) |
| PEG18Gal/PEG12Sia | 0.9 (±0.06) | 0.9 (±0.09) | 0.9 (±0.06) | 1.0 (±0.07) |
| PEG18Gal/PEG18Sia | 0.9 (±0.20) | 1.7 (±0.15) | 1.8 (±0.07) | 2.3 (±0.14) |
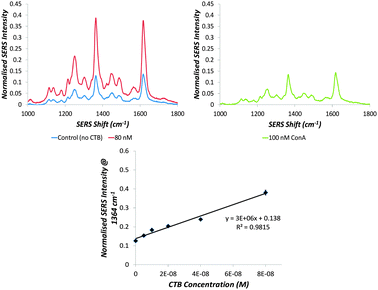
The selectivity of the particles is demonstrated in Fig. 3. Incubating the GSNPs with 100 nM of the glucose-specific plant lectin, Concanavalin A (which interacts specifically with gluconanoparticles, see ESI,† Fig. S6) causes no aggregation and hence gives no appreciable change in SERS intensity even after 30 minutes of incubation. The CTB limit of detection (1 nM or 56 ng mL−1) achieved by using the GSNPs is 50 times greater than that achievable by UV-visible extinction spectroscopy (54 nM or 3 μg mL−1).18 This falls within the recommended detection range of 10 pg mL−1 to 100 ng mL−1 and matches the limits of WHO approved tests including infant rabbit and coagglutination assays, demonstrating the relevancy of this detection level.31,32
The mixed-monolayer glyconanoparticles presented herein react rapidly with CTB in a selective and sensitive manner, giving results in 5 minutes. A low volume is required to aggregate the nanoparticles (10 μL CTB sample in 200 μL of sensor). Previous research has demonstrated that CTB can be measured in both patient and water samples, however this involves PCR or antibody-based methods with multiple assay steps required for eventual detection.33,34 The capability of the rapid and sensitive nanoparticle sensor described herein has been demonstrated in synthetic freshwater samples. By developing our proof-of-concept, we aim to generate a working CTB assay for use in patient samples. Additionally, the availability of portable Raman spectrometers could allow for field testing with the GSNPs. A similar preparation strategy could be employed to generate particles which mimic the G1b gangliosides (GT1b, GD1b and GQ1b) for the detection of tetanus and botulism toxins by SERS.
The authors acknowledge the Carnegie Trust for the Universities of Scotland for funding. DG acknowledges support from the Royal Society through a Wolfson Research Merit Award.
Notes and references
- C. R. Steven, G. A. Busby, C. Mather, B. Tariq, M. L. Briuglia, D. A. Lamprou, A. J. Urquhart, M. H. Grant and S. V. Patwardhan, J. Mater. Chem. B, 2014, 2, 5028–5042 RSC.
- R. Rojanathanes, A. Sereemaspun, N. Pimpha, V. Buasorn, P. Ekawong and V. Wiwanitkit, Taiwan. J. Obstet. Gynecol., 2008, 47, 296–299 CrossRef PubMed.
- L. E. Jamieson, A. Jaworska, J. Jiang, M. Baranska, D. J. Harrison and C. J. Campbell, Analyst, 2015, 140, 2330–2335 RSC.
- A. Lesniewski, M. Los, M. Jonsson-Niedziolka, A. Krajewska, K. Szot, J. M. Los and J. Niedziolka-Jonsson, Bioconjugate Chem., 2014, 25, 644–648 CrossRef CAS PubMed.
- C. A. Mirkin, R. L. Letsinger, R. C. Mucic and J. J. Storhoff, Nature, 1996, 382, 607–609 CrossRef CAS PubMed.
- C. Zhang, Z. Zhang, B. Yu, J. Shi and X. Zhang, Anal. Chem., 2001, 74, 96–99 CrossRef.
- D. C. Hone, A. H. Haines and D. A. Russell, Langmuir, 2003, 19, 7141–7144 CrossRef CAS.
- S. Schlücker, Angew. Chem., Int. Ed., 2014, 53, 4756–4795 CrossRef PubMed.
- D. Graham, G. McAnally, J. C. Jones and W. Ewen Smith, Chem. Commun., 1998, 1187–1188 RSC.
- K. Saha, S. S. Agasti, C. Kim, X. Li and V. M. Rotello, Chem. Rev., 2012, 112, 2739–2779 CrossRef CAS PubMed.
- G. V. P. Kumar, S. Shruthi, B. Vibha, B. A. A. Reddy, T. K. Kundu and C. Narayana, J. Phys. Chem. C, 2007, 111, 4388–4392 CAS.
- K. Faulds, R. P. Barbagallo, J. T. Keer, W. E. Smith and D. Graham, Analyst, 2004, 129, 567–568 RSC.
- D. Craig, J. Simpson, K. Faulds and D. Graham, Chem. Commun., 2013, 49, 30–32 RSC.
- E. Smith and G. Dent, Modern Raman Spectroscopy: A Practical Approach, John Wiley & Sons Ltd., Chichester, West Sussex, 1st edn, 2005 Search PubMed.
- K. Faulds, F. McKenzie, W. E. Smith and D. Graham, Angew. Chem., 2007, 119, 1861–1863 CrossRef.
- R. A. Halvorson and P. J. Vikesland, Environ. Sci. Technol., 2010, 44, 7749–7755 CrossRef CAS PubMed.
- K. L. Rule and P. J. Vikesland, Environ. Sci. Technol., 2009, 43, 1147–1152 CrossRef CAS PubMed.
- C. L. Schofield, R. A. Field and D. A. Russell, Anal. Chem., 2007, 79, 1356–1361 CrossRef CAS PubMed.
- N. C. Reichardt, M. Martin-Lomas and S. Penadés, Chem. Soc. Rev., 2013, 42, 4358–4376 RSC.
- B. K. Gorityala, Z. Lu, M. L. Leow, J. Ma and X.-W. Liu, J. Am. Chem. Soc., 2012, 134, 15229–15232 CrossRef CAS PubMed.
- D. Craig, S. McAughtrie, J. Simpson, C. McCraw, K. Faulds and D. Graham, Anal. Chem., 2014, 86, 4775–4782 CrossRef CAS PubMed.
- T. K. Lindhorst, Essentials of Carbohydrate Chemistry and Biochemistry, Wiley-VCH Verlag GmbH & Co. KGaA, Weinheim, 3rd edn, 2007 Search PubMed.
- L. Chen and F. Li, J. Virol., 2013, 87, 4118–4120 CrossRef CAS PubMed.
- E. A. Merritt, S. Sarfaty, F. V. D. Akker, C. L'Hoir, J. A. Martial and W. G. J. Hol, Protein Sci., 1994, 3, 166–175 CrossRef CAS PubMed.
- R. J. Pieters, Org. Biomol. Chem., 2009, 7, 2013–2025 CAS.
- M. Bergstrom, S. Liu, K. L. Kiick and S. Ohlson, Chem. Biol. Drug Des., 2009, 73, 132–141 Search PubMed.
- J. Sanchez and J. Holmgren, Indian J. Med. Res., 2011, 133, 153–163 CAS.
- G. McAnally, C. McLaughlin, R. Brown, D. C. Robson, K. Faulds, D. R. Tackley, W. E. Smith and D. Graham, Analyst, 2002, 127, 838–841 RSC.
- C. Sonnichsen, B. M. Reinhard, J. Liphardt and A. P. Alivisatos, Nat. Biotechnol., 2005, 23, 741–745 CrossRef PubMed.
- L. Guerrini, F. McKenzie, A. W. Wark, K. Faulds and D. Graham, Chem. Sci., 2012, 3, 2262–2269 RSC.
- CDC, Centre for Disease Control and Prevention – Laboratory Testing for Cholera, 1993, pp. 62–88.
- D. L. Evers, J. He, J. T. Mason and T. J. O'Leary, J. Clin. Microbiol., 2010, 48, 4620–4622 CrossRef PubMed.
- U. Tuteja, S. Kumar, J. Shukla, J. Kingston and H. V. Batra, J. Med. Microbiol., 2007, 56, 1340–1345 CrossRef CAS PubMed.
- V. L. Schmit, R. Martoglio and K. T. Carron, Anal. Chem., 2012, 84, 4233–4236 CrossRef CAS PubMed.
Footnote |
| † Electronic supplementary information (ESI) available. See DOI: 10.1039/c5nh00036j |
| This journal is © The Royal Society of Chemistry 2016 |