Efficient degradation of sulfamethoxazole and the response of microbial communities in microbial fuel cells†
Lu Wanga,
Yicheng Wub,
Yue Zhenga,
Lidan Liua and
Feng Zhao*a
aKey Laboratory of Urban Pollutant Conversion, Chinese Academy of Sciences Institute of Urban Environment, Chinese Academy of Sciences, Xiamen, 361021, China. E-mail: fzhao@iue.ac.cn; Fax: +86 592 6190766; Tel: +86 592 6190766
bXiamen University of Technology, Xiamen, 361021, China
First published on 12th June 2015
Abstract
Sulfamethoxazole (SMX) is an extensively consumed sulfonamide antimicrobial agent and is frequently detected in surface water. This work studied the degradation process of SMX in anodic chamber of microbial fuel cell (MFC) reactors. We found that the biodegradation of SMX could be achieved after acclimation and even high concentrations of SMX (e.g. 200 ppm) could be rapidly degraded. Excitation and emission matrix fluorescence spectroscopy analysis revealed that the chemical structure of SMX was altered during the process. Q-Exactive hybrid quadrupole-Orbitrap mass spectrometry was used to identify the degradation byproducts of SMX. The activity of electrode biofilm was examined afterwards and it was found that the microbe was in an active state. High-throughput sequencing analysis suggested that the microbial community structure was greatly changed during the process; some reported SMX scavengers, such as Achromobacter and Pseudomonas, were abundant in the reactors. Some metazoans were also recognized in the biofilm samples, which indicates that the operation of the MFC reactors was in a steady state. This study discusses the degradation mechanism of SMX and explores the microbial community response during the process, which provides useful information for the application of MFC in antibiotic elimination.
1 Introduction
Due to the wide application of antibiotics, large amounts of unutilized antibiotics were discharged into the sewage system. Since the biodegradation of antibiotics is difficult, they cannot be effectively removed through conventional wastewater treatment technologies. Sulfamethoxazole (SMX) is an extensively used antibiotic and always detected in environmental samples.1–3 Schaider et al. detected SMX in US drinking water supply wells at a concentration of up to 113 ppt.4 The concentration of SMX in surface water ranges from 7.9 to 1900 ppt5–7 and in groundwater ranges from 38 to 450 ppt.8,9To eliminate the negative effects of SMX, biodegradation is the preferred process. Biodegradation holds advantages in operation costs, robustness and application scope compared to chemical oxidation methods, but biodegradation is a time consuming process and it is influenced by the concentration levels of organics. Many researchers studied the biodegradation process of SMX. Jiang et al.10 isolated Pseudomonas psychrophila HA-4 which can utilize SMX as sole carbon and energy source, and 34.3% of SMX can be degraded after 8 days of operation. Bouju et al.11 isolated five strains of SMX-degrading microbe from a membrane bioreactor and found that SMX could be partly mineralized after 16 days of operation. García-Galán et al.12 found that white-rot fungus Trametes versicolor can degrade 9 ppm SMX after 20 hours of culture and Müller et al.13 reported that SMX could be effectively degraded in an activated sludge system. However, there are few reports about the rapid degradation of SMX in sewage treatment reactors and the effect of SMX concentration towards SMX degradation is ambiguous.
Microbial fuel cell (MFC) technology has aroused increasing interest in recent years due to its performance in energy recycling and pollutant degradation. Many published works revealed that MFCs can improve the removal rate of pollutants in sewage14 and soil.15,16 In our previous study, we also found that utilizing MFCs to degrade SMX was a feasible solution.17 However, whether the degradation of SMX relays on its original concentration and the long term performance of MFCs towards SMX removal still bothered us. Moreover, the microbial community structure changes during the process may also offer useful information for further studies about the degradation of SMX.
In this work, we initially studied the removal process of SMX and explored the relationship between removal rate and SMX concentration. Then, the degradation byproducts were examined based on mass spectrometry and a metabolic pathway for SMX has been proposed. To further understand the mechanism, the activity of MFC biofilm was examined by fluorescent staining method, and high-throughput sequencing was employed to analyze the microbial community composition. The microbial community shift during the different operation stages revealed how the microbial community responded to substrate changes. These results afford useful information for studies about SMX-degrading microbial strains.
2 Materials and methods
2.1 Chemicals
Sulfamethoxazole was ordered from Sigma-Aldrich (Munich, Germany); all other chemicals were purchased from Aladdin Industrial Corporation (Shanghai, China). Ultrapure water was prepared by a Milli-Q 18 MΩ system (Millipore, Germany). Water samples were filtered through 0.22 μm-pore-size filters (Millipore, Germany) before test. SMX was dissolved in methanol with a concentration of 25 mg mL−1 and stored as a stock solution in dark place.2.2 Reactor operation
Two-chamber perspex MFC reactors were used in this study. The working volume of each chamber was 115 mL. The anodic and cathodic chambers were separated by a cation exchange membrane (4.0 × 4.0 cm2, Zhejiang Qianqiu Group Co., Ltd., China). Carbon felt (4.0 × 4.0 cm2, Haoshi Carbon Fiber Co., Ltd., China) was used as the electrode and connected with titanium wire (1 mm in diameter, Shanghai GuiTai Titanium Group Co., Ltd., China). Anaerobic sludge was injected into the anodic chamber and the anolyte was changed in batch mode. The cathodic chamber was filled with potassium ferricyanide (100 mM, pH 7.0) and changed biweekly. The voltage of the reactors were monitored by a digital multi-meter (Keithley Instruments, Inc., USA). All reactors were set in a thermostat container (35 ± 2 °C).Initially, artificial wastewater containing sodium acetate (NaAc, 1 g L−1), phosphate buffered solution (20 mM, pH = 7.0) and trace element solution18 was injected into the anodic chamber to start the MFC reactors. After one month’s operation, the voltage output of reactors was maintained at a steady level (590 ± 30 mV). Then, the SMX stock solution (final concentration: 20 ppm) was injected into the reactors instead of NaAc, and the anolyte was changed biweekly. Six months later, the added SMX could be degraded within a week and the anolyte was changed weekly. Twelve months later, we found that SMX could be rapidly removed in all reactors (within 3 days) and the concentration of SMX was increased from 20 ppm to 50 ppm. Fourteen months later, the concentration of SMX in the reactors was increased to 150 ppm and furthermore, to 200 ppm after eighteen months of operation.
2.3 Morphology analysis of the biofilm
Field emission scanning electron microscopy (S4800, Hitachi, Japan) was used to study the morphology of the electrode biofilm. MFC anode samples were rinsed with 20 mM phosphate-buffered saline (PBS) (pH 7.0) 3 times, fixed by 5 wt% glutaric aldehyde in 50 mM PBS (pH 7.0), and dehydrated in ethanol aqueous solutions (25%, 15 min; 50%, 15 min; 75%, 15 min; 90%, 20 min; 95%, 20 min and 100%, 20 min). Then the samples were dried supercritically and coated with gold for SEM observation.SYBR® Green I and Propidium Iodide (PI, Invitrogen, Carlsbad, USA) were utilized to stain the electrode biofilm. Ten microliters of 30 mM PI was dissolved in DMSO and mixed with 1000 μL of SYBR® Green I. The mixture was diluted 100-times as a stock solution. Biofilm samples were torn from the MFC anode and rinsed with 20 mM PBS (pH 7.0) three times to remove loose biomass debris and extracellular polymeric substances. Then the samples were stained with 10 μL stock solution and incubated in the dark at room temperature for 20 mins, and rinsed again with ultrapure water to remove any dye excess. An inverted fluorescence microscope (IX71, Olympus, Japan) was used to detect the samples.
2.4 Three dimensional excitation and emission matrix (3D-EEM) fluorescence spectroscopy analysis
EEM spectra were measured with an F-4600 fluorescence spectrophotometer (Hitachi, Tokyo, Japan). EEM spectra were obtained by recording scanning emission spectra (from 230 to 600 nm at 10 nm intervals) at varied excitation wavelengths (from 200 to 500 nm at 10 nm intervals). Excitation and emission slits were set at 5 nm, and the scanning speed was maintained at 240 nm min−1. The PMT voltage was set at 700 V. The spectrum of 20 mM PBS was recorded as a blank control.2.5 Analysis of SMX degradation byproducts
The reactor effluent samples were analyzed with a Thermo Scientific Q-Exactive hybrid quadrupole-Orbitrap mass spectrometer (San Jose, CA, USA) under positive electrospray mode. Chromatography was performed with an ultra-performance liquid chromatography (UPLC, Waters, USA) system at a flow rate of 200 μL min−1 on a Waters C18 column (Acquity UPLC BEH C18, 2.1 × 100 mm, 1.7 μm). Acetonitrile was used as the mobile phase. The collision energy was set as 28.0 for HCD fragmentation.2.6 Microbial community analysis
In order to study the microbial community change during the SMX removal process, bio-samples at different operation stages were collected. Stage 1, raw sludge, which was injected into the reactors. Stage 2, MFC biofilm samples utilizing NaAc as the substrate. The samples were collected from the anode after one month of operation. Stage 3, MFC biofilm samples using SMX as the substrate. The samples were collected from the anode after 10 months of operation. All samples were rinsed with ultrapure water and stored at −20 °C for further analysis.Electrode biofilm sample was torn from the top, middle and bottom part of the carbon felt. After that, the samples were mixed and minced with a sterile scalpel. DNA extraction and purification were carried as previously reported.19 Extracted DNA samples were quantified with a micro-ultraviolet spectrophotometer (Nanodrop Inc., USA). Purified DNA samples were used as templates for PCR amplification as previously reported.20 The PCR products were quantitated with QuantiFluor™-ST and mixed for Illumina Miseq sequencing.
Quantitative Insights Into Microbial Ecology software (QIIME, version 1.8) was used to process the Miseq data of 16S rRNA gene sequences. Miseq data was merged before data analysis and the completeness of the barcodes and primer sequencing was checked by QIIME. The reads which were shorter than 200 bp and quality score below 25 were removed. After that, the sample sequences were labeled with 6 bp barcodes respectively. Only those sequences with above 97% identity were divided into OTUs (Operational Taxonomic Units) for further data analysis. The most abundant sequence of each OTU was defined as the representative sequence and assigned by PyNAST, and these representative sequences were employed for the classification of the taxonomy according to the Greengenes database. We randomly chose 13![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 056 sequences per sample to explore the α-diversity, and compared the β-diversity between samples. The count of OTUs, Chao1 index, Shannon index and phylogenetic diversity index of each sample was calculated.
056 sequences per sample to explore the α-diversity, and compared the β-diversity between samples. The count of OTUs, Chao1 index, Shannon index and phylogenetic diversity index of each sample was calculated.
3 Results and discussion
3.1 Reactor startup and operation
When considering the biodegradation of recalcitrant contaminants, the function of microbe is vital. Anaerobic sludge was collected from a coal gasification wastewater treatment plant21 which mainly disposes phenolics, poly-nuclear aromatic hydrocarbons and heterocyclic compounds. The anaerobic sludge was injected into the MFC reactors and NaAc was fed as the substrate to start the MFC reactors. Fig. 1A illustrates the reactor’s voltage fluctuation curve during the process. Reactor voltages soared to 260 ± 50 mV after several days of operation. Since the voltage of the MFC is related to the activity of the electrochemically active biofilm, the rapid increasing of the cell voltage revealed the success startup of MFC system. After several operation cycles, the effluent was turbid and bubbles were formed on the anode. During this period, the output voltage was maintained at 580 ± 30 mV, indicating that an electrochemically active biofilm was forming and the microbe was active.After a month of operation, the output voltage of the reactors was in a steady state. Then, NaAc was replaced with the SMX stock solution (20 ppm SMX). We found that the turbidity of the anolyte was greatly reduced and the effluent became odorless, with the voltage dropping to 15 ± 6 mV. As a broad-spectrum antibacterial agent, it seemed that SMX had greatly inhibited the activity of the microorganisms. However, in the following 2–4 months of operation, the voltage of the reactors was gradually increased. Five months later, the voltage stably raised to 640 ± 40 mV at each batch operation, indicating that the microbes in the MFCs could utilize SMX for metabolism and electricity production.
The surface morphology of the anode after 10 months of operation is shown in Fig. 1B. The biofilm was uniformly attached on the carbon felt and bubbles would occasionally form on the biofilm. We tore some biofilm from the carbon felt electrode and examined it with a microscope (Fig. 1C). It was found that the biofilm was firmly attached to the carbon felt fibers. The strong connection between biofilm and electrode fiber would favor the energy production of MFC system. Fig. 1D shows the microscopic morphology of the electrode biofilm. Different genera of microorganisms (Sarcina, Diplococcus, Coccus, Bacillus etc., mixed with cell debris and secretion) can be observed in the image, suggesting a rich community was formed.
The concentration of SMX in the supernatant was monitored during the whole experiment. After months of operation, the removal ability towards SMX gradually increased. After analyzing and fitting the data, and on comparison with previous studies22,23 about the biodegradation of recalcitrant chemicals, we inferred that the concentration change of SMX followed (pseudo) first order kinetics. Fig. 2A shows the half-life time of SMX in the MFC reactors as a function of time. Four months later, 50% SMX (20 ppm) could be removed in MFCs within 10 days. Ten months later, the half-life of SMX in reactors decreased to 1 day, and 20 ppm SMX could be fully removed within 3 days. These data suggested that the high removal efficiency of SMX can be achieved through acclimation.
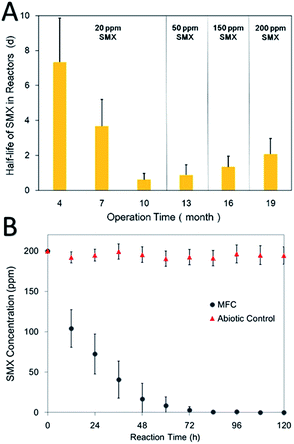 | ||
| Fig. 2 Half-life time of SMX in reactors as a function of time (A) and concentration change of 200 ppm SMX in MFCs and abiotic control after 18 months of operation (B). | ||
Since the SMX (20 ppm) was rapidly removed in the reactors, whether the reactors could remove higher concentrations of SMX aroused our interest. The concentration was gradually increased to 200 ppm after 18 months of operation. Around 70% SMX could be removed within 24 hours and its concentration was below the detection limit after 96 hours operation, as shown in Fig. 2B. However, the concentration of SMX showed no obvious change in the abiotic control reactor. When the concentration was increased from 20 ppm to 200 ppm, the half-life time of SMX in the reactors merely increased from 1 day to 3 days. Xu et al. also noticed that the degradation of SMX in their bioreactors was not depended on its original concentration, and speculated that co-metabolism may occur in the biodegradation of SMX.24
3.2 Degradation mechanism of SMX
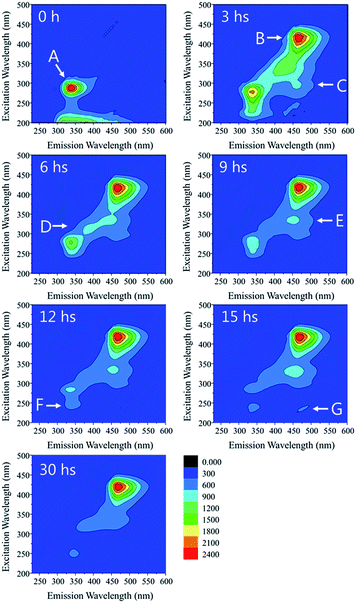
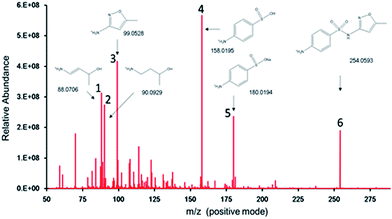
In above section we discussed the concentration change of SMX in reactors. However, whether the SMX had been degraded was still uncertain. Fluorescence EEM analysis was carried to study the structural change of SMX. Fig. 3 illustrates the fluorescence of the reactor supernate sampled at different times. Peak A (λex/λem = 290/340) can be attributed to SMX since the influent contained no other organics and there was no fluorescence peak in the blank sample (i.e. the reactor influent without SMX). The intensity of peak A decreased and completely vanished after 30 hours of operation, while the intensity of peak B (λex/λem = 420/470) was increased. This phenomenon revealed that the chemical structure of SMX and its metabolism byproducts changed as a function of time, and this could be attributed to the degradation process. A few scattered peaks (C, D, E and G) appeared and then vanished during the process, indicating the transformation of the fluorescent chemicals. Previous studies25,26 reported that peak B belongs to coenzyme F420, a type of coenzyme which acts as an electron carrier in both anabolic and catabolic redox reactions27 of methanogenic bacteria. Considering the fact that the operation of the MFCs is generally accompanied with the formation of electron transfer mediators, we speculate that some degradation products of SMX might be utilized by the microbe to synthesise exoenzymes or other electrochemically active substances. Moreover, coenzyme F420 may enhance the bioelectrochemical process in our system as an electron transfer mediator.To further understand the degradation pathway of SMX, Orbitrap mass spectrometry was employed to detect the degradation byproducts. Since the detection was carried under positive mode, organics were combined with a cation and the detected m/z values were 1 unit (–NH2 combined with H+), 22 units (–COOH combined with Na+) or 38 units (–COOH combined with K+) higher than the original molecules. Several potential metabolites were identified from the total ion current chromatogram spectra (Fig. 4A). Previous studies reported that 3-amino-5-methylisoxazole (3A5MI) appeared during the degradation of SMX, and we also found 3A5MI in the samples (peak 3, m/z 99.0528). The m/z difference between peak 4 and peak 5 is 22.0001, indicating that chemical 5 may be the sodium salt of chemical 4, and that chemical 4 is an organic acid. We speculate that chemical 4 is 4-amino benzene sulfinic acid. During the degradation process, we noticed that 3A5MI could be further degraded in the reactors, and speculated that chemical 1 was a degradation product of 3A5MI. Based on the identified degradation byproducts, the degradation pathway of SMX in MFC reactors is proposed as follows. SMX is initially hydrolyzed into 4-amino benzene sulfinic acid and 3A5MI. Then, the isoxazole ring of 3A5MI is opened and its nitrogen atom removed, resulting in the formation of 4-amino-2-butanol. Since 4-amino-2-butanol is a chain structure, it is easily degraded by microbes.
3.3 Microbial analysis
During acclimation, the electrode biofilm got thicker and thicker, which may influence the activity of microbe and hinder the mass transfer from solution into the biofilm (bred in the middle part of the carbon felt electrode). The activity of the biofilm was estimated by a fluorescent staining method (Fig. 5). Most of the cells presented intense green fluorescence, indicating that their morphologic structure was intact and that they were in an active state. The orange spots represent damaged cells which are dead or in an apoptotic state, but there were only a few microbes in this state. Moreover, a layer of active microbes were uniformly attached to the surface of the carbon felt fiber (with some floc twisted around it). This phenomenon indicates that the microbes bred in the MFC carbon felt presented a great activity.Interestingly, during the microscopic examination of the biofilm samples, we noticed that some suspected metazoans such as roundworms (Fig. 5A and S1†) were presented in the reactors. Generally, the appearance of metazoans reveals that a food chain has been created in the bioreactors and the water quality is not bad.28 Besides, metazoans can devour cell debris and facilitate the regeneration of the biofilm. It seems that the degradation products in the MFCs may not hinder the growth and breed of metazoans.
According to the high-throughput sequencing results in Fig. 6, the OTU number of raw sludge was the largest compared to other samples, indicating that the microbial composition of raw sludge was rather complex. Since the raw sludge originated in coal gasification wastewater, diverse microbial species were conducive for the degradation of complex substrates. However, when the substrate was changed to NaAc or SMX while operating the reactor under MFC mode, the diversity of the microbial community was largely decreased. This may be partly ascribed to the selectivity of the MFC system, as the substrate was solely for microbe degradation and hence influenced the community structure. In addition, the bio-samples in NaAc and SMX stages were the biofilms, while the sample of stage 1 was the sludge, therefore the difference in the operation mode could also greatly influence the community structure. According to PCoA results, we noticed that when different reactors were fed with same substrate (NaAc or SMX), the bio-samples tended to cluster together. This indicates that certain species may be favoured for the consumption of different substrates and they may gradually dominate the microbial community. On the other hand, the distance between the samples fed with SMX or NaAc is rather large, revealing that the substrate has a great influence on the composition of the microbial community.
 | ||
| Fig. 6 Weighted principle coordinate analysis (PCoA) of the microbial communities sampled at different stages. The size of the dot represents the OTU number of each sample. | ||
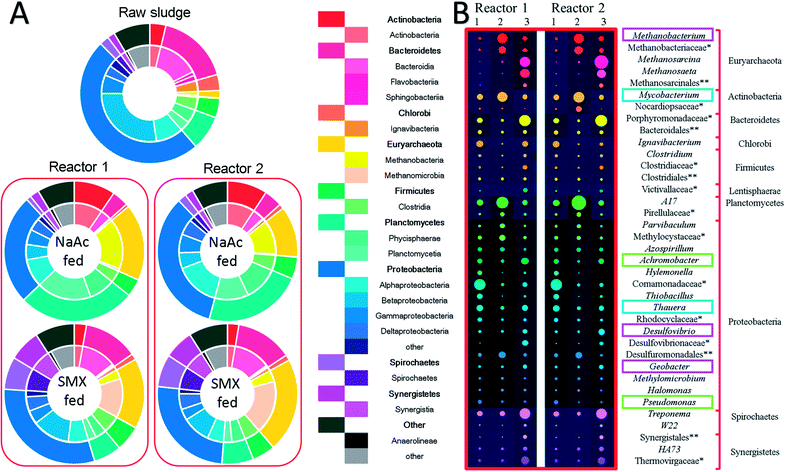
Raw sludge, NaAc-fed and SMX-fed bio-samples were examined by high-throughput sequencing method to study the shift in the microbial community structure. Fig. 7A shows the microbial phyla of bio-samples collected at different stages. In raw sludge, Proteobacteria (49%) was the largest phylum, suggesting that Proteobacteria may contribute a lot in the disposal of coal gasification wastewater. However, when the substrate was changed and operated in closed circuit mode, the community share of Proteobacteria decreased to 25.4–33.6%. On the other hand, Euryarchaeota (1.7%) was the smallest phylum in raw sludge but it increased to 16.2–21.7% in MFC when the reactor was fed with SMX. The relative abundance of Planctomycetes in raw sludge accounted for merely 8%. When NaAc was fed into the MFCs, the relative abundance of Planctomycetes increased to 25% and its abundance decreased to 7% in the biofilm when SMX was fed as substrate.
The heat map (background image) of Fig. 7B presents the relative abundance of microbial phyla while the size of the bubbles in the chart represent the relative abundance of the different microbial genera identified in the samples. Generally, the relative abundance of the different microbial genera in raw sludge was balanced, but the relative abundance of some genera was largely increased when fed with NaAc (Methanosarcina and Mycobacterium) or SMX (Methanobacterium, Methanosaeta and Treponema). This may be due to the carbon source of the raw sludge being rather complex, and different genera grew together to maintain a balance towards substrate degradation.
On genus level, Methanosarcina, Mycobacterium and other two unnamed genera (respectively belonging to Methanobacteriaceae and Pirellulaceae) were largely bred in the MFCs when fed with NaAc. Theoretically, methane production adversely affects the energy production in MFCs, since the conversion of organics into methane require electrons. On the other hand, it would benefit pollutant degradation, since different consortia have to compete for substrate utilization which may lead to a more effective degradation performance towards recalcitrant chemicals. Moreover, the formation of methane-producing consortia also revealed that macromolecules could be degraded into small organics in the system. When the substrate was changed from NaAc to SMX, the microbial community changed greatly: the number of Methanobacterium, Methanosaeta, Treponema, Achromobacter and an unnamed genus (belonging to the Porphyromonadaceae family) largely increased. As SMX is an effective antimicrobial agent, the appearance of methanogens demonstrates that the biotoxicity of SMX can be effectively reduced.
Methanobacterium and Methanosaeta may also contribute a lot to the mineralization of some degradation products of SMX; the formation of coenzyme F420 based on the 3D-EEM test may also be related to their metabolic activity. Some species of Achromobacter and Pseudomonas hold a strong degradation ability towards SMX according to published results,10,29 and their appearance in reactors supports our previous discussion that SMX could be biodegraded in MFCs. Besides, Mycobacterium, Ignavibacterium, and Thauera also had a higher relative abundance in the biofilm samples. These genera are generally found in phenol-contaminated soils or sewage systems and are capable for the degradation of aromatic hydrocarbons.30–32 The flourishment of these microbes reveal that functional groups useful for SMX degradation formed in the reactor biofilms and this may be related to the rapid degradation performance of SMX in MFCs.
4 Conclusions
In this study, we explored the possibility of utilizing MFC reactors to remove SMX and studied the chemical and biological mechanism during the process. We found that SMX could be biodegraded in MFCs after acclimation and even high concentrations of SMX (i.e. 200 ppm) could be removed. The degradation process was studied by 3D-EEM and Orbitrap mass spectrometery, and a degradation mechanism was subsequently proposed. We then examined the activity of the MFC biofilms and noticed that some metazoans were present in the reactors, demonstrating that the removal performance of MFC was great. High-throughput sequencing technology was used to study the microbial community shift during the SMX elimination process and some reported SMX degradation microbes were identified in the system. This study offers a feasible choice for the elimination of SMX in wastewater which may be extended to other antibiotics.Acknowledgements
This work was supported by NSFC (21322703, 21177122 and 41471260) and the Natural Science Foundation for Distinguished Young Scholars of Fujian Province (2014J06007). We gratefully appreciate the help from Mr Danyang Li for providing the anaerobic sludge.References
- M. Petrović, M. D. Hernando, M. S. Díaz-Cruz and D. Barceló, J. Chromatogr. A, 2005, 1067, 1–14 CrossRef PubMed
.
- T. A. Ternes, TrAC, Trends Anal. Chem., 2001, 20, 419–434 CrossRef CAS
.
- M. J. Hilton and K. V. Thomas, J. Chromatogr. A, 2003, 1015, 129–141 CrossRef CAS
.
- L. A. Schaider, R. A. Rudel, J. M. Ackerman, S. C. Dunagan and J. G. Brody, Sci. Total Environ., 2014, 468–469, 384–393 CrossRef CAS PubMed
.
- K. Choi, Y. Kim, J. Park, C. K. Park, M. Kim, H. S. Kim and P. Kim, Sci. Total Environ., 2008, 405, 120–128 CrossRef CAS PubMed
.
- J. V. Holm, K. Ruegge, P. L. Bjerg and T. H. Christensen, Environ. Sci. Technol., 1995, 29, 1415–1420 CrossRef CAS PubMed
.
- J. M. Conley, S. J. Symes, S. A. Kindelberger and S. M. Richards, J. Chromatogr. A, 2008, 1185, 206–215 CrossRef CAS PubMed
.
- R. Loos, G. Locoro, S. Comero, S. Contini, D. Schwesig, F. Werres, P. Balsaa, O. Gans, S. Weiss and L. Blaha, Water Res., 2010, 44, 4115–4126 CrossRef CAS PubMed
.
- E. Godfrey, W. W. Woessner and M. J. Benotti, Ground Water, 2007, 45, 263–271 CrossRef CAS PubMed
.
- B. Jiang, A. Li, D. Cui, R. Cai, F. Ma and Y. Wang, Appl. Microbiol. Biotechnol., 2014, 98, 4671–4681 CrossRef CAS PubMed
.
- H. Bouju, B. Ricken, T. Beffa, P. F.-X. Corvini and B. A. Kolvenbach, Appl. Environ. Microbiol., 2012, 78, 277–279 CrossRef CAS PubMed
.
- M. J. García-Galán, C. E. Rodríguez-Rodríguez, T. Vicent, G. Caminal, M. S. Díaz-Cruz and D. Barceló, Sci. Total Environ., 2011, 409, 5505–5512 CrossRef PubMed
.
- E. Müller, W. Schüssler, H. Horn and H. Lemmer, Chemosphere, 2013, 92, 969–978 CrossRef PubMed
.
- T. Zhang, S. M. Gannon, K. P. Nevin, A. E. Franks and D. R. Lovley, Environ. Microbiol., 2010, 12, 1011–1020 CrossRef CAS PubMed
.
- X. Wang, Z. Cai, Q. Zhou, Z. Zhang and C. Chen, Biotechnol. Bioeng., 2012, 109, 426–433 CrossRef CAS PubMed
.
- L. Lu, T. Huggins, S. Jin, Y. Zuo and Z. J. Ren, Environ. Sci. Technol., 2014, 48, 4021–4029 CrossRef CAS PubMed
.
- J.-H. Ahn, S. G. Shin and S. Hwang, Chem. Eng. J., 2009, 153, 145–150 CrossRef CAS PubMed
.
- D. R. Lovley and E. J. Phillips, Appl. Environ. Microbiol., 1988, 54, 1472–1480 CAS
.
- Z. H. Yang, Y. Xiao, G. Zeng, Z. Y. Xu and Y. S. Liu, Appl. Microbiol. Biotechnol., 2007, 74, 918–925 CrossRef CAS PubMed
.
- Y. Zheng, C. Wang, Z. Y. Zheng, J. Che, Y. Xiao, Z.-H. Yang and F. Zhao, RSC Adv., 2014, 4, 62544–62549 CAS
.
- C. Cui, F. Ma, Y. Zhang and H. Li, J. Harbin Univ. Civ. Eng. Archit., 2002, 6, 006 Search PubMed
.
- S. Suarez, J. M. Lema and F. Omil, Water Res., 2010, 44, 3214–3224 CrossRef CAS PubMed
.
- B. Li and T. Zhang, Environ. Sci. Technol., 2010, 44, 3468–3473 CrossRef CAS PubMed
.
- B. Xu, D. Mao, Y. Luo and L. Xu, Bioresour. Technol., 2011, 102, 7069–7076 CrossRef CAS PubMed
.
- F. Dong, Q.-B. Zhao, J.-B. Zhao, G.-P. Sheng, Y. Tang, Z.-H. Tong, H.-Q. Yu, Y.-Y. Li and H. Harada, Bioresour. Technol., 2010, 101, 1722–1726 CrossRef CAS PubMed
.
- W. G. Li, C. Y. Su, X. Z. Liu and L. Zhang, Environ. Sci. Pollut. Res., 2014, 21, 8167–8175 CrossRef CAS PubMed
.
- J. Dolfing and J.-W. Mulder, Appl. Environ. Microbiol., 1985, 49, 1142–1145 CAS
.
- N. Kinner and C. Curds, Water Res., 1987, 21, 481–490 CrossRef CAS
.
- P. J. Reis, A. C. Reis, B. Ricken, B. A. Kolvenbach, C. M. Manaia, P. F. Corvini and O. C. Nunes, J. Hazard. Mater., 2014, 280, 741–749 CrossRef CAS PubMed
.
- B. L. Burback and J. J. Perry, Appl. Environ. Microbiol., 1993, 59, 1025–1029 CAS
.
- X. Zhu, J. Tian, C. Liu and L. Chen, Appl. Microbiol. Biotechnol., 2013, 97, 8767–8775 CrossRef CAS PubMed
.
- Y. Mao, X. Zhang, X. Xia, H. Zhong and L. Zhao, J. Ind. Microbiol. Biotechnol., 2010, 37, 927–934 CrossRef CAS PubMed
.
Footnote |
| † Electronic supplementary information (ESI) available. See DOI: 10.1039/c5ra08438e |
| This journal is © The Royal Society of Chemistry 2015 |