Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Evaluation of gas chromatography mass spectrometry and pattern recognition for the identification of bladder cancer from urine headspace
M.
Cauchi
*a,
C. M.
Weber
a,
B. J.
Bolt
a,
P. B.
Spratt
a,
C.
Bessant
b,
D. C.
Turner
c,
C. M.
Willis
d,
L. E.
Britton
d,
C.
Turner
c and
G.
Morgan
c
aCranfield Biotechnology Centre, Cranfield University, Bedfordshire, MK43 0AL, UK. E-mail: m.cauchi@cranfield.ac.uk
bSchool of Biological and Chemical Sciences, Queen Mary University of London, Mile End Road, London E1 4NS, UK
cDepartment of Life, Health and Chemical Sciences, Open University, Milton Keynes, MK7 6AA, UK
dDepartment of Dermatology, Amersham Hospital, Amersham, Buckinghamshire HP7 0JD, UK
First published on 3rd May 2016
Abstract
Previous studies have indicated that volatile organic compounds specific to bladder cancer may exist in urine headspace, raising the possibility that they may be of diagnostic value for this particular cancer. To further examine this hypothesis, urine samples were collected from patients diagnosed with either bladder cancer or a non-cancerous urological disease/infection, and from healthy volunteers, from which the volatile metabolomes were analysed using gas chromatography mass spectrometry. The acquired data were subjected to a specifically designed pattern recognition algorithm, involving cross-model validation. The best diagnostic performance, achieved with independent test data provided by healthy volunteers and bladder cancer patients, was 89% overall accuracy (90% sensitivity and 88% specificity). Permutation tests showed that these were statistically significant, providing further evidence of the potential for volatile biomarkers to form the basis of a non-invasive diagnostic technique.
Introduction
Bladder cancer is the seventh most common cancer in the UK, with over 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 700 new cases diagnosed in 2012.1 As with most cancers, early diagnosis greatly increases the chances of survival; individuals presenting with stage I tumours having a one year relative survival rate of around 97%, compared to 26% for those with stage IV disease.2 For people exhibiting symptoms or requiring surveillance, cystoscopy with biopsy remains the “gold standard” investigative technique for bladder cancer detection, but is invasive, expensive and time-consuming. Urine cytology can be a useful non-invasive adjunct to diagnosis, since it has a high specificity for bladder cancer (96–98%), but its sensitivity is low (22–52%), especially for low-grade tumours which shed proportionally fewer cells into the urine. Furthermore, an experienced cytologist or pathologist is needed to perform the cytological evaluation, making the test relatively expensive and slow.3
700 new cases diagnosed in 2012.1 As with most cancers, early diagnosis greatly increases the chances of survival; individuals presenting with stage I tumours having a one year relative survival rate of around 97%, compared to 26% for those with stage IV disease.2 For people exhibiting symptoms or requiring surveillance, cystoscopy with biopsy remains the “gold standard” investigative technique for bladder cancer detection, but is invasive, expensive and time-consuming. Urine cytology can be a useful non-invasive adjunct to diagnosis, since it has a high specificity for bladder cancer (96–98%), but its sensitivity is low (22–52%), especially for low-grade tumours which shed proportionally fewer cells into the urine. Furthermore, an experienced cytologist or pathologist is needed to perform the cytological evaluation, making the test relatively expensive and slow.3
Utilisation of molecular biomarkers present in urine offers a promising alternative non-invasive approach to diagnosis, which if sufficiently accurate, rapid and cheap has the potential to be used for mass screening of the population. Of the protein markers which have so far been investigated in depth, three have achieved FDA approval as assays for diagnosis and/or follow-up – nuclear mitotic apparatus protein (NMP22), complement factor H-related protein and complement factor H (BTA stat® and BTA TRAK®), and carcinoembryonic antigen combined with two bladder tumour cell-associated mucins (ImmunoCyt™/uCyt+™).4,5 Whilst these are more sensitive than urine cytology, having reported sensitivities of 47–100%, 53–83% and 50–100%, respectively, specificities are significantly lower at 60–90%, 51–75% and 69–79%, respectively.
Recently, it has been suggested that volatile organic compounds (VOCs) present in the headspace of urine from bladder cancer sufferers may be used as diagnostic biomarkers. This concept was initially demonstrated in a canine olfactory proof-of-principle study by Willis et al.6 and subsequently supported by findings using a metal oxide semiconductor (MOS) and field effect transistor (MOSFET) gas sensor array,7 where sensitivity and specificity rates of up to 70% were achieved. A more recent pilot study by Khalid et al.8 involving 24 bladder cancer patients and 74 control patients with non-malignant urological disease, utilised an in-house fabricated combined gas chromatography (GC) MOS-sensor device with pattern recognition, reporting accuracies of between 93% and 100% for the correct assignment of urine samples. Although very promising, the authors acknowledge that larger sample sizes are needed to confirm the results.
Gas sensor arrays undoubtedly offer practical advantages over trained dogs for the detection of the urinary VOCs associated with bladder cancer. However, they currently exhibit performance limitations, including sensor drift and a lack of inter-device reproducibility, and, furthermore, cannot be used to identify the chemical nature of individual volatile biomarkers. In the present study, we apply a more revealing analytical technique; that of gas chromatography mass spectrometry (GC-MS), and further demonstrates the potential for VOCs as a diagnostic approach to bladder cancer. GC-MS has already shown promise in the early diagnosis of lung cancer based on the analysis of VOCs contained in breath samples.9 It is now an important analytical technique in the field of metabolomics due to its high sensitivity, reproducibility and peak resolution.10 As early as 1980, methods had been established that could identify up to 155 metabolites in samples originating from urine.11,12
A number of different mass spectrometry systems are available for such analysis, including time-of-flight (ToF) and quadrupoles coupled with a database containing a library of spectral data for the identification of compounds.13 Recent advances have been seen in the separation of compounds with the advent of GCx-GC coupled with ToF-MS.14 In this regard, copious amounts of data are generated which require a robust statistical analytical approach, such as chemometrics,15 and, in particular, multivariate data analysis. This can sometimes involve an exploratory approach typically using principal components analysis (PCA) to identify possible trends and outlying samples16 which is followed by pattern recognition.17 The latter, in the form of multivariate classification with partial least squares discriminant analysis (PLS-DA), can deduce which type of class a particular sample belongs to, for example, healthy or diseased.18,19 Although there are other machine learning algorithms available, e.g. artificial neural networks (ANNs),20 random forests21 and support vector machines (SVMs),22,23 PLS-DA permits visualisation of the most significant features in a given chromatogram via the PLS loadings.19,24
This paper presents the identification and classification of bladder cancer via the multivariate statistical technique of partial least squares discriminant analysis (PLS-DA) and the machine learning approaches of support vector machines and random forests, on GC-MS data acquired from urine samples.
Experimental
Reagents
Analytical grade reagents and solvents were employed, unless otherwise stated.Participant selection
A total of 72 patients (Table 1) presenting at Buckinghamshire Healthcare NHS Trust with new or recurrent transitional cell carcinoma (TCC) of the bladder donated urine prior to surgical intervention. Grade and stage of the tumour were recorded, and three groups drawn up based on grade: TCC1 – low grade or well differentiated; TCC2 – moderately differentiated; TCC3 – high grade or poorly differentiated. An additional 205 control subjects, categorised into one of three groups (controls 1, 2 and 3, depending upon age and disease status), also provided urine samples. The control groups were split as follows: control group 1 (C1) – no urine abnormality on dipstick analysis; Control group 2 (C2) – any non-urological non-cancerous condition or disease, and/or one or more positive dipstick findings of a minor nature. Menstruating women with blood in their urine were included in this group, for example, as were individuals with suspected urinary tract infection, positive for leucocytes, blood and/or protein.| Group | No. of subjects | No. of males | No. of females | Age range (y) | Median age (y) |
|---|---|---|---|---|---|
| TCC1 | 17 | 12 | 5 | 59–82 | 74.0 |
| TCC2 | 28 | 19 | 9 | 50–86 | 66.5 |
| TCC3 | 27 | 15 | 12 | 56–88 | 75.5 |
| C1 | 70 | 29 | 41 | 18–31 | 26.0 |
| C2 | 71 | 35 | 36 | 18–32 | 25.0 |
| C3 | 46 | 8 | 38 | 50–89 | 66.0 |
Control group 3 (C3) – confirmed non-cancerous urological disease, with or without urine dipstick abnormalities. Urological conditions included renal and ureteric stones, renal cysts and polypoid cystitis.
As criteria for inclusion/exclusion, controls over 32 years of age were required to have had recent cystoscopy to exclude visible bladder malignancy. For both controls and the cancer positive group (TCC), men over 50 years were only included if recent cancer-negative prostate histology had been demonstrated. Individuals with pre-malignant urological disease or a history of urological carcinoma other than TCC were excluded. A history of malignancy in other organ systems (>5 years previously) was acceptable, providing the individual was now considered disease-free. All other past and/or present medical conditions were permissible. There were no exclusions on the basis of medication, menstrual cycle, diet, alcohol consumption, or chemical exposure. However, details of all of these factors were recorded for each participant, should their influence on the composition and odour of the urine need to be considered at any stage. Special attention was paid to smoking habits, with 28% of those with bladder cancer being current cigarette smokers, as compared to 31% control subjects. Finally, in order to ensure that age would not be a main contributory factor when comparing the C3 group against the TCC groups, 18 subjects under the age of 50 were omitted from the C3 group.
The study was given favourable ethical opinion by the Mid and South Buckinghamshire Local Research Ethics Committee (04/Q1607/65), and all participants gave written informed consent; after samples were taken, they and all subsequent data were anonymised.
Analysis and processing of urine samples
Following urinalysis (Multistix 10 SG, Bayer Corporation, NY, USA), fresh urine specimens were refrigerated immediately, and frozen as soon as possible as 0.5 ml aliquots in glass vials. The median time interval between refrigeration and freezing was 3 hours (range 1–24 hours). Samples were then stored at −80 °C until required. It was found in a recent study that the effect of freezing samples had no noticeable effect on the volatile composition of the samples.25 The use of glass vials has recently become of concern due to it being able to absorb volatiles.26 However the absorption of analytes onto the glass is dependent on a very large range of factors including concentration, functional groups, etc. Generally, freezing reduces the likelihood of interaction with the glass vials. Though reduced surface activity (RSA) vials are readily available which significantly reduces silanols and surface ions on the glass surface,27 they were not available during the initial stages of the work and thus glass vials were employed. However, it is stressed that the smallest glass vials were utilised to minimise the headspace and the surface area therefore resulting in minimal losses. Incidentally, plastic vials would not be suitable for GC analyses.Headspace analysis
Gas chromatography mass spectrometry was used to characterise the VOC (volatile organic compound) content of urine. Measurements were performed using the following instrumentation:• CTC CombiPal Autosampler (CTC Analytics, Switzerland): to automatically introduce the sample into the inlet.
• Agilent 6890 GC with S/SL inlet (Agilent Technologies, CA, USA): a gas chromatograph with an injector to introduce the vaporised sample onto the column.
• Leco Pegasus 4D ToFMS (Leco Corp., MI, USA): a time of flight mass spectrometer.
A total of 832 urine (C1, C2, C3, TCC1, TCC2 and TCC3) samples were randomly analysed over 9 batches and interspersed with either a fibre blank (no sample) or sample blank (urine replaced with 0.5 ml deionised water) after every 5 injections. All samples were prepared by placing a 0.5 ml sample in a pre-conditioned 10 ml headspace vial containing 1 g anhydrous sodium sulphate (Fisher Scientific UK Ltd., Loughborough, UK) conditioned overnight at 100 °C and 1.5 ml of 0.1 M hydrochloric acid (Fisher Scientific UK Ltd., Loughborough, UK). An internal standard in the form of deuterated (d6-) phenol (ISOTEC, Miamisburg, Ohio, USA) at a concentration of 100 mg ml−1 was spiked (10 ml) into the vial which was immediately capped. This mixture was pre-equilibrated for 10 minutes at 60 °C. A pre-conditioned 75 μm carboxen/PDMS fiber (Sigma-Aldrich, Dorset, UK) was inserted for 5 minutes to extract the volatile organic compounds and then the fiber was exposed in the GC inlet at 280 °C for 2 minutes under splitless conditions to desorb the analytes onto the column. In this work, only one column was employed in the GC-ToF-MS instrument. The analytes were thus separated on a BP624 30 m × 0.25 mm internal diameter with a 1.4 μm film thickness column (SGE Analytical Science, Victoria, Australia) with the oven programmed from 30 °C (2 minute hold) to 240 °C at 20 °C min−1 (hold 1.5 min). The data were collected at 10 spectra per second across the mass range 33–350 m/z. The mass range started at m/z 33 so as to avoid background interferences and higher baselines from the oxygen (m/z 32) and nitrogen (m/z 28) and using this headspace technique in order that analytes with a molecular weight greater than 350 amu would not be introduced into the GC. The reproducibility of the method was checked before measurements of the samples were made in triplicate.
Finally, the data were stored in NetCDF format (Network Common Data Form). These are binary files (i.e. cannot be opened in a standard text editor, such as NotePad) in which specific information is stored and all zero values are removed in order to minimise the storage space used on a hard drive. All information is stored as row vectors. Information includes some of the following:
• Total_intensity: the sum of the abundances across all of the retention times. The length of the vector is the number of retention time scans.
• Scan_acquisition_time: the vector of retention time values containing the time values in minutes.
• Scan_index: the index values indicating the starting positions of each retention time scan in the mass_values and intensity_values vectors (see below). The length of the vector is the number of retention time scans.
• Point_index: this gives the number of non-zero data points for each retention time value. The length of the vector is the number of retention time scans.
• Mass_values: the actual mass-to-charge (m/z) values corresponding to the non-zero values. The length of the vector is the sum of all the numerical values in the point_index vector.
• Intensity_values: the corresponding intensity values for each of the respective mass values. The length of the vector is the sum of all the numerical values in the point_index vector.
Data analysis
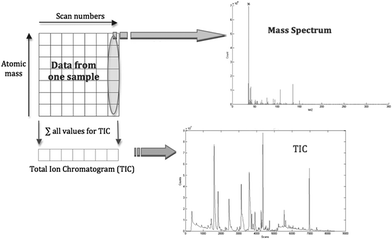
The provided NetCDF data files were processed and analysed using MATLAB (R2011a, MathWorks Inc, USA). Each file contained the information of the full spectral information of one sample, a chromatogram, which was stored in a data matrix of size m/z_values × scans. From a data storage point of view, all samples build a cube – one chromatogram arranged behind the other. Every single entry of the data matrix of one sample represents the abundance of a specific ion at a certain point of time. Each column in the matrix can be interpreted as a mass spectrum. A typical mass spectrum is usually represented as a “stick diagram”, displaying the relative current induced by ions of alternating mass-to-charge ratio. But when it comes to the storage of the data and the computational data processing point of view, each mass spectrum is represented as an array of numbers. The rows of a GC-MS chromatogram represent single ion count (SIC) chromatograms. This fact allows inferring the total ion count (TIC) chromatogram by summing up the columns. This data reduction was necessary, as the majority of multivariate data analysis techniques require two-dimensional data.For each NetCDF data file that was imported into the MATLAB environment, and based on the knowledge of the contents of the NetCDF file given previously, the GC-MS data matrix was reconstructed to the order of m/z_values × scans re-inserting zero values where appropriate into the single ion count (SIC) chromatograms. All of the abundance values were normalised against the abundance values of the deuterated (d6-) phenol internal standard (at m/z 99). The m/z values are summed so that a row vector is generated whose length is the number of scans (i.e. the retention time values). The same process is repeated with the remaining NetCDF files. Finally, all row vectors are combined into a data matrix of the order samples × scans. Fig. 1 illustrates the relationship among the elements within a single data matrix and demonstrates the formation of the dataset containing the TIC of each sample.
As the process required chromatograms to be warped in time to align corresponding peaks, correlation optimised warping (COW) was applied28,29 on these data prior to further data analysis. The “retention time shifts” can be caused by physical changes in the column, mobile phase composition, instrumental drift and interaction between analytes, and these must be corrected.30 Although other warping methods exist,31–34 COW was employed due to the ability to preserve peak shape and area, in addition to the ability to deduce the optimal parameters required for alignment of the retention time peaks.29 The deduced optimal parameters are the segment (the number of data points per interval) and the slack (the extent of warping/shifting of the peaks in any direction). The segment and slack values attained for C1 v TCC, C2 v TCC, C3 v TCC, C3 v TCC1, C3 v TCC2, and C3 v TCC3 were {31, 1}, {23, 1}, {6, 1}, {23, 1}, {19, 1} and {6, 1} respectively.
Exploratory data analysis was accomplished via principal components analysis (PCA) and hierarchical cluster analysis (HCA), which are the most widely used multivariate statistical techniques.15,35 This was performed to reveal natural groupings based on the chromatograms of the GC-MS via the characteristics that cause the greatest variance in the dataset.
Next, three pattern recognition tools were employed via custom-written scripts to build classification models using the cancer status of the samples: partial least squares discriminant analysis (PLS-DA), random forests (RFs) and support vector machines (SVMs). For PLS-DA, the PLS Toolbox 3.5 (Eigenvector Research Inc., USA) was employed in MATLAB R2011a (MathWorks Inc., Nattick, USA); for SVMs the libsvm3.20 toolbox was employed; for RFs, MATLAB was made to call the RandomForest package in R (3.0.2). All three techniques call for information about the parameter of interest (the cancer status) to be known in order to train the algorithm to identify those molecules that differentiate between the classes.
PLS-DA is considered to be a dimensionality reduction method and can be seen as the regression extension of principal components analysis.36 Unlike PCA, which attempts to describe the maximum variation in the measured data, PLS-DA tends to maximise the covariance between the input data and the output class. The information returned by PCA is that which was caused by the attribute with the biggest variance. In contrast, PLS-DA returns only data that were caused by the property under investigation.
It is known that PLS-DA is prone to overestimate the accuracy of classification if it is not accurately validated.37 For this reason the number of latent variables (LVs) was varied from 1 to 20 in each test run. Furthermore a very thorough evaluation process – bootstrapping with optimisation by leave-one-out cross-validation (LOOCV)38,39 – was implemented to assess the performance of the PLS-DA classifier. In each bootstrap evaluation, the dataset was randomly split into two subsets: the first subset was the bootstrap training set which would be used to determine the optimum model parameters via LOO-CV and was made up of 70% of the original dataset; the remaining 30% formed the bootstrap testing set which would be used to evaluate the model at the determined optimum LV. This whole process was repeated for the next bootstrap evaluation until all 150 evaluations had taken place. A set of statistical parameters are then calculated such as the overall accuracy, specificity, sensitivity and the area under the receiver operating characteristic (AUROC) curve which uses the trapezoid rule.40 This method ensures that validation is sequentially performed on each sample using a model that excludes the data from that sample.
Two machine learning algorithms were also employed: random forests21 and support vector machines.23 In order to ensure the optimum number of trees was employed for random forests, they were varied from 50 up to 450 in steps of 100. The linear kernel was employed for SVM. During the optimisation process of the linear kernel the cost values applied were 0.5, 1.0, 2.0, 4.0 and 8.0. These two machine learning approaches were integrated into the bootstrapping procedures described in the previous paragraph.
As final validation of the results, and to attain an indication of the statistical significance of the results, permutation testing involving a Monte Carlo simulation was used to evaluate the obtained results.38 This involved repeated random sampling. In this context a null model was generated from a set of data that was statistically similar to the data under study, but for which it was not expected to be able to build a meaningful classification model. For each of the 6 datasets (C1 v TCC, C2 v TCC, C3 v TCC, C3 v TCC1, C3 v TCC2, and C3 v TCC3), random class assignations were made to the samples in the datasets 300 times. Within each random assignation, the datasets were subjected to the bootstrap procedure described previously. For a disease discriminating model trained on the real sample classes to be considered statistically significant it needs to achieve a classification accuracy towards the extremities of those produced by the null models.
Results and discussion
Exploratory analysis via PCA and HCA
The visual outputs of the two independent exploratory techniques of principal components analysis (PCA) and hierarchical cluster analysis (HCA) did not disclose any separation by cancer status of the samples, in any of the experiments. Other influences such as age, diet or gender may be responsible for the groupings obtained. However, this does not mean that the data do not contain any information concerning bladder cancer. The PCA was able to demonstrate that the cancer status was not responsible for the bigger part of the variance, captured by the first two or three principal components (PCs). Nevertheless, investigating principal components of lower variance did not lead to an explicitly disease-related differentiation, either.Pattern recognition via PLS-DA, SVMs and RFs
Table 2 compares the results attained via the machine learning algorithms of support vector machines (SVMs) and random forests (RFs) along with the multivariate statistical technique of partial least squares discriminant analysis (PLS-DA). Each chromatogram contained approximately 8400 data points, i.e. all of the features. This enables multivariate methods such as PLS-DA to be able to detect “hidden features” that are crucial for the model to distinguish between cancer and control samples, which univariate methods are not able to identify properly.| Dataset | Model comparison | % overall | % spec | % sens | LV or tree | AUROC |
|---|---|---|---|---|---|---|
| C1 v TCC | PLS-DA | 87.53 | 87.23 | 87.82 | 16 | 0.906 |
| SVM Lin | 88.99 | 88.84 | 89.13 | — | 0.935 | |
| RF | 80.91 | 80.28 | 81.75 | 450 | 0.892 | |
| C2 v TCC | PLS-DA | 88.35 | 88.21 | 88.48 | 12 | 0.928 |
| SVM Lin | 89.18 | 88.00 | 90.33 | — | 0.922 | |
| RF | 82.70 | 82.93 | 82.72 | 450 | 0.865 | |
| C3 v TCC | PLS-DA | 83.01 | 66.06 | 88.66 | 8 | 0.8680 |
| SVM Lin | 83.48 | 44.36 | 96.52 | — | 0.9023 | |
| RF | 83.57 | 42.90 | 86.99 | 150 | 0.8427 | |
| C3 v TCC1 | PLS-DA | 69.18 | 66.18 | 73.29 | 13 | 0.7424 |
| SVM Lin | 67.30 | 86.15 | 41.38 | — | 0.6363 | |
| RF | 67.33 | 77.63 | 54.03 | 450 | 0.7102 | |
| C3 v TCC2 | PLS-DA | 80.51 | 71.39 | 88.23 | 7 | 0.8985 |
| SVM Lin | 81.44 | 72.15 | 89.31 | — | 0.9040 | |
| RF | 75.87 | 64.31 | 86.66 | 350 | 0.8642 | |
| C3 v TCC3 | PLS-DA | 79.70 | 73.48 | 85.17 | 20 | 0.8580 |
| SVM Lin | 81.46 | 73.91 | 88.11 | — | 0.9283 | |
| RF | 74.44 | 66.76 | 81.64 | 350 | 0.8098 |
It is clear to see that the C3 v TCC1 dataset has been the most difficult to classify due to the nature of the datasets: TCC1 being the low grade and C3 other urological diseases. The random forests and support vector machines algorithms have not performed as well as the partial least squares discriminant analysis algorithm in this instance. As far as the classification models are concerned, the classifiers were trained with the two most disparate groups: control 1 (C1), representing healthy males or females, and the TCC groups incorporating people suffering from bladder cancer. Since group C1 possesses the most differences compared to the cancer group, the classification outcome of this sample set was expected to be the best. However this was surprisingly not the case. A mean total accuracy of 87.5%, 89.0% and 80.9% were attained for PLS-DA, SVM and RF respectively.
Next, the classifier with control 2 (C2) and the cancer group (TCC) data was trained. Urine samples within this control subgroup showed similar abnormalities on dipstick analysis to some cancer samples, such as blood, for example, and were therefore more difficult to distinguish from cancerous samples than control 1 (C1) samples. However, the achieved specificity contradicts this (for example, PLS-DA at 88.2% compared with 87.2% for C1). The overall classified accuracies attained were greater for each classifier than C1.
In the third experiment, the classifier had to distinguish between samples with confirmed non-cancerous urological diseases (control 3) and cancerous samples (TCC). This was expected to be the most difficult combination, as disease markers not specific to bladder cancer are likely to be present. The achieved total accuracies appeared to perform better than expected as they attained values of 83.0%, 83.5% and 83.6% for PLS-DA, SVM and RF respectively. However it is noted that the specificities attained were especially poor for SVM Lin and RF (<50%) yet PLS-DA was at 66.1% suggesting that PLS-DA is the better algorithm. The specificity values attained can be attributed to the unbalanced nature of the data since the TCC subgroup is far greater (combining TCC1, TCC2 and TCC3) than the C3 subgroup (Table 1) suggesting that the models learn better the patterns attributed to the TCC group more so than the C3 group.
The remaining experiments focusing on C3 versus the TCC cancer grades (TCC1, TCC2 and TCC3) show that SVM-Lin was better than PLS-DA and RF at discriminating the control (C3) from the TCC grades due to the overall and sensitivities attained for C3 v TCC2 and C3 v TCC3 (SVM > PLS-DA > RF). However, for C3 v TCC1, PLS-DA was shown to be better than SVM and RF, especially as the latter two only achieved sensitivities of 41.4% and 54.0% respectively. This suggests that the PLS-DA classifier was able to distinguish to a certain extent the C3 control from the low grade TCC (TCC1) whilst SVM and RF could not. From a clinical perspective, the ability to distinguish between the C3 control and TCC1 is of paramount importance.
To assess the significance of the presented results, permutation testing via a Monte Carlo simulation was carried out. Fig. 2 shows the results attained for each of the six experiments each with 300 random runs (dark grey vertical bars) for the PLS-DA classifier. It also shows the respective distributions of the observed analytical accuracies attained via the 150 classification models generated (light grey vertical bars) during the analysis.
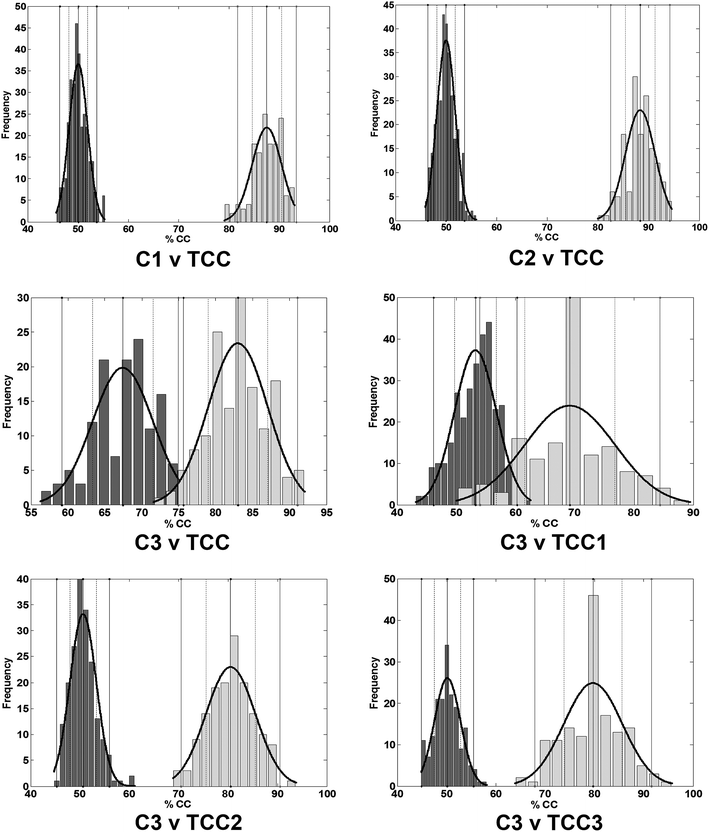 | ||
| Fig. 2 Distribution of the overall percentage classified after randomised assignation of classes to the samples (dark grey vertical bars) corresponding to each of the six experiments via the PLS-DA classifier. Number of runs: 300. The light grey vertical bars denote distribution of the observed accuracies attained via the classification models (150 runs). It can be seen that the respective means of the accuracy attained (the maxima of the rightmost distribution curve) is beyond two standard deviations of the respective permutation means (the maxima of the leftmost distribution curve) indicating that statistically significant results had been achieved at the 95% confidence level. This is further corroborated in Table 3. Confidence intervals (CI) for evaluations: (C1 v TCC: mean: 87.5%; CI (95%): 82–94%); (C2 v TCC: mean: 88.4%; CI (95%): 82–95%); (C3 v TCC: mean: 83.0%; CI (95%): 75–91%); (C3 v TCC1: mean: 69.2%; CI (95%): 54–84%); (C3 v TCC2: mean: 80.5%; CI (95%): 70–90%); (C3 v TCC3: mean: 79.7%; CI (95%): 68–91%). | ||
Although overlap had been observed in the distributions for C3 v TCC and C3 v TCC1 (Fig. 2), the Z-test41 was carried out to test for significance between the means of the two distributions. As Table 3 shows, all calculated probability (p) values were lower than the critical value (α = 0.05) indicating that the means of the two distributions are statistically significantly different. This implies that the controls can be distinguished from TCC as well as C3 against all of the TCC grades. Furthermore, the area under the receiver operating characteristic (AUROC) curve values calculated for each of the experiments (Table 2) give further support to the findings with values ranging from 0.93 for C2 v TCC to 0.74 for C3 v TCC1 for the PLS-DA classifier.
| Case | Overall accuracy (%) | Z value (Zcrit = 1.96) | p-Value (α = 0.05) | Significant difference |
|---|---|---|---|---|
| C1 v TCC | 87.53 | 143.61 | <0.0001 | Yes |
| C2 v TCC | 88.35 | 147.54 | <0.0001 | Yes |
| C3 v TCC | 83.01 | 32.02 | <0.0001 | Yes |
| C3 v TCC1 | 69.18 | 24.42 | <0.0001 | Yes |
| C3 v TCC2 | 80.51 | 66.07 | <0.0001 | Yes |
| C3 v TCC3 | 79.70 | 56.70 | <0.0001 | Yes |
Diagnostic potential
By combining gas chromatography mass spectrometry with pattern recognition techniques, progress towards a new instrumental method of bladder cancer detection based on volatile biomarkers has been made. The obtained results confirm that there is a clear relationship between the acquired GC-MS data and the cancer status of the respective samples. This relationship shows promise as the basis of a non-invasive diagnostic technique. As many as 88.5% of cancer patients and 88.2% of non-cancerous subjects were correctly classified when the classifier was trained with a combination of TCC positive urine samples and samples from healthy control groups containing patients diagnosed with some form of non-cancerous disease such as urinary tract infections (C2).Samples from group C2 showed abnormalities such as blood, for example haematuria – blood in the urine – is the most common symptom of bladder cancer. Samples containing traces of blood therefore represent a challenge for the distinction between control samples and bladder cancer samples. However, the major contributor to this classification outcome was control group 3. All subjects within this subgroup had confirmed non-cancerous urological disease, the pathological effects of which are likely to be similar to the secondary effects of bladder cancer. Within both these groups, varying amounts of metabolic products associated with inflammation, infection and/or necrosis will almost certainly be present. Because of this, control 3 samples form the most important control subset and contain the most relevant information. Training the classifier with this kind of data is therefore fundamental in order to be able to subtract general disease compounds present in the urine from those specific for bladder cancer. Accurate diagnosis of the control subjects is, of course, paramount to this process, since the inclusion of false negative individuals would lead to incorrect classification rules.
Interestingly, within the TCC sample group, the majority of those incorrectly classified as negative were from patients with more advanced tumours. In these cases, it is possible that metabolic products generated secondarily to the tumour may overwhelm or mask the volatile cancer biomarkers within the urine, giving rise to a urine headspace more closely resembling that of control 3 samples. Canine olfactory studies support this hypothesis; high grade TCCs with a significant level of invasion are missed more frequently by trained dogs than low-grade superficial tumours.42
Fig. 2 also showed the increase in complexity of the control samples (C1 to C3) as reflected in the poorer performing models with overall classifications of ∼80%, ∼80% and ∼73% for C1, C2 and C3 respectively. In most cases, the best performing models were shown to achieve an overall classification of ∼95% for both C1 and C2, and 92% for C3. More so, Fig. 2 clearly illustrates the difficulty in distinguishing the C3 control group from the TCC1 cancer group via PLS-DA. This was also observed via support vector machines (SVMs) and random forests (RFs) suggesting that a more rigorous modelling algorithm/machine learning technique is warranted in conjunction with data pre-processing and pre-treatment methods.
Visualisation of the PLS-DA loadings revealed a number of possible metabolites/compounds which could be potential biomarkers for the determination of TCC. These are summarised in Table 4. As is often the case with complex samples analysed by GC-MS, the identity of some compounds determined through using NIST (National Institute of Standards and Technology) and MassBank (http://www.massbank.jp) is less certain due to incomplete separation and similar library spectra for different (but related) compounds. However, based on the most likely compound identification, the list does not seem to concur with the list of biomarkers suggested by Pasikanti et al.43 Yet some of the suggested compounds in Table 4 have been identified as being significant in colo-rectal cancer, i.e. 2-pentanone, hexanal and 2,3-butanedione44 (suggested in Table 4 to decrease from C3 to TCC); 3-hydroxyanthranilic acid has been found in bladder cancer45 (suggested in Table 4 to increase from C3 to TCC). In addition, 4-heptanone (suggested in Table 4 to decrease from C3 to TCC) was reported to be a marker for bladder cancer when human urine was analysed via headspace GC-MS.46 Other chemicals have been reported in the medical literature, but not as cancer markers. For example, piperitone has been reported to inhibit the cervical cancer cell-line growths,47 benzoic acid (suggested in Table 4 to increase from C3 to TCC) reduces bladder cancer when as a functional group within the retinoid-related molecule AGN193198,48 and butyrophenone (suggested in Table 4 to increase from C3 to TCC) is employed in the treatment of schizophrenia and other central nervous disorders49 though it is unclear if any patients were taking this medication.
| Compound | Database | Change |
|---|---|---|
| 2-Pentanone | NIST & MassBank | Decrease |
| 2,3-Butanedione | MassBank | Decrease |
| 4-Heptanone | MassBank | Decrease |
| Dimethyl disulphide | NIST | Decrease |
| Hexanal | NIST | Increase |
| Benzaldehyde | MassBank | Increase |
| Butyrophenone | MassBank | Increase |
| 3-Hydroxyanthranilic acid | MassBank | Increase |
| Benzoic acid | MassBank | Increase |
| trans-3-Hexanoic acid | MassBank | Increase |
| cis-3-Hexanoic acid | MassBank | Increase |
| 2-Butanone | NIST | Increase |
| 2-Propanol | NIST | Decrease |
| Acetic acid | NIST | Decrease |
| Piperitone | MassBank | Decrease |
| Thujone | MassBank | Decrease |
It should be noted that some biomarkers are almost ubiquitous biomarkers and can be seen as volatile compounds emanating from biological systems; examples include: dimethyl disulphide, 2-butanone, 2-propanol, acetic acid, etc. However, their relative concentrations may alter due to the presence of abnormal metabolism, and this may give information about changes occurring in that system. Though use of an internal standard had been employed (deuterated phenol), it may not have accounted for differing concentrations – where it had been observed during sample preparation that some urine samples were very watery whilst others more concentrated. However the same volume of urine was always taken therefore it is possible to make use of a naturally occurring internal standard such as creatinine. Furthermore, the concentration of acetic acid in the headspace may increase if the pH surrounding a tumour is lowered because it pushes the chemical equilibrium away from the acetate ion and to the acetic acid molecule which is much more volatile and hence detectable by this method. For this reason, it is quite reasonable that some “cancer biomarkers” are in fact compounds found under non-cancerous circumstances, but with varying relative concentrations; these can still form the basis for a diagnostic test.
Although Pasikanti and colleagues claim 100% sensitivity in identifying human bladder cancer,43 there is no specific mention of identifying transitional cell carcinoma (TCC) in conjunction with applying any retention time shift corrections. The authors have also not specified the clinical diagnoses of any of their controls (only that they had bladder cancer symptoms, but were cystoscopy negative), so the nature, severity or chronicity of their urological conditions are currently not known.
Though the article by Khalid et al.8 reported a success of 96% accuracy using two alternative statistical approaches, the first involving a simple linear discriminant analysis on 9 selected time points, and the second employing PLS-DA on all time points, both approaches only employed leave-one-out cross-validation. This has been shown to give overoptimistic results and it is thus recommended to employ a more thorough validation approach employing cross-model validation and permutation testing37 as has been employed in this work, and thus permitting greater confidence and reliability in the results presented. Finally, recent work has been reported in which nanoparticles are employed in conjunction with cystoscopy to improve the recognition of tumours, for example distinguishing flat lesions from non-malignant cells, yet though outcomes are positive, there is still an invasive element to the procedure.50
Finally, in a recent paper by Aggio et al., it was reported that a GC-sensor was able to distinguish in urine prostate cancer from controls, bladder cancer from controls, and bladder cancer from prostate cancer via an in-house data processing and analysis pipeline reporting very high (≫90%) accuracies, sensitivities and specificities.51 It was stated that “different VOCs are associated with the two urological disorders” however it must be suggested that it is very likely that there will also be the same VOCs present in both cancers. Both statements can be corroborated via the use of mass spectrometry in order to identify compounds, the potential of which have been demonstrated in this work, and are acknowledged by the authors for their future work.
Conclusions
PLS-DA-derived models gave a mean accuracy for patients presenting with other non-cancerous urological disease of 88.4%, with 88.5% sensitivity and 88.2% specificity for C2 versus TCC (TCC1, TCC2 and TCC3 combined). SVM-derived models had given a mean accuracy of 89.2%, with a sensitivity of 90.3% and specificity of 88.0%. Although the specificities achieved were marginally less than that of conventional urine cytology (typically >90% specificity), sensitivity was very close to typical range of 80–90% for high-grade tumours52 and thus better than the typical range of 20–50% for low-grade tumours,3 case in point, the sensitivity attained for C3 v TCC1 was 73.3% which is considerably better than the “gold-standard” of 20–50%. Of course, further improvement is still highly warranted.Acknowledgements
The authors would like to thank the staff of the Urology Department, Buckinghamshire Healthcare NHS Trust for their enthusiastic support.References
- CRUK, Bladder cancer incidence statistics, http://www.cancerresearchuk.org/health-professional/cancer-statistics/statistics-by-cancer-type/bladder-cancer/incidence#heading-Zero, accessed 13 January 2016, 2016.
- CRUK, Bladder cancer survival statistics, http://www.cancerresearchuk.org/health-professional/cancer-statistics/statistics-by-cancer-type/bladder-cancer/survival#heading-Three, accessed 13 January 2016, 2016.
- P. Bassi, V. De Marco, A. De Lisa, M. Mancini, F. Pinto, R. Bertoloni and F. Longo, Urol. Int., 2005, 75, 193–200 CrossRef PubMed.
- Z. L. Smith and T. J. Guzzo, F1000Prime Rep., 2013, 5, 21 Search PubMed.
- E. Xylinas, L. A. Kluth, M. Rieken, P. I. Karakiewicz, Y. Lotan and S. F. Shariat, Urol. Oncol.: Semin. Orig. Invest., 2013, 32, 222–229 CrossRef PubMed.
- C. M. Willis, S. M. Church, C. M. Guest, W. A. Cook, N. McCarthy, A. J. Bransbury, M. R. T. Church and J. C. T. Church, BMJ, 2004, 329, 712–716 CrossRef PubMed.
- C. M. Weber, M. Cauchi, M. Patel, C. Bessant, C. Turner, L. E. Britton and C. M. Willis, Analyst, 2011, 136, 359–364 RSC.
- T. Khalid, P. White, B. De Lacy Costello, R. Persad, R. Ewen, E. Johnson, C. S. Probert and N. Ratcliffe, PLoS One, 2013, 8, e69602 CAS.
- G. Song, T. Qin, H. Liu, G.-B. Xu, Y.-Y. Pan, F.-X. Xiong, K.-S. Gu, G.-P. Sun and Z.-D. Chen, Lung Cancer, 2009, 67, 227–231 CrossRef PubMed.
- K. K. Pasikanti, P. C. Ho and E. C. Y. Chan, J. Chromatogr. B: Anal. Technol. Biomed. Life Sci., 2008, 871, 202–211 CrossRef CAS PubMed.
- K. Tanaka, D. G. Hine, A. West-Dull and T. B. Lynn, Clin. Chem., 1980, 26, 1839–1846 CAS.
- K. Tanaka, A. West-Dull, D. G. Hine, T. B. Lynn and T. Lowe, Clin. Chem., 1980, 26, 1847–1853 CAS.
- E. J. Want, A. Nordstrom, H. Morita and G. Siuzdak, J. Proteome Res., 2007, 6, 459–468 CrossRef CAS PubMed.
- W. Welthagen, R. Shellie, J. Spranger, M. Ristow, R. Zimmermann and O. Fiehn, Metabolomics, 2005, 1, 65–73 CrossRef CAS.
- R. G. Brereton, Applied Chemometrics for Scientists, Wiley, Chichester, 2007 Search PubMed.
- S. Wold, K. Esbensen and P. Geladi, Chemom. Intell. Lab. Syst., 1987, 2, 37–52 CrossRef CAS.
- M. Otto, Chemometrics: Statistics and Computer Applications in Analytical Chemistry, Wiley-VCH, Weinheim, 2nd edn, 2007 Search PubMed.
- M. Barker and W. Rayens, J. Chemom., 2003, 17, 166–173 CrossRef CAS.
- S. Wiklund, E. Johansson, L. Sjostrom, E. J. Mellerowicz, U. Edlund, J. P. Shockcor, J. Gottfries, T. Moritz and J. Trygg, Anal. Chem., 2007, 80, 115–122 CrossRef PubMed.
- M. T. Hagan, H. B. Demuth and M. Beale, Neural Network Design, International Thompson Publishing, Boston, 1996 Search PubMed.
- L. Breiman, Mach. Learn., 2001, 45, 5–32 CrossRef.
- M. Sattlecker, C. Bessant, J. Smith and N. Stone, Analyst, 2010, 135, 895–901 RSC.
- V. Vapnik, The Nature of Statistical Learning Theory, Springer, New York, 1st edn, 1995 Search PubMed.
- J. Trygg, E. Holmes and T. r. Lundstedt, J. Proteome Res., 2007, 6, 469–479 CrossRef CAS PubMed.
- S. Smith, H. Burden, R. Persad, K. Whittington, B. d. L. Costello, N. M. Ratcliffe and C. S. Probert, J. Breath Res., 2008, 2, 037022 CrossRef CAS PubMed.
- MTC-USA, Compound-Dependent Vial Adsorption Studies - Comparison to Conventional glass, http://www.microsolvtech.com/PDF/No-277-Compound-Dependant-Adsorption-Studies-RSA-DH-C18-ANP.pdf, accessed 12 December 2015, 2015.
- MTC-USA, RSA – Reduced Surface Activity Glass, http://wwwmicrosolvtech.com/rsa_chart.asp, accessed 12 December 2015, 2015.
- T. Skov, F. van den Berg, G. Tomasi and R. Bro, J. Chemom., 2006, 20, 484–497 CrossRef CAS.
- G. Tomasi, F. van den Berg and C. Andersson, J. Chemom., 2004, 18, 231–241 CrossRef CAS.
- N.-P. V. Nielsen, J. M. Carstensen and J. r. Smedsgaard, J. Chromatogr. A, 1998, 805, 17–35 CrossRef CAS.
- N. Hoffmann and J. Stoye, Bioinformatics, 2009, 25, 2080–2081 CrossRef CAS PubMed.
- K. J. Johnson, B. W. Wright, K. H. Jarman and R. E. Synovec, J. Chromatogr. A, 2003, 996, 141–155 CrossRef CAS PubMed.
- A. Kassidas, J. F. MacGregor and P. A. Taylor, AIChE J., 1998, 44, 864–875 CrossRef CAS.
- B. Walczak and W. Wu, Chemom. Intell. Lab. Syst., 2005, 77, 173–180 CrossRef CAS.
- R. O. Duda, P. E. Hart and D. G. Stork, Pattern Classification, Wiley, Chichester, 2001 Search PubMed.
- K. Yuan, H. Kong, Y. Guan, J. Yang and G. Xu, J. Chromatogr. B: Anal. Technol. Biomed. Life Sci., 2007, 850, 236–240 CrossRef CAS PubMed.
- J. Westerhuis, H. Hoefsloot, S. Smit, D. Vis, A. Smilde, E. van Velzen, J. van Duijnhoven and F. van Dorsten, Metabolomics, 2008, 4, 81–89 CrossRef CAS.
- R. G. Brereton, Chemometrics for Pattern Recognition, Wiley-Blackwell, Chichester, 2009 Search PubMed.
- T. Hastie, R. Tibshirani and J. Friedman, The Elements of Statistical Learning: Data Mining, Inference and Prediction, Springer, Berlin, 2001 Search PubMed.
- Q. Rahman and G. Schmeisser, Numerische Mathematik, 1990, vol. 57, pp. 123–138 Search PubMed.
- M. J. Campbell and D. Machin, Medical Statistics: a Common Sense Approach, John Wiley & Sons Ltd, Chichester, UK, 3rd edn, 1999 Search PubMed.
- C. M. Willis, R. Harris, L. E. Britton, C. M. Guest and J. J. Wallace, Cancer Markers, 2010, 8, 145–153 CAS.
- K. K. Pasikanti, K. Esuvaranathan, P. C. Ho, R. Mahendran, R. Kamaraj, Q. H. Wu, E. Chiong and E. C. Y. Chan, J. Proteome Res., 2010, 9, 2988–2995 CrossRef CAS PubMed.
- R. Arasaradnam, P. M. J. McFarlane, C. Ryan-Fisher, E. Westenbrink, P. Hodges, M. G. Thomas, S. Chambers, N. O'Connell, C. Bailey, C. Harmston, C. U. Nwokolo, K. D. Bardhan and J. A. Covington, PLoS One, 2014, 9, e108750 Search PubMed.
- Y.-S. Tsai, Y.-C. Jou, Y.-P. Tsai, B.-D. Liu, H.-I. Lin, C.-L. Wei, S.-Y. Chen, H.-T. Tsai, C.-H. Ou, W.-H. Yang and Z.-S. Tzai, Urol. Sci., 2015, 26, S36–S49 CrossRef.
- F.-Y. Zhu, A.-N. Yu, Y.-H. Qiu, Y.-L. Sa and F. Wang, Chin. J. Anal. Chem., 2007, 35, 1132–1136 CAS.
- R. Ali, Z. Mirza, G. M. D. Ashraf, M. A. Kamal, S. A. Ansari, G. A. Damanhouri, A. M. Abuzenadah, A. G. Chaudhary and I. A. Sheikh, Anticancer Res., 2012, 32, 2999–3006 CAS.
- A. Reitmair, D.-L. Shurland, K.-Y. Tsang, R. Chandraratna and G. Brown, Int. J. Cancer, 2005, 115, 917–923 CrossRef CAS PubMed.
- R. Cacabelos, P. Cacabelos and G. Aliev, Open J. Psychiatr., 2013, 3, 46–139 CrossRef.
- B. Tomlinson, L. Tzu-yin, M. Dall'Era and C.-X. Pan, Nanomedicine, 2015, 10, 1189–1201 CrossRef CAS PubMed.
- R. B. M. Aggio, C. Ben de Lacy, P. White, T. Khalid, N. M. Ratcliffe, R. Persad and C. S. J. Probert, J. Breath Res., 2016, 10, 17106–17121 CrossRef PubMed.
- P. S. Sullivan, J. B. Chan, M. R. Levin and J. Rao, Am. J. Transl. Res., 2010, 2, 412–440 Search PubMed.
| This journal is © The Royal Society of Chemistry 2016 |