Reaching beyond HIV/HCV: nelfinavir as a potential starting point for broad-spectrum protease inhibitors against dengue and chikungunya virus†
Abstract
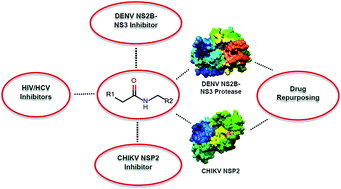
Drug repurposing or re-profiling has become an effective strategy to identify novel indications for already-approved drugs. In this study, peptidomimetic FDA-approved HIV/HCV inhibitors were explored for their potential to be repurposed for the inhibition of the replication of dengue (DENV) and chikungunya virus (CHIKV) by targeting the NS2B-NS3 and NSP2 protease, respectively. MM/GBSA-based binding free energy results put nelfinavir forward as a potential inhibitor of both dengue and chikungunya virus, which subsequently was further explored in a virus-cell-based assay for both viruses. Nelfinavir showed modest antiviral activity against CHIKV (EC50 = 14 ± 1 μM and a selectivity index of 1.6) and was slightly more active against DENV-2 (EC50 = 3.5 ± 0.4 μM and a selectivity index of 4.6). Even though the antiviral potency was limited, the fact that some activity was observed in these assays made it worthwhile exploring the potential and properties of nelfinavir as a stepping-stone compound: a more detailed computational analysis was performed to understand the binding mode, interaction, hydrogen bond distance, occupancy and minimum pharmacophoric features. The comprehensive data set that resulted from these analyses may prove to be useful for the development of novel DENV and CHIKV protease inhibitors.

 Please wait while we load your content...
Please wait while we load your content...