Superparamagnetic core/shell nanostructures for magnetic isolation and enrichment of DNA
R. M. Patila,
P. B. Shetea,
S. M. Patilb,
S. P. Govindwarc and
S. H. Pawar*a
aCenter for Interdisciplinary Research, D. Y. Patil University, Kolhapur-416006, MS, India. E-mail: pawar_s_h@yahoo.com; Fax: +91-231-260159; Tel: +91-231-2601202
bDepartment of Biotechnology, Shivaji University, Kolhapur-416004, MS, India
cDepartment of Biochemistry, Shivaji University, Kolhapur-416004, MS, India
First published on 12th October 2015
Abstract
Fe3O4 magnetic nanoparticles (MNPs) are promising candidates for various biomedical applications due to their extraordinary properties. Such MNPs, surface modified with chitosan–glutaraldehyde (Fe3O4–CH/GLD) i.e. magnetic core/shell nanostructures, were used in the present study to investigate isolation and enrichment of bacterial DNA. Isolation was carried out in comparison with an organic method. FTIR was used to confirm biding of DNA onto the surface of the core–shell structures. The concentration of isolated DNA (yield) was 14.90 and 17.55 μg mL−1 for phenol/chloroform and magnetic isolation methods, respectively. The purity of isolated DNA was found to be 1.69 and 1.71 for phenol/chloroform and magnetic isolation methods, respectively. The present study firstly reports the comparison between magnetic and organic isolation of DNA. From both results (yield and purity), it was found that magnetic isolation of DNA was superior to the general organic method used for bacterial DNA isolation. Experiments for DNA enrichment were performed in batch mode and the effects of core/shell concentration, pH of the sample solution and temperature were optimized. The formation energy required for adsorption of DNA was found to be −55.56 × 10−23 J per molecule (−34.70 × 10−4 eV per molecule). The negative value indicates energy was utilized (endothermic process) for the adsorption of DNA onto the magnetic core/shells. The magnetic isolation method used in the present study was simple, fast, robust and ecofriendly (it does not require organic solvents or sophisticated equipment).
1. Introduction
Currently there is huge interest in the design of nanobiocomposites with MNPs and nucleic acids. Their application is of vast importance for growth in the field of nanoelectronics, biomedical diagnosis and therapy.1 The use of DNA molecules in designing nanodevices is very interesting. The rapid advancement of ideas on the utilization of MNPs in biomedical research began in the 1960s.2 The unique properties of magnetic materials in the nanostate provide the possibility of detecting structures based on MNPs and to control them by an external magnetic field. Conjugation of MNPs with biological molecules, nucleic acids in particular, allows development of various nanobiohybrid systems that possess unique magnetic properties and biological selectivity to improve the efficiency of diagnosis and therapy of various diseases.3Various approaches to conjugate nucleic acids with MNPs have been proposed. A nucleic acid molecule can directly bind to MNPs; otherwise, formation of chemical bonds requires surface modification of MNPs. However, prevention of nonspecific interactions between nucleic acids and MNPs and determination of component ratios in complexes still remains topical for nanobiofabrication.4 A research on mechanisms of the DNA–MNP interaction is of paramount importance for assessment of the influence of MNPs on genetic material in biological systems, which is directly related to human safety.5
The affinity of ferrite MNPs towards DNA is well reported. Interaction between DNA and any other substrate is given by the forces existing between them. Frey et al.6 investigated the physics of DNA using single molecule manipulation technique. He used superparamagnetic beads as force probe for DNA molecule which is attached to AFM cantilever. The force exerted on DNA molecule is determined by Hooke's law,
| F = −K′z | (1) |
Adsorption is a multistep process involving transport of solute particles from solution to surface of the solid particles followed by diffusion into the interior of the pores. Lagergren's pseudo-first-order7 and Ho's pseudo-second-order8 kinetic models were used in order to study the controlling mechanisms of adsorption process. Lagergren's pseudo-first-order differential equation is:
| qt = qe(1 − e−k1t) | (2) |
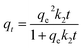 | (3) |
Liu et al. studied the isotherm parameters for plasmid DNA adsorption using acridine orange MNPs (ACO-MNPs).9 The analysis of the isotherm data also helps in finding how the adsorbate molecules distribute between the liquid phase and the solid phase when the adsorption process reaches an equilibrium state. This can be done by fitting them to different isotherm models. The adsorption of DNA onto both MNPs is described well by the Langmuir model than Freundlich model. Taylor et al.10 investigated binding of DNA to MNPs and showed that regularities of adsorption/desorption on MNPs differ from those established for classical adsorbents (e.g., silica).
Covalent immobilization is often attained with thiolated and aminated molecules. Here, amino, carboxyl, sulfhydryl and azido groups are formed on the surface of MNPs. Covalent bonds are formed using traditional methods of bioconjugation and ‘click’-chemistry approaches including thiol-disulfide exchange, carbodiimide activation, azide–alkyne cycloaddition, aldehyde–amine condensation, etc. Bioconjugation of DNA with MNPs is very often attained using cross-linkers.11
The present manuscript focuses on magnetic isolation and enrichment of bacterial DNA. E. coli DNA is magnetically isolated and the results are compared with routinely used DNA isolation i.e. phenol/choloroform method. The DNA is also studied for its adsorption on magnetic core–shells.
2. Materials
Ferrous chloride (FeCl2·4H2O), hydrochloric acid (HCl), glacial acetic acid, sodium hydroxide (NaOH) were procured from HiMedia, India. Chitosan and glutaraldehyde were purchased from Sigma-Aldrich, USA. Double distilled water was used throughout the procedure.3. Experimental
Fe3O4–CH/GLD core/shells were synthesized as reported earlier.12 These core/shells were studied for isolation and enrichment of bacterial DNA. E. coli BL21 was chosen for the experiments as it is well known and easily available microorganism. E. coli culture was maintained on MacConkey's agar. MacConkey agar is a selective and differential culture medium designed to selectively isolate Gram negative bacteria and enteric bacilli and differentiate them based on lactose fermentation as per the protocol.13,14 It contains bile salts (to inhibit most Gram positive bacteria), crystal violet dye (which also inhibits certain Gram positive bacteria), neutral red dye (which turns pink if the microbes are fermenting lactose).Isolation of bacterial DNA was done using phenol/chloroform method already reported with slight modification to purify genomic DNA. The procedure followed for magnetic isolation is described in brief as follows. The cells were lysed using lysis buffer by incubation at 37 °C for 1 h. After cell lysis, 10 mg of magnetic core/shells were mixed in sample so that DNA molecules can attach to magnetic core–shells. This suspension was kept at room temperature for 10 min on thermoshaker. The MNPs were then separated out using bar magnets. The adsorbed DNA was eluted by 1 mL of Tris buffer (pH 7.5 and 8.5).
Gel electrophoresis of isolated DNA samples was carried out for separation and visualization under UV light. The protocol can be divided into three stages: (1) a gel is prepared with an agarose concentration appropriate for the size of DNA fragments to be separated; (2) the DNA samples are loaded into the sample wells and the gel is run at a voltage and for a time period that will achieve optimal separation; and (3) the gel is stained or, if ethidium bromide has been incorporated into the gel and electrophoresis buffer, visualized directly upon illumination with UV light.15
The concentration and purity of DNA sample were checked by the use of UV spectrophotometry. DNA absorbs UV light very efficiently making it possible to detect and quantify either at concentrations as low as 2.5 ng μL−1. The nitrogenous bases in nucleotides have an absorption maximum at about 260 nm. These procedures were carried out on Eppendorf Biospectrometer.
The adsorption of DNA on Fe3O4–CH/GLD core/shells was investigated in order to study exact nature of DNA binding with core/shells and influence of various parameters (such as pH, temperature and concentration of core/shells) on enrichment of DNA. Firstly, the amount of adsorbent dosage i.e. magnetic core/shells (2, 4, 6, 8 and 10 mg) were examined. The standard DNA concentration was kept constant at 15 μg mL−1. The experiments were carried out at room temperature with pH 4. The effect of pH on adsorption was investigated at pH range of 3–8, keeping 6 mg amount of magnetic core/shells at room temperature. The initial pH of the solution was adjusted by using 0.1 M HCl or 0.1 M NaOH. The standard DNA concentration was kept constant at 15 μg mL−1. Again, temperature range of 10 to 50 °C was studied at pH 4 and 6 mg amount of magnetic core–shells. The elution of adsorbed DNA from 1 mL aqueous solution was carried out in 100 μL of Tris base of pH 8.5.
4. Results and discussion
4.1. Isolation and detection of DNA
E. coli DNA was successfully isolated by phenol/chloroform method and by magnetic isolation. The protocols used for DNA isolation by phenol/chloroform method and magnetic isolation method are shown in Fig. 1. As bare MNPs were coated with CH/GLD, their surface has amino groups. These free amino groups form bonds with phosphate groups of DNA molecules. There is also a possibility of adsorption of DNA on surface of MNPs. The isolated DNA by both the methods was used for further analysis i.e. for diphenylamine (DPA) test, agarose gel electrophoresis, UV-Vis spectroscopy and FTIR. The morphology of core/shells is given in Fig. 2.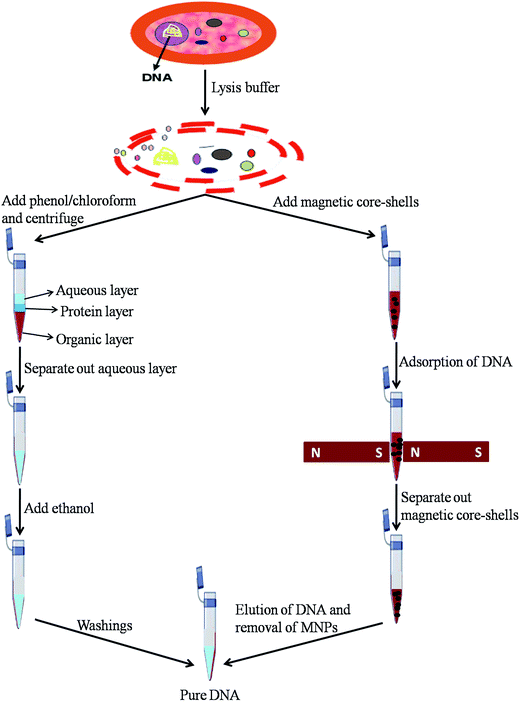 | ||
| Fig. 1 Diagrammatic representation of protocol used for DNA isolation by phenol/chloroform method and magnetic isolation method. | ||
DPA is a qualitative test for detection of DNA. When DNA is treated with DPA under the acidic condition a bluish green colored complex is formed which has an absorption peak at 595 nm. This reaction is given by 2-deoxypentose in general. In acidic solution deoxypentose are converted into a highly reactive β hydroxyl levulinic aldehyde, which reacts with DPA and gives bluish green colored complex. The color intensity was measured using a red filter at 595 nm. After magnetic isolation of DNA, DPA test was performed. The color change from colorless to blue confirms the presence of DNA in the sample.
An absorption maximum (λmax) of the product of DPA reagent and DNA reaction was determined. The optical density was taken from 450 to 650 nm after color development. Optical density of the reaction product is plotted against wavelength in nm and shown in Fig. 3. The λmax was found to be 600 nm and it was in well comparison with the principle of reaction and the literature.16
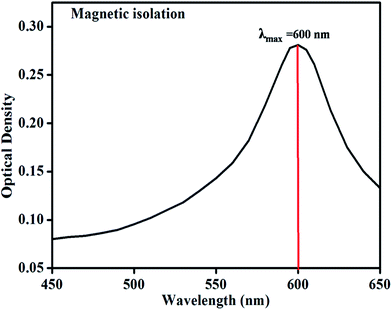 | ||
| Fig. 3 Visible spectra of product of reaction between DNA and DPA reagent for magnetic isolation method. | ||
Adsorption of DNA on core/shells was confirmed by FTIR spectroscopy. Fig. 4 shows the FTIR spectra of Fe3O4–CH/GLD core shells and DNA adsorbed core/shells (DNA@MNPs) over the range of 400 to 4000 cm−1. As stated earlier in chapter 5, the band observed at 565 cm−1 corresponds to the intrinsic stretching vibration (Fetetra–O) of metal–oxygen at tetrahedral site, whereas the band observed at 455 cm−1 corresponds to the stretching vibration (Feocta–O) of metal–oxygen at octahedral site. The band observed at 3376 cm−1 corresponds to surface-adsorbed water molecules on Fe3O4. The bands observed at 2854 and 2924 cm−1 relate to C–H stretching vibrations. The broad band observed at around 3391 cm−1 corresponds to stretching vibration of N–H and O–H. The band observed at 1624 cm−1 in Fe3O4–CH/GLD and DNA@MNPs was corresponds to imine (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N), which was due to the cross linking of CH with GLD.12 This confirmed that core–shell structures were not disturbed by the adsorption of DNA.
N), which was due to the cross linking of CH with GLD.12 This confirmed that core–shell structures were not disturbed by the adsorption of DNA.
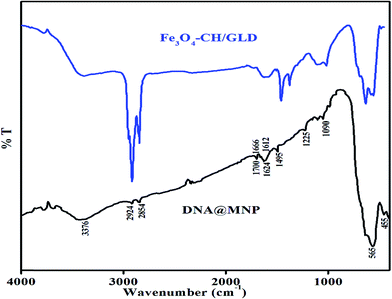 | ||
| Fig. 4 FTIR spectra of Fe3O4–CH/GLD core/ shells and DNA adsorbed core/shells (DNA@MNPs) over the range of 400 to 4000 cm−1. | ||
The FTIR spectrum of DNA in the region of 400 to 1800 cm−1 contains a variety of information on the conformational arrangement. Polymers with amide or amine groups can interact with DNA via electrostatic attractions after their protonation. However, the N–H stretching vibration of nucleic acid bases in the region 1500 to 1700 cm−1 overlaps with the amine signals of the polymers. Hence, the bands for C–H and N–H bending were not clearly observed in DNA@MNPs spectra. The bands observed at 1700, 1612, 1666 and 1495 cm−1 were corresponds to guanine, adenine, thymine and cytosine respectively. The slight shifts in these bands may be attributed to the interaction of DNA with the core–shell of MNPs. The slight shifts in bands were in agreement with the literature.17 The bands observed at 1090 and 1225 cm−1 are attributed to the symmetric and asymmetric PO2− stretching vibrations. The marker band for PO2− is normally observed at 1236 cm−1,18,19 but in this case it was observed at 1225 cm−1. This shift was may be due to the interaction of negatively charged phosphate with the positively charged amino groups in CH. The carbonyl stretching vibration of deoxyribose sugar of the DNA appeared as a strong band at 1064 cm−1.18 In this case it was shifted to 1051 cm−1. This shift may be due to the interaction between the CH/GLD and deoxyribose sugar of the DNA backbone, which results in hydrogen bonding between the sugar and polymer. The shifting in major bands of nucleotides, phosphate and deoxyribose sugar showed the DNA had electrostatic interaction and hydrogen bonding with the Fe3O4–CH/GLD core–shells.
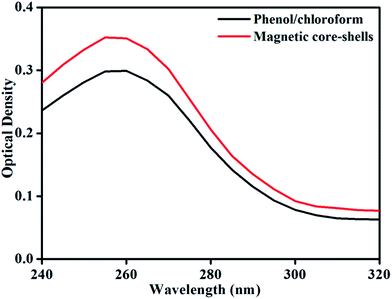
To determine the concentration (yield) and purity of isolated DNA by both the methods, UV-Vis spectroscopy of the samples was done. Fig. 5 shows UV spectra of the extracted DNA samples by phenol/chloroform and magnetic isolation method. Concentration of the DNA in the samples was calculated by measuring the absorption at 260 nm and using the following equation
| DNA concentration (μg mL−1) = (OD260) × (dilution factor) × (50 μg DNA per mL)/(1OD260 unit) | (4) |
Using a 1 cm light path, the extinction coefficient for nucleotides at wavelength 260 nm is 20. Based on this extinction coefficient, absorbance at 260 nm in a 1 cm quartz cuvette of a 50 μg mL−1 solution of double stranded DNA is equal to 1. The absorbance of a DNA sample at 280 nm gives an estimate of the protein contamination of the sample. The ratio of the absorbance at 260 nm/absorbance at 280 nm is a measure of the purity of a DNA sample; it should be between 1.65 and 1.85.
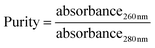 | (5) |
The concentration of isolated DNA (yield) was 14.90 and 17.55 μg mL−1 for phenol/chloroform and magnetic isolation method respectively. Purity of isolated DNA was found to be 1.69 and 1.71 for phenol/chloroform and magnetic isolation method respectively. From both the results (yield and purity) it was found that magnetic isolation of DNA is superior over the general organic method used for bacterial DNA isolation. Magnetic isolation is also fast and simple to perform.
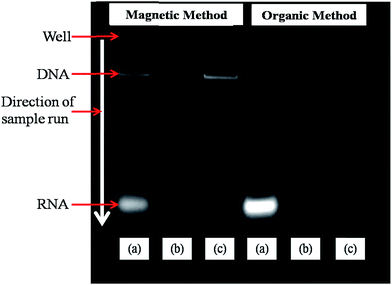
Fig. 6 shows the gel electrophoresis pattern obtained from DNA isolated with phenol/chloroform method and magnetic isolation method. Fig. 6(a) shows patterns for DNA sample without treatment of RNase, where a bright band of RNA can easily be seen at the end of gel, in addition to the band for DNA at the middle of gel. The sample was eluted at pH 8.5. Fig. 6(b) shows pattern for DNA sample with treatment of RNase, where RNA band is vanished as RNase breaks down all RNA in the sample. The sample used for this was eluted at pH 7.5. Fig. 6(c) shows pattern for DNA sample with treatment of RNase, where also RNA band is vanished. The sample used for this was eluted at pH 8.5. The results show that brighter bands are obtained when DNA were magnetically isolated implying higher yield of DNA as compared to samples obtained from general procedure.
4.2. Magnetic enrichment
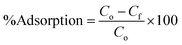 | (6) |
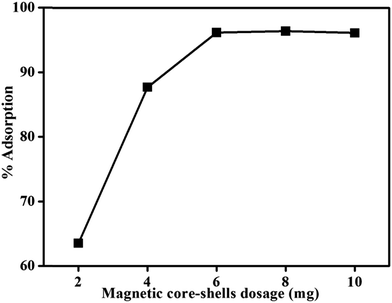
From Fig. 7, it can be seen that adsorption of DNA increases as amount of magnetic core/shells increases upto 6 mg; after which it remains constant. As the sample contains fixed number of DNA concentration, further increase in core/shells concentration does not affect the percentage adsorption anymore. It shows that 6 mg of magnetic core/shells were sufficient to adsorb the entire DNA present in the sample.
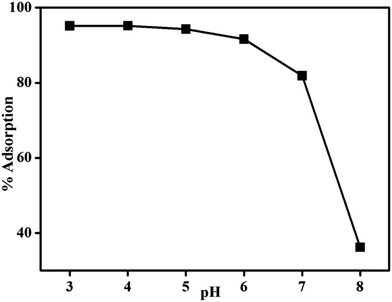
The effect of pH on adsorption of DNA onto magnetic core/shell surfaces was assessed at different pH values, ranging from 3 to 8. The magnetic core/shells amount and temperature were set at 6 mg and 30 °C respectively. The experiments were performed in a batch technique and each solution was stirred for 5 min on thermoshaker. The percentage adsorption of DNA with pH is in Fig. 8 indicating that there is a general decrease in percent adsorption of DNA as pH increases. The results are in good agreement with the adsorption of DNA on silica, TiO2 and CoFe2O4 at different pH showing that adsorption was increased by decreasing the pH.20–23
It seems that both electrostatic interaction and hydrogen bonding are responsible for adsorption of DNA at different pH values. At pH ≤ 6, the presence of protonated amino groups, i.e. –NH3+ and hydroxyl at the surfaces of MNPs, not only develops favorable hydrogen bonding with DNA24 but also causes electrostatic attraction with polyanion DNA (phosphate groups). By increasing pH, eventually, a sharp diminish of protonated amino and hydroxyl groups on the MNPs surfaces is expected. Form the results it was observed that more than 95% DNA was adsorbed at pH 3 and 4. At pH 7 and 8, % adsorption of DNA was very low due to absence of protonated amino groups. The observed data are in good agreement with similar results reported for the adsorption of DNA on metal oxides surfaces.20
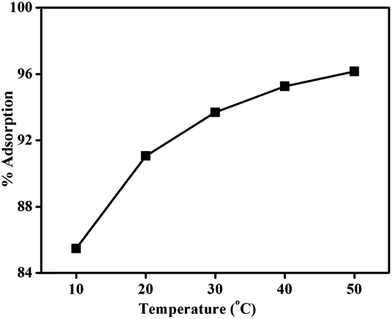
For MNPs, percent adsorption increases with increasing temperature, indicating the endothermic (ΔH > 0) nature of the adsorption process. This nature can be explained theoretically by using the following equation
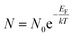 | (7) |
Further, this equation can be modified as,
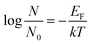 | (8) |
From the above equation, N/N0 indicates adsorption of DNA on magnetic core/shells and is inversely proportional to the temperature, showing that as temperature increases there is an increase in adsorption of DNA on magnetic core–shells. Negative sign indicates that heat is used (i.e. endothermic) in the adsorption process.
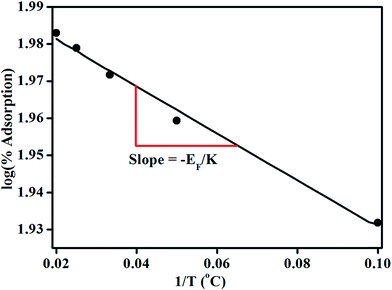
According to equation (8) the log of percentage adsorption of DNA on magnetic core/shells verses 1/T is plotted and shown in Fig. 10. Linear nature of the plot shows that the adsorption process obeys equation (8). From the slope, value of EF is calculated and found to be −55.56 × 10−23 J per molecule (−34.70 × 10−4 eV per molecule). The negative value indicates energy was utilized for the adsorption of DNA onto the magnetic core/shells.
5. Conclusions
The magnetic core/shells were quite efficient as novel magnetic nano-adsorbents for fast adsorption of DNA from aqueous solutions. The electrostatic interactions of magnetic core/shells with phosphate groups of DNA strand may be the basis of adsorption of DNA. By the virtue of this property, DNA could be isolated and enriched magnetically more efficiently than the general methods. A high yield and pure DNA was obtained by magnetic isolation. Effect of various parameters like pH, temperature and amount of magnetic core/shells affected the % adsorption of DNA. The study showed that pH 4, 30 °C temperature and 6 mg concentration of magnetic core/shells were optimum to bind 15 μg of DNA from 1 mL aqueous solution. The present report is very basic study of magnetic core/shells for the application of DNA isolation. This research may open the doors for superparamagnetic core/shell nanostructures to the world of medical diagnosis and forensic sciences. The DNA–magnetic core/shell conjugates may promise important application in medical science, such as forensically challenged samples rapid pathogen detection, gene delivery and magnetofection.References
- K. M. Abu-Salah, A. A. Ansari and S. A. Alrokayan, J. Biomed. Biotechnol., 2010, 7, 15295 Search PubMed.
- L. Merhari, Hybrid Nanocomposites for Nanotechnology: Electronic, Optical, Magnetic and Biomedical Applications, Springer, New York, 2009 Search PubMed.
- B. Haun, T. J. Yoon, H. Lee and R. Weissleder, Wiley Interdiscip. Rev.: Nanomed. Nanobiotechnol., 2010, 2, 291 CrossRef PubMed.
- N. Geerts and E. Eiser, Soft Matter, 2010, 6, 4647 RSC.
- N. Singh, G. Jenkins, R. Asadi and S. H. Doak, Nano Rev., 2010, 1, 5358 Search PubMed.
- E. W. Frey, A. A. Gooding, S. Wijeratne and C. H. Kiang, Frontiers of Physics, 2012, 7(5), 576 CrossRef PubMed.
- S. Lagergren, Zur theorie der sogenannten adsorption gelöster stoffe, Kungliga Svenska Vetenskapsakademiens Handlingar, 1898, vol. 24, pp. 1–39 Search PubMed.
- Y. S. Ho, D. A. J. Wase and C. F. Forster, Environ. Technol., 1996, 17, 71 CrossRef CAS PubMed.
- C. H. Liu, S. L. Sahoo and M. H. Tsao, Colloids Surf., B, 2014, 115, 150 CrossRef CAS PubMed.
- J. I. Taylor, C. D. Hurst, M. J. Davies, N. Sachsinger and I. J. Bruce, J. Chromatogr., 2000, A890159 Search PubMed.
- G. Spoto and R. Corradini, Detection of Non-Amplified Genomic DNA. Dordrecht, Springer, 2012, pp. 67–201 Search PubMed.
- R. M. Patil, P. B. Shete, N. D. Thorat, S. V. Otari, K. C. Barick, A. Prasad, R. S. Ningthoujam, B. M. Tiwale and S. H. Pawar, J. Magn. Magn. Mater., 2014, 355, 22 CrossRef CAS PubMed.
- J. M. Cunningham, et al., Cancer Epidemiol., Biomarkers Prev., 2013, 22, 2047 CrossRef CAS PubMed.
- F. D. Pietro, F. Ortenzi, M. Tilio, F. Concetti and V. Napolioni, Mol. Cell. Probes, 2011, 25, 44 CrossRef PubMed.
- D. Voytas, Current Protocols in Molecular Biology, 2001 DOI:10.1002/0471142727.mb0205as51.
- W. W. Ackermann, F. Sokol and A. J. Brandau, Anal. Biochem., 1965, 12, 332337 CrossRef.
- K. H. Ghanbari, S. Z. Bathaie and M. F. Mousavi, Biosens. Bioelectron., 2008, 23, 1825 CrossRef CAS PubMed.
- M. M. Mady, W. A. Mohammed, N. M. El-Guendy and A. A. Elsayed, Int. J. Phys. Sci., 2011, 6(32), 7328 CAS.
- E. Lipiec, J. Kowalska, J. Lekki, A. Wiecheć and W. M. Kwiatek, Acta Phys. Pol., A, 2012, 121, 506 Search PubMed.
- T. Amano, T. Toyooka and Y. Ibuki, Sci. Total Environ., 2010, 408, 480 CrossRef CAS PubMed.
- A. G. Pershina, A. E. Sazonov and L. M. Ogorodova, Bioorg. Khim., 2009, 35, 674 CAS.
- T. Geng, N. Bao, O. Z. Gall and C. Lu, Chem. Commun., 2009, 7, 800 RSC.
- A. P. Tiwari, R. K. Satvekar, S. S. Rohiwal, V. A. Karande, A. V. Raut, P. G. Patil, P. B. Shete, S. J. Ghosh and S. H. Pawar, RSC Adv., 2015, 5, 8463 RSC.
- N. Takezawa, Bull. Chem. Soc. Jpn., 1971, 44, 3177 CrossRef CAS.
| This journal is © The Royal Society of Chemistry 2015 |