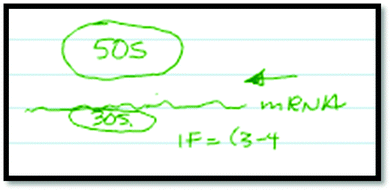What do biochemistry students pay attention to in external representations of protein translation? The case of the Shine–Dalgarno sequence
Thomas J.
Bussey
*a and
MaryKay
Orgill
b
aDepartment of Chemistry and Biochemistry, University of California, San Diego, USA. E-mail: tbussey@ucsd.edu
bDepartment of Chemistry and Biochemistry, University of Nevada, Las Vegas, USA
First published on 8th April 2015
Abstract
Biochemistry instructors often use external representations—ranging from static diagrams to dynamic animations and from simplistic, stylized illustrations to more complex, realistic presentations—to help their students visualize abstract cellular and molecular processes, mechanisms, and components. However, relatively little is known about how students use and interpret external representations in biochemistry courses. In the current study, variation theory was used to explore the potential for student learning about protein translation from a stylized, dynamic animation. The results of this study indicate that students learned from this animation, in that they noticed many critical features of the animation and integrated those features into their understandings of protein translation. However, many students also focused on a particular feature of the animation, the Shine–Dalgarno sequence, that their instructors did not feel was critical to promote an overall understanding of this metabolic process. Student attention was focused on this feature because of the design of the animation, which cued students to notice this feature by significantly varying the appearance of the Shine–Dalgarno sequence.
Introduction
Educators have long relied on images, diagrams, models, and other external representations to illustrate a collective, common meaning of specific content (Pinar et al., 2004). The imagery of science presented in the classroom, informal learning environments, the media, and popular culture can convey fundamental ideas about science, thereby allowing students to construct a deeper understanding of a particular topic. However, students may develop varied understandings of the same image based on variations in their level of prior knowledge of the material (Cook, 2006; Cook et al., 2008; Harle and Towns, 2012a; Sim and Daniel, 2014). For example, previous research has shown that students with lower levels of prior knowledge tend to focus on superficial aspects of external representations (Heyworth, 1999; Lowe, 2003, 2004); and the mental models they develop from those representations tend to be overly simplistic (Snyder, 2000).External representations in science range from symbolic representations of chemical equations to graphical diagrams of empirical data, from static images to dynamic animations, from simplistic illustrations to complex multimedia presentations. External representations have the potential to make the unseen, seen; they can depict intricate relationships or deconstruct complicated processes; they can often negate the constraints of size, time, and space.
“Scientific images do not, of course, aim to record what is visible, their purpose is to make visible. This applies to the ordinary enlargement as well as to the miracle of the electron-scanning microscope which has enabled scientists to answer so many questions—always presupposing that they know the specifications of the instrument, its magnification, power of resolution, and so on.” (Gombirch, 1980, p. 185)
External representations have been shown to be an extremely efficient means of communication “if the viewer understands the rules of construction” (Perino, 2001, p. 16). A photograph, for example, is an image in the same way a great painting is an image; they are both subject to distortion and interpretation and separate from the physical reality they are meant to capture, depict, or evoke. However, if the viewer understands the constraints of the medium, they are able to draw conclusions from the image (Gombirch, 1980). Similarly, scientists have used instrumentation to create artifacts—external representations—and make observations of phenomena that could otherwise not be recorded or observed. As with a photograph, the external representations generated through the use of scientific instrumentation aim to capture some aspect(s) of an object or phenomenon as filtered through the instrument and presented in a chosen medium and format. In the same way that an electron micrograph is a representation of a molecule, thereby making the data collected by an electron microscope visible to a scientist, external representations—broadly defined—can be used in an educational setting to make non-experiential scientific concepts visible to a student. However, a teacher's ability to effectively use external representations and students' ability to understand and make meaning from those representations can be affected by an array of factors, such as students' prior knowledge, the type (mode) of representation, or the level of abstraction depicted in the representation (Cook, 2006; Schönborn and Anderson, 2006; Cook et al., 2008; Harle and Towns, 2012a, 2012b, 2013). In essence, external representations are only as useful as students' ability to understand them.
Visual literacy and external representations in biochemistry education
In step with the development of new technologies, the use of educational imagery—imagery used in an educational context to convey information—has grown exponentially (Schönborn and Anderson, 2006). Computer technologies and Internet applications have become ubiquitous in the educational landscape (Gray et al., 2010). Moreover, these technologies are increasingly being used to design more intricate and complex representations. For example, movie animation technology has fostered the development of molecular animations of biological phenomena (Olsen, 2010). In particular, biochemistry educators have become highly dependent on a variety of external representations—including, for example, symbolic representations and reaction schemes—for both instructional and assessment purposes (Linenberger and Holme, 2014, 2015).Avgerinou and Ericson (1997) and others have argued that visual literacy—the ability to interpret and use external representations—is a skill that must be explicitly “identified and taught” to biochemistry students (p. 288). Biochemists demonstrate expert visual literacy by decoding, evaluating, interpreting, manipulating, and constructing external representations in order to explore and explain biochemical and cellular phenomena (Schönborn and Anderson, 2006, 2010). They also display other cognitive skills related to visual literacy, including the ability to translate between multiple external representations and between the various levels of organization, as well as the ability to “visualize orders of magnitude, relative size, and scale” (Schönborn and Anderson, 2010, p. 349). Clearly, the ability to construct meaning from visual representations is a necessary skill for biochemists. It is also, therefore, a necessary skill for biochemistry students, who will be presented with a large number and varying types of representations over the course of their educational careers. However, biochemistry instructors often do not make these skills explicit to students (Linenberger and Holme, 2015). As a consequence, students often interpret the external representations presented to them in their biochemistry classes in a manner different than that which was intended by their instructors (e.g., Harle and Towns, 2013). In the current study, we examine students' interpretations of an animation of protein translation and propose an explanation as to why students may be focusing on certain aspects of the external representation over others.
In order to convey the structure and function of proteins and their associated biochemical pathways—including protein translation—to students, graphic designers, textbook authors, and other instructional materials designers rely on external representations. However, the variety of types of external representations and the range of symbolism used to represent biochemical phenomena pose an obstacle to biochemistry students (Schönborn and Anderson, 2006). For example, Schönborn et al. (2002) asked 151 students, most of who were second year biochemistry students, to explain a textbook diagram of an immunoglobin G (IgG) protein. The researchers assessed student understanding of the diagram and its content through a series of written probes. Subsequently, ten participants volunteered to be interviewed. They concluded that students displayed several difficulties in understanding the diagram, including difficulties interpreting the dynamic aspect of the process being depicted as well as the structural features of the IgG protein. They noted that a possible source of student difficulties “is the fact that biochemistry textbooks often use more than one convention to represent a single structural feature of a molecule […]” (Schönborn et al., 2002, p. 96). They observed that the diagram of IgG they showed to students used a black line to represent a disulfide bond, whereas, other diagrams use a yellow line or –S–S– to represent a disulfide bond. The researchers also noted that, in the diagram, straight black lines represented not only disulfide bonds, but also the polypeptide chains of IgG. If students confused the disulfide bonds for peptide chains or vice versa, this would significantly alter their understanding of the structure of IgG. Therefore, the researchers concluded that one of the factors contributing to student difficulties in understanding the external representation was students' inability to decipher the symbols used to depict the structural features of the IgG protein.
Other research has also demonstrated that students' inability to correctly decipher the symbols and conventions of scientific diagrams negatively affects students' ability to interpret and use those diagrams. For example, Harrison and Treagust (1996) suggest that students may develop alternative conceptions of atoms due to inaccurate decoding and interpretation of external representations of atoms. It is, therefore, reasonable to assume that students' inability to decipher the symbols and conventions in representations of biochemical processes, like protein translation, will likewise influence what students can and do learn from these representations.
Purpose of the current study
Even though protein translation is an important foundational biochemical concept, the authors have not been able to find any study that specifically examines students' understanding of external representations of protein translation. The study reported here is part of a larger project about what students can learn about protein translation from common external representations of the process (Bussey, 2013). The focus of the current study was to answer the research question “Why do students learn about certain features of protein translation depicted in a given representation and not others?” Here, we present students' interpretations of a particular animation of protein translation and propose reasons why students may have focused on one feature of the animation more than they focused on others. There are three main sources of data that will be described in this study: the selected animation of protein translation, interviews with biochemistry instructors, and interviews with biochemistry students. For our purposes, we will focus only on the data that is relevant to students' interpretation of the specific animation (see Bussey, 2013, for a more detailed description of the larger data set).Methodology
The theoretical framework used in this study is variation theory. Studies informed by variation theory look at why people experience a phenomenon differently, i.e., what causes the variation in perception of a phenomenon? For any given phenomenon, there are many different aspects to which an individual could pay attention. The individual's experience with a given phenomenon depends on the particular set of aspects to which they attend. Thus, any variation in the aspects of a phenomenon to which an individual attends will also cause variation in the way the individual perceives that phenomenon. Moreover, the more a particular aspect of a phenomenon is varied, the more likely an individual is to notice that aspect (see Bussey et al., 2013, for a more detailed description of variation theory).In the context of educational research, a study informed by variation theory focuses on an object of learning, in other words, a phenomenon that is to be learned by a student. The object of learning for the current study is protein translation. The object of learning is examined from three different perspectives. The intended object of learning focuses on what students should learn about protein translation as defined by the instructor. The enacted object of learning focuses on what students can learn as constrained by the learning environment. In this case, the learning environment is defined as the space of learning created by the animation of protein translation presented to students. Finally, the lived object of learning focuses on what students actually learn as described by the students. This framework was particularly useful to examine why students seemed to be learning about certain features depicted in a given external representation because it allowed us to triangulate instructors' perceptions with the features found in the representation and with students' perceptions of protein translation.
In order to determine how biochemistry students should perceive a particular external representation of protein translation—or what they should pay attention to in the external representation—data was obtained from interviews with five biochemistry instructors from two universities. Two of the instructors were from a large Midwestern university, and three instructors were from a large Southwestern university. All instructors were university faculty members who had taught at least one college-level biochemistry course that had addressed the topic of protein translation. The instructors interviewed had an average of 20.6 years of teaching experience.
All interviews were semi-structured and lasted about an hour. The instructor interview protocol (see Appendix A: Instructor Interview Guide) was adapted from Schönborn and Anderson's Three-Phase Single Interview Technique (3P-SIT) (Schönborn et al., 2007; Schönborn and Anderson, 2009). The 3P-SIT model was chosen because it addresses both participant knowledge of the content and their understanding of external representations depicting the content. All interviews were audio-taped and transcribed verbatim. As the intention was to describe instructors' perceptions of what students should learn about translation from the animation, a grounded theory approach was used to analyze this data. Interview transcripts and artifacts were coded to identify critical features of the conceptual knowledge of translation and the animation, i.e., information that could be used to answer the following questions: what should students know about translation, and what should students know in order to understand this external representation of translation? These coded features were used as the basis for the subsequent analysis of the external representation and the student interviews.
The coding scheme used in this study was developed and revised using the constant comparative method throughout the coding process. Highly similar codes were collapsed until each defined code described a unique feature of protein translation. The coding scheme was validated using an external coder. The external coder was a postdoctoral researcher with expertise in biochemistry education. She coded excerpts from the instructor interviews using a code list developed by the first author. She was asked to apply codes as they were defined on the code list and to identify any areas of confusion or possible new codes. Inconsistent coding and ambiguous code definitions were revised in collaboration with the external coder. A final revised code list containing an operational definition of each code was then created and can be found in Appendix D.
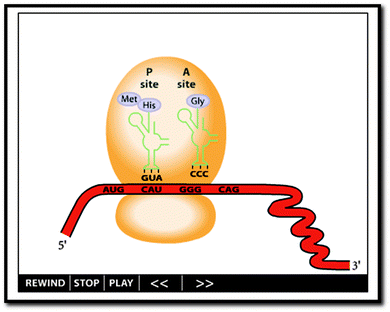
In order to define the learning environment for the biochemistry students interviewed in this study, a stylized dynamic external representation (referred to from here on out as “the animation”) was selected using input from the interviewed biochemistry instructors (Fig. 1, http://www.biostudio.com/demo_freeman_protein_synthesis.htm). The animation was presented to participants during the interview without auditory cues. In other words, narration and other presentation sounds were not presented to participants, so as to focus participant attention on the visual information and to eliminate potential complicating variables introduced by variations in narrated text and sound.
In order to determine how students interpreted and what they learned from the animation, data was collected from interviews with twenty undergraduate second-semester biochemistry students from a large Southwestern university. Students were recruited during the semester in which they were enrolled in a second semester biochemistry course and participated on a voluntary basis. Students were recruited in class, with the instructor's permission, following their exam that covered protein translation. The second semester biochemistry course was chosen specifically because this was the first time students had been exposed to a description of protein translation in the context of a biochemistry course. Previous discussions of protein translation in high school and/or undergraduate biology courses would have exposed students to a cursory explanation of translation from a biological perspective; however, the biochemical components and interactions of this metabolic pathway are discussed in greater detail in a second semester biochemistry course. Additionally, all students enrolled in the second semester biochemistry course had met the course prerequisites. Specifically, students had taken first semester biochemistry; and had, therefore, been exposed to an introduction to biochemistry. Also, student participants had taken similar amounts and types of undergraduate coursework. This ensured that all students had been exposed to a similar level of prior instruction regarding the components and processes of protein translation.
The semi-structured student interview protocol for students (see Appendix B: Student Interview Guide) was also adapted from Schönborn and Anderson's Three-Phase Single Interview Technique (3P-SIT) (Schönborn et al., 2007; Schönborn and Anderson, 2009). Students were asked to describe their understandings of translation before, during, and after they were shown the animation. This allowed us to assess the students' prior knowledge of translation as well as their interpretations of the animation and what they were able to learn from the animation.
Because students' construction of knowledge is influenced both by a learning event (in this case, being exposed to the animation of protein translation) and their prior knowledge, we felt that it was essential to determine what students knew about protein translation before being exposed to the animation during the student interviews. All student participants had some level of prior knowledge of the concept of protein translation. In order for us to determine if students learned from their experience with the animation, we first needed to account for what they already knew about protein translation before being exposed to the animation.
If students already knew about translation, what do we mean when we say that students learned from their experience of the animation, though? Marton et al. (1993) have identified six conceptions of learning. They describe learning as (1) increasing one's knowledge, (2) memorizing and reproducing, (3) applying, (4) understanding, (5) seeing something in a different way, and (6) changing as a person. Within the context of this project, we will use the term learning to refer to seeing something in a different way. This means that although students may have already had prior knowledge pertaining to an object of learning (in this case, protein translation), if their experience of the animation allows them to see the object of learning in a different way, then we claim that students are learning. Thus, while students may already know about protein translation, their ability to recall the details about this object of learning, as defined by their instructors, may be limited and cursory. If the animation of protein translation allows students to see translation in a different way, students are learning. Specifically, we measured learning as a progression of students' stated or depicted knowledge towards a more scientifically accurate understanding of protein translation.
All student interviews were audio-taped and transcribed verbatim. Any artifacts created by the students during the interview were also collected. In particular, students were asked to draw their initial and final understandings of protein translation. Prior research has shown that having students draw their understanding of biochemical structures and interactions is an effective method for exploring their understanding of fundamental biochemical concepts (Harle and Towns, 2012b, 2013). These drawings were collected through the use of a Livescribe Echo™ Smartpen (Linenberger and Bretz, 2012).
Interview transcripts and artifacts were coded for critical features of the conceptual knowledge of translation and the animation, i.e., what did students notice about translation, and what did students notice about the representational design of the animation? The critical features of students' conceptual knowledge of translation before and after exposure to the animation were compared in order to assess the degree of learning. Finally, the critical features identified by students were compared to the critical features identified by the instructors. Alignment between instructors' and students' critical features was used to assess the degree to which students held an expert view of translation and the effect of the animation in promoting an expert view of translation. As mentioned previously, the study described here is part of a larger project. The data presented in the sections that follow are limited to those that describe what students should, could, and did learn about protein translation from one particular animation of the process. Specifically, we will focus why students learned more about the Shine–Dalgarno sequence than any other feature presented in the animation and the educational implications of this finding.
Results and discussion
Students could have demonstrated learning about a variety of features of protein translation as a result of exposure to the animation. Instructors identified 16 primary critical features that students should learn about protein translation, most of which focused on structural aspects of the component parts of translation (Table 1). Primary critical features were defined as components or processes of protein translation that all five instructors described as important features that students should learn about and included biochemical structures that participate in translation—such as the large and small subunits of the ribosome and the codon on an mRNA molecule—and interactions between structures that participate in translation—such as the interaction between a codon on an mRNA molecule and an anti-codon on a tRNA molecule.| Theme | General feature | Coded feature |
|---|---|---|
| a Note: primary critical features were identified as those features described unanimously by the instructors interviewed in this project. Other features were identified by some of the instructors but not all and were not classified as primary critical features. All critical features can be found in Appendix C. Definitions of all coded features can be found in Appendix D. | ||
| Component/structure | mRNA | General molecule M |
| Codon | ||
| Ribosome | General molecule R | |
| Large subunit | ||
| Small subunit | ||
| Aminoacyl (A) site | ||
| Peptidyl (P) site | ||
| Exit (E) site | ||
| tRNA | General molecule(s) T | |
| Anti-codon loop | ||
| P-site tRNA | ||
| Amino acids | General molecule AA | |
| Polypeptide chain | General molecule P | |
| Interactions/chemistry | Initiation | Codon/anti-codon base pairing I |
| Elongation | Peptide bond formation | |
| Codon/anti-codon base pairing E | ||
Forty-three additional critical features, including the Shine–Dalgarno sequence, were also identified. These additional critical features were categorized as secondary or tertiary critical features to indicate a decrease in the level of agreement amongst the instructors interviewed in this study. Secondary critical features (2°) were described by a majority (3–4) of instructors but not all. Tertiary critical features (3°) were described by a minority (1–2) of instructors (see Appendix C, the Categorization of Critical Features of Protein Translation, Appendix D for the definition of all codes, and Bussey, 2013, for a more detailed description of these critical features). Of the 59 critical features instructors identified as being important for students to learn about protein translation, the animation only presented students with 41 features (Table 2). Thus, exposure to the animation could potentially cause students to learn about the 41 features present in the animation. On the other hand, exposure to the animation would not be expected to cause students to learn about the 18 features that were not present in the animation.
| Theme | General feature | Coded feature |
|---|---|---|
| a Note: definitions of all coded features can be found in Appendix D. | ||
| Components/structure | mRNA | General molecule M |
| Codon | ||
| Nucleotide sequence (multiple codons) | ||
| Shine–Dalgarno sequence | ||
| 5′ end M | ||
| 3′ end M | ||
| Start codon | ||
| Stop codon | ||
| Nucleotide sequence (start codon) | ||
| Nucleotide sequence (stop codon) | ||
| Ribosome | General molecule(s) R | |
| Large subunit | ||
| Small subunit | ||
| Aminoacyl (A) site | ||
| Peptidyl (P) site | ||
| 16S rRNA | ||
| tRNA | General molecule(s) T | |
| Anti-codon loop | ||
| P-site tRNA | ||
| A-site tRNA | ||
| E-site tRNA | ||
| 2D shape | ||
| Nucleotide sequence (anti-codon loop) | ||
| Amino acids | General molecule(s) AA | |
| Methionine | ||
| Sequential AA | ||
| Polypeptide chain | General molecule P | |
| Primary structure | ||
| Release factors | General molecule(s) RF | |
| Interactions/chemistry | Initiation | Codon/anti-codon base pairing I |
| General process I | ||
| Hydrogen bonding (mRNA/ribosome) I | ||
| Initial tRNA/ribosome/mRNA | ||
| Elongation | Codon/anti-codon base pairing E | |
| General process E | ||
| Incoming tRNA/ribosome/mRNA | ||
| Exiting tRNA/ribosome/mRNA | ||
| Ribosomal translocation | ||
| AA/AA interaction | ||
| Termination | General process T | |
| General considerations | Reaction kinetics | |
Of the 41 critical features presented in the animation, students exposed to the animation demonstrated learning about 23 of those features (Table 3). In other words, although students had not originally included these features in their descriptions/depictions of translation, they did so after being exposed to the animation.
| Theme | General feature | Coded feature | Percentage of students who learned about the feature after viewing the animation (%) |
|---|---|---|---|
| a Note: definitions of all coded features can be found in Appendix D. | |||
| Components/structure | mRNA | Shine–Dalgarno sequence | 25 |
| 5′ end M | 5 | ||
| 3′ end M | 5 | ||
| Start codon | 15 | ||
| Stop codon | 10 | ||
| Nucleotide sequence (start codon) | 10 | ||
| Ribosome | Aminoacyl (A) site | 5 | |
| Peptidyl (P) site | 5 | ||
| 16S rRNA | 5 | ||
| tRNA | 2D shape | 5 | |
| Amino acids | General molecule(s) AA | 5 | |
| Methionine | 5 | ||
| Sequential AA | 5 | ||
| Release factors | General molecule(s) RF | 15 | |
| Interactions/chemistry | Initiation | Codon/anti-codon base pairing I | 5 |
| General process I | 10 | ||
| Elongation | Incoming tRNA/ribosome/mRNA | 5 | |
| Exiting tRNA/ribosome/mRNA | 5 | ||
| Ribosomal translocation | 15 | ||
| AA/AA interaction | 10 | ||
| Termination | General process T | 15 | |
| General considerations | Reaction kinetics | 5 | |
While students demonstrated learning about a variety of features present in the animation of protein translation, certain features, such as the Shine–Dalgarno sequence, were more likely to be noticed and learned about than others (see Table 3 for a list of the percentage of students who were shown the animation and demonstrated learning about specific critical features). Why is this? Why would students be more likely to demonstrate learning about one feature—the Shine–Dalgarno sequence—over others? What we will show in the following sections is that students were able to demonstrate learning about the Shine–Dalgarno sequence and propose reasons why students may have preferentially learned about this particular feature over others.
Students demonstrate learning about the Shine–Dalgarno sequence from the animation
As noted earlier, this study is part of a larger project in which students were exposed to several external representations of protein translation, including the animation discussed here. Some critical features of protein translation were presented exclusively in one representation. Therefore, any inclusion of these features in students' final descriptions of translation could be attributed to that particular representation as long as that student did not initially describe that feature prior to viewing the representations. Such is the case with the Shine–Dalgarno sequence. This feature was only explicitly presented in the animation we discuss here, and only students who had been exposed to the animation demonstrated learning about this feature.In order to demonstrate learning about this feature, students first had to demonstrate a lack of or inappropriate knowledge of the Shine–Dalgarno sequence. For example, Student 25(B) initially made no reference to the Shine–Dalgarno sequence in his description or drawing of initiation (Fig. 2).
Student 25(B): “Ok, so you have a smaller part of the ribosome down here [draws the small subunit on the bottom], which is going to be like a 30S here [labels the small subunit], and then you have a larger one up here [draws the large subunit on the top], which is like a 50S [labels the large subunit]. Um, and your mRNA is going to be through here [draws mRNA strand between large and small subunits] in this direction. What ends up happening is, ah, this is what I was saying, the initiation factors, and there's 3 or 4 of them around, and I don't remember the names [labels IF = 3–4 indicating the number of initiation factors involved]. Ah, they're going to come in and bind the mRNA to this small subunit before the 50S is able to come down because if they clamp down before [referring to the subunits coming together], from what I know you wouldn't be able to have the mRNA come in.”
Although he mentions the binding between the mRNA and the small ribosomal subunit, he does not identify the role of the Shine–Dalgarno or any other nucleotide sequence in the assembly process. However, while viewing the animation, he identifies not only the presence of the Shine–Dalgarno sequence depicted in the animation but describes its role in orienting the ribosome at the proper start site for translation to begin.
Student 25(B): “[describing the animation as it is playing] Ok, so you've the small, um, part of the ribosome that's binding to the mRNA which bends the mRNA. […] It goes to find the specific sequence, in this case the Shine–Dalgarno or start site, to know where to actually start the translation […].”
This discussion of the specific role of the Shine–Dalgarno sequence in the protein translation process indicates that Student 25(B) was previously aware of this feature; however, his lack of inclusion of this feature in his initial description suggests that he did not initially find the Shine–Dalgarno sequence to be a salient feature of the process. After watching the animation, he includes the Shine–Dalgarno sequence in both his verbal description as well as in his drawing of initiation.
Researcher: “So we're going to come back to your drawing over here. So I'm going to have you, one more time, just re-explain to me the process of translation. You can add, change, edit, keep the same anything you did before.”
Student 25(B): “Ok, um, I don't know. I'm pretty happy with my explanation here. […] I think the only thing that I didn't really mention was once you have the large subunit and small subunit bind or bound together, um, your mRNA has, like I said, the Shine–Dalgarno or a, or a start site, something that triggers the actual start of [translation].”
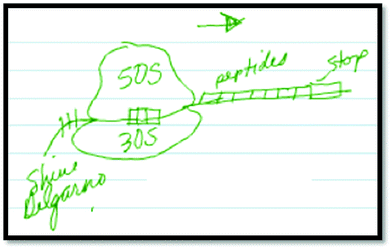
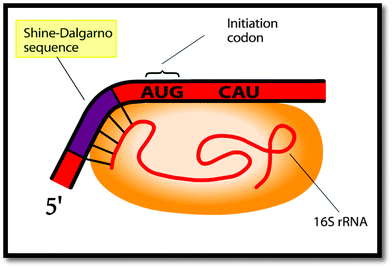
Although not quite accurately, he then draws in the Shine–Dalgarno sequence as part of his final drawing to explain the alignment of the ribosome on the mRNA (Fig. 3). He goes on to label the Shine–Dalgarno sequence in a similar fashion to the animation (Fig. 4).
It is arguable whether or not this student has gained a greater understanding of the process of translation or more specifically the role of the Shine–Dalgarno sequence in the initiation of the process of translation as a result of his experience of the animation. However, the inclusion of the Shine–Dalgarno sequence in his final explanation indicates that he perceives this feature as more important, and therefore worthy of inclusion, in his description of the process as compared to his initial explanation.
The design of the animation influences students' perception of which features are important to learn
Although many students demonstrated learning about the Shine–Dalgarno sequence, the instructors generally did not feel that this feature was necessary for students to learn in order to demonstrate a sound biochemical understanding of protein translation. So why did so many students pick up on a feature that the instructors did not feel was that important? Instructor 5 notes that the animation highly emphasized the alignment of the mRNA and small ribosomal subunit through the Shine–Dalgarno sequence interaction and that this emphasis would cause students to notice this feature.Instructor 5: “The best part of [the animation], though, had nothing to do with translation itself, but was the locking of the Shine–Dalgarno sequence to the 16S ribosomal RNA in the small subunit. That's really, that was very good. Yeah, that was nicely done. That was very clear, and that, that made a lot of sense.”
Researcher: “If a student were watching this, what do you think they would take away from it?”
Instructor 5: “That. They would, they would like that. It had a nice emphasis. Yeah it was very clear. They actually stalled out on it a little bit and they go ‘oh yeah I see all the hydrogen bonding between them.’”
According to this instructor, the design of the animation—the way in which features have been presented—is cueing students to notice this non-primary critical feature.
As previously noted, an individual's experience with a given phenomenon is dependent on the features to which they attend. According to variation theory, people attend to features that vary more than they attend to features that do not vary. There are several ways that students' attention could be drawn to particular features of the animation—or to features of any external representation. In the current study, variation in the position, size, or labeling of a depicted feature were all identified as possible means by which to draw students' attention to those features. Given that students demonstrated learning about the Shine–Dalgarno sequence more than they learned about any other feature presented in the animation, we would expect that the presentation of this feature is highly varied in the animation. Indeed, this is the case. Table 4 lists the mean percent variation in position, size, and labeling for the top five features students demonstrated learning about after viewing the animation.
| Coded feature | Percentage of students who learned about the feature after viewing the animation (%) | Number of frames in which the feature appears (out of 43 total frames) | Mean percent variation in position, size, and labeling in the frames in which the feature appears (%) |
|---|---|---|---|
| a Note: ‘Frames’ refers to snapshots of the animation taken every 2 seconds of the animation. | |||
| Shine–Dalgarno sequence | 25 | 7 | 73 |
| Start codon | 15 | 32 | 52 |
| Release factors | 15 | 8 | 68 |
| Ribosomal translocation | 15 | 20 | 32 |
| Termination | 15 | 5 | 32 |
Although the Shine–Dalgarno sequence is shown in only seven of the 43 frames of the animation, its presentation is the most highly varied. On average, the position, size, and/or labeling of the Shine–Dalgarno sequence varies 73% of the time in the frames in which it appears. This high level of variation in the presentation may have cued students to pay attention to this particular feature (Bussey et al., 2013). In the sections that follow, we examine the animation in more detail in order to determine which of these variations might have caused students to preferentially notice—and learn about—the Shine–Dalgarno sequence.
Instructor 3: “[In response to the animation] I like the fact, though, that, that it has more, it's, it's moving along the mRNA, so you can see one amino acid at a time, one code being deciphered at a time. […] That's actually nice. If you have multiple steps, you can see one step at a time.”
Thus, as suggested by Instructor 3, as features or structures move in, around, and out of the animation, the variation in position could potentially cue students to notice those features. However, the depiction of the Shine–Dalgarno sequence in the animation is predominantly stationary. The animation begins with the mRNA in a stationary position as the small ribosomal subunit moves towards the mRNA. To indicate the alignment of the Shine–Dalgarno sequence with the 16s rRNA, the animation then shows the 5′ end of the mRNA moving down slightly to fit the curve of the small ribosomal subunit. The limited movement of the Shine–Dalgarno sequence in the animation might not significantly cue students to notice this feature. Other aspects of variation, then, should account for the increase in students’ learning about the Shine–Dalgarno sequence.
 | ||
| Fig. 5 The zoom-in effect depicted during the animation: frames 7–9. Note: ‘Frames’ refers to snapshots of the animation taken every 2 seconds of the animation. | ||
Instructor 3: “[In response to the animation] The, the binding to the, to the beginning of the mRNA, it's the Shine–Dalgarno sequence, eh, eh, it's too much. There's now too much information in there, which distracts from the most important part of the system. [Students] would think this is the most important part, and now that part, which is the actual reaction of the system, is not shown very well.”
Although Instructor 3 does not like that the animation has zoomed in on the particular interaction between the Shine–Dalgarno sequence and the small ribosomal subunit, he acknowledges that this change in the size of the component features would cause students to attend to those features and potentially notice them more after they have returned to their original size because their change in size has indicated that they are an important component of the representation. He goes on to note that the interaction he feels is more important, peptide bond formation, is not emphasized in the animation. To remedy this lack of focus on the chemistry, he suggests that a second zoomed-in scene be added to the animation in order to focus students' attention on that interaction as well.
Instructor 3: “[In response to the animation] I would have included, at least in the first peptide, peptide bond synthesis, a blow up showing how the reaction actually happens.”
Instructor 2 makes a similar recommendation when asked how the animation could be improved to focus students' attention on features he deemed to be important.
Instructor 2: “[In response to the animation] The first time one of [the amino acids] moves over, maybe have a frame come up and, and hone in on that and show the chemistry, show why it's moving over. Um, that might be a nice thing to do. […] In terms of the basics, though, [the animation is] not bad. Um, it might be, the only things I would add is, is a frame that really showed the chemistry going on.”
Thus, change in size was identified by the instructors as an aspect of variation of the animation that would call students' attention to particular components or interactions of protein translation, important or otherwise. In the case of the Shine–Dalgarno sequence, the instructors noted that the zoom in effect would focus students' attention on this feature even though they didn't consider this to be an important feature to be learned.
Students also discussed the “zoom in” representational strategy. For example, Student 4(A) described that he did not like the animation because it because it didn't show the mechanism of peptide bond formation and suggested including a zoom in feature in the animation that showed more of the “actual chemistry.”
Student 4(A): “If you were able to see the actual, like, chemistry binding, like they zoomed in and then showed that it was binding to it and then it transfers instead of just putting in the next tRNA and it just moves and then it pops off that one and move to the next one. It kind of leaves you with ‘well how did that happen?’”
Interestingly, though, the Shine–Dalgarno sequence is not the only feature cued in this way in the animation. The 16S rRNA is also zoomed in on; however, no students described or drew the 16S rRNA in their final descriptions. This is interesting because the 16S rRNA is the ribosomal complement to the Shine–Dalgarno sequence, yet it is not being learned to the same extent as the Shine–Dalgarno sequence. Simply zooming in on a particular component might not be sufficient to draw students' attention to that component. Many students, like Student 25(B), seemed to be familiar with the Shine–Dalgarno sequence and less familiar with the 16S rRNA. Therefore, perhaps the zooming-in, combined with students' undeclared prior knowledge of the Shine–Dalgarno sequence, is leading to the incorporation of that feature into students' final descriptions of protein translation.
Instructor 1: “Uh one of things that I discovered about [representations] that distinguishes good ones from bad ones is that […] students really like it if a process, a complicated process like translation, is, uh, described in numbered steps, so your eyes look at this complicated piece of art and it follows, and at each point you are able to say yea I see, I see that. […] Your eye just can't be every place all at the same time, and, uh the temptation with modern molecular graphics is that they're so damned beautiful that you try to show everything all at once, and it doesn't work.”
We observed that the individual components/structures depicted as the animation zooms in on the initial interaction between the mRNA and small ribosomal subunit were labeled. However, instead of labeling all of the features at the same time, the animation only labels three features at any one time. Those labels were then individually highlighted in yellow, potentially calling attention to each component of the interaction and cueing students to notice those features. In the case of the Shine–Dalgarno sequence, this label was shown in the upper left-hand corner and highlighted first (see Fig. 4). This may be seen as giving the Shine–Dalgarno sequence priority over the other features labeled around the same time, including the 16S rRNA which was highlighted last and with a label shown in the lower right-hand corner of the frame. It is possible that the sequence and relative position of the labeling may have altered the perceived importance of that information and may provide part of the reason why students demonstrated more learning about the Shine–Dalgarno sequence than about the 16S rRNA. Further research is needed to examine why some highly cued features, such as the Shine–Dalgarno sequence, were more frequently articulated in students' final descriptions or depicted in students' final drawings while other similarly cued features, such as the 16S rRNA, were not.
Conclusions
External representations in biochemistry, and more broadly in science, education have the potential to be powerful teaching tools. For example, previous research has shown that external representations can be used to help students integrate new knowledge with prior knowledge and make abstract content more coherent (Winn, 1991). Similarly, research has shown that external representations can be a valuable tool for students to construct new or more intricate knowledge (e.g., Harrison and Treagust, 2000) and that “certain diagrams can help individuals to reason more rapidly and more accurately” (Bauer and Johnson-Laird, 1993, p. 378). As such, external representations can be an extremely beneficial resource to allow students to develop their conceptual knowledge and reasoning abilities.However, research has also shown that the results of using external representations are not always positive. For example, Lowe (1996) notes that differences in content knowledge can lead to differences in understanding of a content-related representation. He goes on to suggest that the representational design of the diagram “implies [that] viewers possess appropriate background knowledge concerning the depicted situation” (Lowe, 1996, p. 377). If viewers fail to possess the “appropriate background knowledge,” it is not simply that they will not be able to understand the representation; instead, viewers can develop alternative understandings of what is being depicted in the representation.
From the results of the current study and the larger study to which it pertains, it appears that external representations are most useful at supporting student learning about biochemical concepts when students notice the critical features of the object of learning—those that are necessary to develop correct understandings of the concept—and pay less attention to features of the representation that are not critical to developing correct understandings of the biochemical concept. Unfortunately, the results of the current study also suggest that variations in the way features are depicted, such as the zooming in or labeling of features, may cue students to pay attention to particular features of the animation, whether or not these features are important to an understanding of the portrayed process, as in the case of the Shine–Dalgarno sequence. It is possible that students with low prior content knowledge or who lack the knowledge of the symbols and conventions used in biochemical representations may instead pay attention to the salient features of the representation emphasized by variation whether instructors want them to pay attention to those features or not.
These findings are consistent with variation theory in that features depicted in external representations that exhibit more variation are more likely to be noticed and learned by students. In particular, the types and context of the variation seems to preferentially cue students to certain critical features. Therefore, instructors should give careful consideration to how and when features of a representation are varying and should design or select representations that align with their intentions for student learning. If variation might help a student to learn about a critical feature, then that variation is well utilized within the educational objectives for the representation. However, if a representation displays variation in features that are not deemed critical, then a less distracting representation might need to be sought out in order to help best direct students' attention on the features of most importance to the instructor. Overall, it is important that instructors align their intentions for student learning with the possibilities for learning created by the external representation, a thought best articulated by Instructor 2.
Instructor 2: “I think the key thing is trying to design [a representation] so students can get the information that they're supposed to get out of [it].”
Appendix A: Instructor Interview Guide
Demographics
• Tell me a little bit about yourself.– Where did you go to school?
– How long have you been at (the school name)?
– Tell me a little bit about your research. What are you currently working on?
– Tell me about your teaching experience. What courses do you teach?
Phase 1 – intended object of learning of translation
• Today I would like to talk about translation. Some people might refer to it as protein synthesis … [long pause] … can you explain to me how the process works? … What are the essential components of the process? Feel free to draw if that makes it easier to explain. [Paper and pencil will be provided to all interviewees.]• Do you teach (or have you taught) translation?
• If you were going to explain this process to your students, how would you go about doing so? What things would you emphasize to them so they understand the process?
• What do you see as being the main idea? What's the “big thing” students should come away with after learning about translation?
• What prior knowledge do your students need to have in order to understand translation?
• What difficulties do you think students might encounter when trying to understand this topic?
• Do you think translation is an important biochemical process for student to understand? Explain your answer.
Phase 2 – instructor evaluation of the static external representations of translation
• I'm going to show you a series of external representations. An external representation is an image or animation, just some type of representation. I'm going to show them to you two at a time. For each set, I would like you to pick the one that you would show to your students to explain translation … so here's #1 … and here's #2 … which one would you choose to show your students? Why did you choose # ____? What didn't you like about the other representation?• Consider the external representation you didn't like.
– Why didn't you like it?
– What could students learn from that representation?
– What things would they notice? (What would stick out to them?)
– What would you change about this representation to make it better?
• Consider the external representation you did like.
– Why did you like it?
– What could students learn from this representation?
– What things would they notice?
– What could students learn from this representation that they wouldn't get from the other one?
– Is there anything on this representation in particular that you don't like or think might be confusing to students?
– What do you think this external representation is not showing? Explain your answer.
– Consider yourself a graphic designer or textbook author. If you could change this external representation in any way, what would you do to improve it, if anything?
Phase 3 – instructor evaluation of the stylized, dynamic external representations of translation
• Again, for this set, I would like you to pick the one that you would show to your students to explain translation … so here's #3 … and here's #4 … which one would you choose to show your students? Why did you choose # ____? What didn't you like about the other representation?• Consider the external representation you didn't like.
– Why didn't you like it?
– What could students learn from that representation?
– What things would they notice? (What would stick out to them?)
– What would you change about this representation to make it better?
• Consider the external representation you did like.
– Why did you like it?
– What could students learn from this representation?
– What things would they notice?
– What could students learn from this representation that they wouldn't get from the other one?
– Is there anything on this representation in particular that you don't like or think might be confusing to students?
– What do you think this external representation is not showing? Explain your answer.
– Consider yourself a graphic designer or textbook author. If you could change this external representation in any way, what would you do to improve it, if anything?
Phase 4 – instructor evaluation of the realistic, dynamic external representations of translation
• For the last set, I would like you to again pick the one that you would show to your students to explain translation … so here's #5 … and here's #6 … which one would you choose to show your students? Why did you choose # ____? What didn't you like about the other representation?• Consider the external representation you didn't like.
– Why didn't you like it?
– What could students learn from that representation?
– What things would they notice? (What would stick out to them?)
– What would you change about this representation to make it better?
• Consider the external representation you did like.
– Why did you like it?
– What could students learn from this representation?
– What things would they notice?
– What could students learn from this representation that they wouldn't get from the other one?
– Is there anything on this representation in particular that you don't like or think might be confusing to students?
– What do you think this external representation is not showing? Explain your answer.
– Consider yourself a graphic designer or textbook author. If you could change this external representation in any way, what would you do to improve it, if anything?
Phase 5 – instructor perceptions of external representations of translation
• So you chose #s _____, _____, and _____. Having now seen all of the representations, would you change anything about your responses?• If you were to show your students all three external representations, what order would you show them in? Why?
• Do you think order matters? In other words, do you think student's understanding would change if you changed the order in which they see them? Why or why not?
• If you had to choose only one external representation to show to your students to explain translation, which one would you choose? Why # _____?
• Do you think students' understanding would change if the type of representation changed? In other words, do you think students would understand translation differently if they were only shown an illustration or an animation, or if they were only shown a stylized representation or a realistic one? Why or why not?
• What would your ideal representation for teaching translation look like? Don't feel bound by traditional presentation formats like textbooks or powerpoint slides. If anything were possible, how would you show students protein translation?
• Do you have any final thoughts about external representations or translation? Thank you so much for your time today.
Appendix B: Student Interview Guide
Demographics
• Tell me a little bit about yourself.– What year are you in school? (Sophomore, Junior, …)
– What is your major?
– What do you want to do after graduate?
– Have you taken Biochem II? If so, when?
– What are your feelings towards biochemistry? Why do you say that?
– How is biochemistry similar to other science classes you have taken? How is it different?
Phase 1 – students' prior knowledge of translation (prior lived object of learning)
• Today I would like to talk about translation. Some people might refer to it as protein synthesis … [long pause] … take your time and start thinking about this process and its components and the sequence of events. Take as much time as you want, don't rush, just relax and think about it for a while [long pause] … think about everything you know about this process, what parts are involved [long pause] … slowly, let your thoughts flow … [silence]. When you are ready, go ahead and tell me what you know about translation … if you want, you can draw things too [paper and pencil will be provided to all interviewees] … [after a while] … ok, what are you thinking about now …• If you were going to explain this process to another student, how would you go about doing so? What things would you emphasize to them so they could understand the process?
• What do you see as being the main idea? What's the “big thing” students should come away with after learning about translation?
• What do you think is the most difficult thing about learning or understanding translation?
Phases 2–6 – students' experiences of external representations of translation (post lived object of learning)
• Go ahead and take a look at the first external representation … tell me what you're seeing.
• What can you tell me about what is happening in this external representation?
• Tell about all of the different parts you see. What are they? What do they do? How do they interact? What are the most important parts?
• What do you think this external representation is not showing? Explain your answer.
• Do you think this is a good and clear representation? Give reasons for your answer.
• Consider yourself a graphic designer or textbook author. If you could change this external representation in any way, what would you do to improve it, if anything?
• Here's the second external representation … tell me what you're seeing.
• What can you tell me about what is happening in this external representation?
• Tell about all of the different parts you see. What are they? What do they do? How do they interact? What are the most important parts?
• What do you think this external representation is not showing? Explain your answer.
• Do you think this is a good and clear representation? Give reasons for your answer.
• Consider yourself a graphic designer or textbook author. If you could change this external representation in any way, what would you do to improve it, if anything?
• Do you think both representations contained the same amount of information? Why do you say that?
• Which external representation do think was easier to understand? What made # _____ easier?
• If you were going to compare them, which representation do you think was the best? What about # _____ makes it better than # _____?
• If you were going to explain translation to another student, would you choose to show them one of these external representations?
– If yes, which one? Why # _____? What would the student have to know before you showed them the representation? What things would you point out to them better understand the representation?
– If no, why not? Would you show them something different? How would you get them to understand translation?
• Do you find it helpful or not to use external representations when you learn about things like translation? Why do you say that?
• Earlier you said that translation was ________________ (repeat what they said in Phase 1, show any drawings they made). Do you still agree? Is there anything you would add or change?
• Do you have any final thoughts about external representations or translation? Thank you so much for your time today.
Appendix C: categorization of critical features of protein translation
| Theme | General feature | Coded feature | Type of feature |
|---|---|---|---|
| Note: classification of the type of feature refers to the extent to which instructors agreed that a particular feature was important for students to learn. Primary critical features (1°) were described by all five instructors interviewed. Secondary critical features (2°) were described by a majority (3–4) of instructors but not all. Tertiary critical features (3°) were described by a minority (1–2) of instructors. | |||
| Component/structure | mRNA | General molecule M | 1° |
| Codon | 1° | ||
| Nucleotide sequence (multiple codons) | 2° | ||
| Shine–Dalgarno sequence | 2° | ||
| 5′ end M | 2° | ||
| 3′ end M | 2° | ||
| 3′ poly A tail | 2° | ||
| 5′ methylated cap | 3° | ||
| Start codon | 3° | ||
| Stop codon | 3° | ||
| Nucleotide sequence (start codon) | 3° | ||
| Ribosome | General molecule R | 1° | |
| Large subunit | 1° | ||
| Small subunit | 1° | ||
| Aminoacyl (A) site | 1° | ||
| Peptidyl (P) site | 1° | ||
| Exit (E) site | 1° | ||
| Tunnel | 2° | ||
| 16S rRNA | 3° | ||
| tRNA | General molecule(s) T | 1° | |
| Anti-codon loop | 1° | ||
| P-site tRNA | 1° | ||
| A-site tRNA | 2° | ||
| 3′ end T | 2° | ||
| E-site tRNA | 3° | ||
| 3D shape | 3° | ||
| 2D shape | 3° | ||
| 5′ end T | 3° | ||
| Nucleotide sequence (anti-codon loop) | 3° | ||
| Amino acids | General molecule AA | 1° | |
| Methionine | 2° | ||
| Sequential AA | 3° | ||
| Polypeptide chain | General molecule P | 1° | |
| Primary structure | 2° | ||
| Secondary structure | 3° | ||
| Initiation factors | General molecule(s) IF | 3° | |
| Elongation factors | General molecule(s) EF | 2° | |
| EF-Tu | 3° | ||
| EF-Ts | 3° | ||
| EF-G | 3° | ||
| Release factors | General molecule(s) RF | 2° | |
| Interactions/chemistry | Activation | General process A | 2° |
| Regeneration of activated tRNAs | 3° | ||
| Initiation | Codon/anti-codon base pairing I | 1° | |
| General process I | 2° | ||
| Hydrogen bonding (codon/anti-codon) I | 3° | ||
| Elongation | Peptide bond formation | 1° | |
| Codon/anti-codon base pairing E | 1° | ||
| General process E | 2° | ||
| Incoming tRNA/ribosome/mRNA | 2° | ||
| Exiting tRNA/ribosome/mRNA | 2° | ||
| Ribosomal translocation | 2° | ||
| GTPase activity of EFs | 3° | ||
| Hydrogen bonding (codon/anti-codon) E | 3° | ||
| General considerations | Reaction kinetics | 2° | |
| Evolution | 2° | ||
| Regulation | 2° | ||
| Random motion of components | 2° | ||
| Energetics | 3° | ||
Appendix D: definition of codes
16S rRNA: a description or depiction of the ribosomal ribonucleic acid component of the small ribosomal subunit of prokaryotes.2D shape: a description or depiction of the two dimensional cloverleaf shape of a transfer ribonucleic acid.
3′ end M: a description or depiction of the 3′ hydroxyl end of a messenger ribonucleic acid.
3′ end T: a description or depiction of the 3′ hydroxyl end of a deacylated transfer ribonucleic acid.
3′ poly A tail: a description or depiction of the polyadenylated 3′ tail of a mature eukaryotic messenger ribonucleic acid.
3D shape: a description or depiction of the three dimensional L-shape of a transfer ribonucleic acid.
5′ end M: a description or depiction of the 5′ phosphate end of a messenger ribonucleic acid.
5′ end T: a description or depiction of the 5′ phosphate end of a transfer ribonucleic acid.
5′ methylated cap: a description or depiction of the 7-methylguanosine cap at the 5′ end of a mature eukaryotic messenger ribonucleic acid.
A-site tRNA: a description or depiction of a transfer ribonucleic acid occupying the aminoacyl (A) site of the ribosome.
AA/AA interaction: a description or depiction of two or more amino acids interacting with one another. A peptide bond does not need to be indicated.
Aminoacyl (A) site: a description or depiction of the animoacyl (A) site of the ribosome.
Anti-codon loop: a description or depiction of the anti-codon loop of a transfer ribonucleic acid. The nucleotide base sequence does not need to be indicated.
Codon: a description or depiction of a codon on the messenger ribonucleic acid. The nucleotide base sequence does not need to be indicated.
Codon/anti-codon base pairing E: a description or depiction of the interaction between start codon of the messenger ribonucleic acid and the complementary anti-codon on the anti-codon loop of the initiator transfer ribonucleic acid. This interaction occurs only during the initiation phase of protein translation. Hydrogen bonding does not need to be indicated.
Codon/anti-codon base pairing I: a description or depiction of the interaction between a codon of the messenger ribonucleic acid and a complementary anti-codon on the anti-codon loop of a transfer ribonucleic acid. This interaction occurs only during the elongation phase of protein translation. Hydrogen bonding does not need to be indicated.
E-site tRNA: a description or depiction of a transfer ribonucleic acid occupying the exit (E) site of the ribosome.
EF-G: a description or depiction of the prokaryotic elongation factor G.
EF-Ts: a description or depiction of the prokaryotic elongation factor Ts.
EF-Tu: a description or depiction of the prokaryotic elongation factor Tu.
Energetics: a description or depiction of thermodynamic considerations of the process of protein translation.
Evolution: a description or depiction of the impact of the process of protein translation on evolution.
Exit (E) site: a description or depiction of the Exit (E) site of the ribosome.
Exiting tRNA/ribosome/mRNA: a description or depiction of the relationship between the messenger ribonucleic acid, the ribosome, and the transfer ribonucleic acid that has exited the ribosome.
General molecule M: a description or depiction of the messenger ribonucleic acid as a general entity.
General molecule P: a description or depiction of the polypeptide chain as a general entity.
General molecule(s) AA: a description or depiction of one or several amino acids as a general entity.
General molecule(s) EF: a description or depiction of one or several elongation factors as a general entity.
General molecule(s) IF: a description or depiction of one or several initiation factors as a general entity.
General molecule(s) R: a description or depiction of the ribosome as a general entity.
General molecule(s) RF: a description or depiction of one or several release factors as a general entity. The term termination factor(s) will be included in the code.
General molecule(s) T: a description or depiction of one or several transfer ribonucleic acids as a general entity.
General process A: a description or depiction of the process of activation of transfer ribonucleic acids as a general entity. This code also includes charging of transfer ribonucleic acids.
General process E: a description or depiction of the process of elongation as a general entity. This code includes indications of polypeptide chain growth.
General process I: a description or depiction of the process of initiation as a general entity.
General process T: a description or depiction of the process of termination as a general entity.
GTPase activity of EFs: a description or depiction of the process of hydrolysis of guanosine triphosphate to guanosine diphosphate and phosphate as carried out by an elongation factor as a general entity.
Hydrogen bonding (codon/anticodon) E: a description or depiction of the hydrogen bonds formed between the codon of a messenger ribonucleic acid and the anti-codon of a complementary transfer ribonucleic acid during elongation.
Hydrogen bonding (codon/anticodon) I: a description or depiction of the hydrogen bonds formed between the start codon of the messenger ribonucleic acid and the anti-codon of the initiator transfer ribonucleic acid during initiation.
Hydrogen bonding (mRNA/ribosome) I: a description or depiction of the hydrogen bonds formed between the Shine–Dalgarno sequence of prokaryotic messenger ribonucleic acid and the 16S ribosomal ribonucleic acid of the small ribosomal subunit during initiation.
Incoming tRNA/ribosome/mRNA: a description or depiction of the relationship between the messenger ribonucleic acid, the ribosome, and the transfer ribonucleic acid that is about to enter the aminoacyl (A) site the ribosome.
Initial tRNA/ribosome/mRNA: a description or depiction of the relationship between the messenger ribonucleic acid, the ribosome, and the initiator transfer ribonucleic acid during the formation of the initiation complex.
Large subunit: a description or depiction of the large ribosomal subunit.
Methionine: a description or depiction of the amino acid, methionine, of the initiator transfer ribonucleic acid in eukaryotes. This code also includes formylmethionine on the initiator transfer ribonucleic acid in prokaryotes.
Nucleotide sequence (anti-codon loop): a description or depiction of the nucleotide sequence of the anti-codon loop of a transfer ribonucleic acid.
Nucleotide sequence (tRNA): a description or depiction of the nucleotide sequence of a transfer ribonucleic acid. This code is inclusive of the anti-codon loop and refers to the entire sequence of the transfer ribonucleic acid.
Nucleotide sequence (multiple codons): a description or depiction of the nucleotide sequence of multiple codons of the messenger ribonucleic acid. This code is inclusive of the start and stop codons and refers to a series of codon on the messenger ribonucleic acid.
Nucleotide sequence (start codon): a description or depiction of the nucleotide sequence of the start codon of the messenger ribonucleic acid (AUG).
Nucleotide sequence (stop codon): a description or depiction of the nucleotide sequence of the stop codon of the messenger ribonucleic acid.
P-site tRNA: a description or depiction of a transfer ribonucleic acid occupying the peptidyl (P) site of the ribosome.
Peptide bond formation: a description or depiction of the nucleophilic attack by the lone pair electrons on the amine group of the amino acid of the A-site tRNA on the carbonyl carbon of the amino acid of the P-site tRNA resulting in the formation of a peptide bond. This is a mechanistic description or depiction.
Peptidyl (P) site: a description or depiction of the peptidyl (P) site of the ribosome.
Primary structure: a description or depiction of the primary sequence of amino acids of the polypeptide chain. The identity of the amino acids does not need to be indicated.
Random motion of cellular components: a description or depiction of the random motion of cellular components in the cytoplasm. Indications of how those random motions can lead to component collisions and subsequent chemical reaction may also be included.
Reaction kinetics: a description or depiction of the rate or speed of the process of protein translation.
Regeneration of activated tRNAs: a description or depiction of the process of regenerating transfer ribonucleic acids with an appropriate amino acid that corresponds to the anti-codon sequence.
Regulation: a description or depiction of various aspects of cellular regulation on the process of protein translation.
Ribosomal translocation: a description or depiction of the movement of the ribosome from one codon to the next from the start codon to the stop codon of the messenger ribonucleic acid.
Secondary structure: a description or depiction of the folding of the polypeptide chain as it interacts with the cytoplasm.
Sequential AA: a description or depiction of the amino acid attached to the incoming transfer ribonucleic acid.
Shine–Dalgarno sequence: a description or depiction of the Shine–Dalgarno region of the prokaryotic messenger ribonucleic acid located upstream of the start codon.
Small subunit: a description or depiction of the small ribosomal subunit.
Start codon: a description or depiction of the start codon of the messenger ribonucleic acid. The nucleotide sequence does not need to be indicated.
Stop codon: a description or depiction of the stop codon of the messenger ribonucleic acid. The nucleotide sequence does not need to be indicated.
Tunnel: a description or depiction of the channel through the large ribosomal subunit from the P-site to the cytoplasm though which the growing polypeptide chain exits the ribosome.
References
- Avgerinou M. and Ericson J., (1997), A review of the concept of visual literacy, Brit. J. Educ. Technol., 28(4), 280–291.
- Bauer M. I. and Johnson-Laird P. N., (1993), How diagrams can improve reasoning, Psychol. Sci., 4(6), 372–378.
- Bell E., (2001), The future of education in the molecular life sciences, Nat. Rev. Mol. Cell Biol., 2, 221–225.
- Bussey T., (2013), What can biochemistry students learn about protein translation? Using variation theory to explore the space of learning created by some common external representations, unpublished Doctoral Dissertation, University of Nevada, Las Vegas, Las Vegas, NV.
- Bussey T. J., Orgill M. and Crippen K. J., (2013), Variation theory: a theory of learning and a useful theoretical framework for chemical education research, Chem. Educ. Res. Pract., 14, 9–22.
- Cloud-Hansen K. A., Kuehner J. N., Tong L., Miller S. and Handelsman J., (2008), Money, sex, and drugs: a case student to teach the genetics of antibiotic resistance, CBE-Life Sci. Educ., 7, 302–309.
- Cook M. P., (2006), Visual representations in science education: the influence of prior knowledge and cognitive load theory on instructional design principles, Sci. Educ., 90, 1073–1091.
- Cook M. P., Wiebe E. N. and Carter G., (2008), The influence of prior knowledge on viewing and interpreting graphics with macroscopic and molecular representations, Sci. Educ., 92, 848–867.
- Fisher K. M., (1985), A misconception in biology: amino acids and translation, J. Res. Sci. Teach., 22(1), 53–62.
- Gombrich E. H., (1980), Standards of truth: the arrested image and the moving eye, in Mitchell W. J. T. (ed.), The language of images, Chicago, IL: The University of Chicago Press, pp. 181–218.
- Gray L., Thomas N., Lewis L. and Tice P., (2010), Teachers' use of educational technology in U.S. public schools: 2009, Washington, DC: National Center for Education Statistics, Institute of Education Sciences, U.S. Department of Education.
- Harle M. and Towns M. H., (2012a), Students' understanding of external representations of the potassium ion channel protein, part I: affordances and limitations of ribbon diagrams, vines, and hydrophobic/polar representations, Biochem. Mol. Biol. Educ., 40, 349–356.
- Harle M. and Towns M. H., (2012b), Students' understanding of external representations of the potassium ion channel protein, part II: structure–function relationships and fragmented knowledge, Biochem. Mol. Biol. Educ., 40, 357–363.
- Harle M. and Towns M. H., (2013), Students' understanding of primary and secondary protein structure: drawing secondary protein structure reveals student understanding better than simple recognition of structures, Biochem. Mol. Biol. Educ., 41, 396–376.
- Harrison A. G. and Treagust D. F., (1996), Secondary students' mental models of atoms and molecules: implications for teaching chemistry, Sci. Educ., 80(5), 509–534.
- Harrison A. G. and Treagust D. F., (2000), Learning about atoms, molecules, and chemical bonds: a case study of multiple-model use in grade 11 chemistry, Sci. Educ., 84(3), 352–381.
- Heyworth R. M., (1999), Procedural and conceptual knowledge of experts and novice students for the solving of a basic problem in chemistry, Int. J. Sci. Educ., 21(2), 195–211.
- Klymkowsky M. W., (2007), Teaching without a textbook: strategies to focus learning on fundamental concepts and scientific processes, CBE-Life Sci. Educ., 6(3), 190–193.
- Linenberger K. J. and Bretz S. L., (2012), A novel technology to investigate students' understanding of enzyme representations, J. Coll. Sci. Teach., 42(1), 45–49.
- Linenberger K. J. and Holme T. A., (2014), Results of a national survey of biochemistry instructors to determine the prevalence and types of representations used during instruction and assessment, J. Chem. Educ., 91, 800–806.
- Linenberger K. J. and Holme T. A., (2015), Biochemistry instructors' views toward developing and assessing visual literacy in their courses, J. Chem. Educ., 92, 23–31.
- Lowe R., (1996), Background knowledge and the construction of a situational representation from a diagram, Eur. J. Psychol. Educ., 11(4), 377–397.
- Lowe R., (2003), Animation and learning: Selective processing of information in dynamic graphics, Learn. Instr., 13, 157–176.
- Lowe R. (2004). Interrogation of a dynamic visualization during learning, Learn. Instr., 14, 257–274.
- Marton F., Beaty E. and Dall'Alba G., (1993), Conceptions of learning, Int. J. Educ. Res., 19, 277–300.
- Miskowski J. A., Howard D. R., Abler M. L. and Grunwald S. K., (2007), Design and implementation of an interdepartmental bioinformatics program across life science curricula, Biochem. Mol. Biol. Educ., 35(1), 9–15.
- Nelson M. M. and Cox D. L., (2000), Lehninger Principles of Biochemistry, 3rd edn, New York, NY: W. H. Freeman.
- Olsen E. (Producer), (2010, November 15), The animators of life, The New York Times, video retrieved November 17, 2010, from http://video.nytimes.com/video/2010/11/15/science/1248069334032/the-animators-of-life.html?scp=1&sq=animations%<?pdb_no 20of?>20of<?pdb END?>%20life&st=cse.
- Perino G. H., (2001), Reductive reasoning: a cognitive barrier to visual literacy, Journal of Visual Literacy, 21(1), 15–30.
- Pinar W. F., Reynolds W. M., Slattery P. and Taubman P. M., (2004), Understanding curriculum: an introduction to the study of historical and contemporary curriculum Discourses, New York, NY: Peter Lang Publishing, Inc.
- Rotbain Y., Marbach-Ad G. and Stavy R., (2006), Effect of bead and illustrations models on high school students' achievement in molecular genetics, J. Res. Sci. Teach., 43(5), 500–529.
- Schönborn K. J. and Anderson T. R., (2006), The importance of visual literacy in the education of biochemists, Biochem. Mol. Biol. Educ., 34(2), 94–102.
- Schönborn K. J. and Anderson T. R., (2009), A model of factors determining students ability to determine external representations in biochemistry, Int. J. Sci. Educ., 31(2), 193–232.
- Schönborn K. J. and Anderson T. R., (2010), Bridging the educational research-teaching practice gap: foundations for assessing and developing biochemistry students' visual literacy, Biochem. Mol. Biol. Educ., 38(5), 347–354.
- Schönborn K. J., Anderson T. R. and Grayson D. J., (2002), Student difficulties with the interpretation of a textbook diagram of immunoglobulin G (IgG), Biochem. Mol. Biol. Educ., 30(2), 93–97.
- Schönborn K. J., Anderson T. R. and Mnguni L. E., (2007), Methods to determine the role of external representations in developing understanding in biochemistry, in Lemmermöhle D. et al. (ed.), Professionell lehren—erfolgreich lernen, New York: Waxmann, pp. 291–301.
- Sim J. H. and Daniel E. G. S., (2014), Representational competence in chemistry: a comparison between students with different levels of understanding of basic chemical concepts and chemical representations, Cogent Educ., 1, 1–17.
- Snyder J. L., (2000), An investigation of the knowledge structures of experts, intermediates and novices in physics, Int. J. Sci. Educ., 22(9), 979–992.
- Winn W., (1991), Learning from maps and diagrams, Educ. Psychol. Rev., 3(3), 211–247.
| This journal is © The Royal Society of Chemistry 2015 |