Sulforaphane and erucin, natural isothiocyanates from broccoli, inhibit bacterial quorum sensing†
Hadas
Ganin
,
Josep
Rayo
,
Neri
Amara
,
Niva
Levy
,
Pnina
Krief
and
Michael M.
Meijler
*
Department of Chemistry and National Institute for Biotechnology in the Negev, Ben-Gurion University of the Negev, Be'er-Sheva 84105, Israel. E-mail: meijler@bgu.ac.il; Fax: +972 8 6472943; Tel: +972 8 6472751
First published on 31st August 2012
Abstract
Sulforaphane and erucin, two natural isothiocyanates that are highly abundant in broccoli and other cruciferous vegetables, were found to strongly inhibit quorum sensing and virulence in Pseudomonas aeruginosa. Mechanistic evaluations of these effects suggest that these isothiocyanates are antagonists of the transcriptional activator LasR.
In recent decades there has been a strong rise in interest and awareness regarding the personal control of our health and prevention of disease, especially in the area of nutrition. In this respect, a well-studied group of vegetables that has raised significant interest is the brassicaceae family, which includes cruciferous vegetables such as broccoli, Brussels sprouts, cabbage, cauliflower, kohlrabi and kale.1 Recent reports have attributed to these food sources a chemoprotective effect against different types of cancer, such as prostate, bladder, lung, colorectal, and pancreatic cancer,2–4 through inhibition of tumor cell proliferation and induction of apoptosis,5 as well as antibacterial effects, such as inhibition of Helicobacter pylori growth.6 The beneficial properties of these vegetables have mostly been attributed to abundant secondary metabolites known as glucosinolates and their enzymatic hydrolysis products, isothiocyanates.7,8 The enzyme myrosinase (thioglucoside glucohydrolase) transforms glucoraphanin (β-thioglucoside-N-hydroxysulfate) into the bioactive compound sulforaphane (1) upon damaging the plant, such as chewing or chopping.9 Posner and coworkers first identified sulforaphane from broccoli as a potent inducer of anticarcinogenic enzymes and elucidated its structure (Fig. 1).
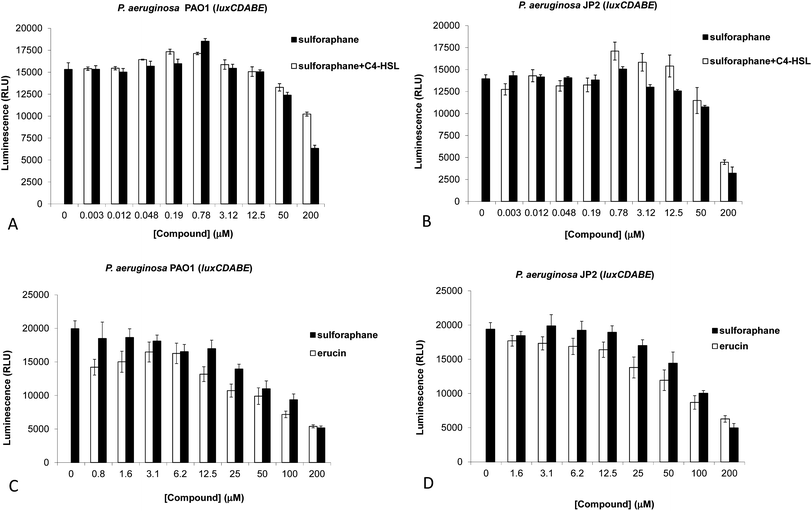 | ||
Fig. 1 Inhibition of QS by different concentrations of sulforaphane, with or without C4-HSL (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 concentration), in P. aeruginosa PAO1-luxCDABE (A) or PAO-JP2-luxCDABE (B) (triplicates, after 5 h of incubation). Activity of different concentrations of sulforaphane and erucin in P. aeruginosa PAO1-luxCDABE (C) or PAO-JP2-luxCDABE (D) (triplicates, after 5 h of incubation). Activity is shown as relative luminescence units (RLU – luminescence divided by the OD600 of the bacteria). Measurements in (A) and (B) were performed using a Spectramax M2 (Molecular Devices) plate reader, and in (C) and (D) using a Varioskan Flash Multimode (Thermo Scientific) plate reader. 1 concentration), in P. aeruginosa PAO1-luxCDABE (A) or PAO-JP2-luxCDABE (B) (triplicates, after 5 h of incubation). Activity of different concentrations of sulforaphane and erucin in P. aeruginosa PAO1-luxCDABE (C) or PAO-JP2-luxCDABE (D) (triplicates, after 5 h of incubation). Activity is shown as relative luminescence units (RLU – luminescence divided by the OD600 of the bacteria). Measurements in (A) and (B) were performed using a Spectramax M2 (Molecular Devices) plate reader, and in (C) and (D) using a Varioskan Flash Multimode (Thermo Scientific) plate reader. | ||
Sulforaphane inhibits Phase I enzymes, which are responsible for activation of carcinogens, and it induces Phase II detoxification enzymes, which are able to detoxify carcinogens.4,10 Additional health benefits that have been attributed to sulforaphane are anti-inflammatory as well as antibiotic.6,11 Due to the increase in the prevalence of antibiotic resistance among pathogenic bacteria, such as the methicillin-resistant Staphylococcus aureus (MRSA) and Pseudomonas aeruginosa,12,13 significant research efforts have been devoted in recent years to the development of alternative strategies to inhibit bacterial growth and virulence.
One of these strategies is to inhibit bacterial quorum sensing. This phenomenon is the process in which bacteria communicate in order to coordinate gene expression. It allows bacteria to control the population-wide expression of important genes through secretion and sensing of small signaling molecules termed autoinducers (AIs). Among the activities controlled by QS are generation of bioluminescence, conjugation, antibiotic production, motility, sporulation, biofilm formation and siderophore production.14 These coordinated behaviors allow bacteria to compete with multicellular organisms and survive harsh and sudden environmental changes.15 Thus, inhibition of this crucial process has been proposed as a promising strategy to combat bacterial infections and several successful applications have been reported.16–18
Autoinducers (AIs) can be grouped into distinct classes according to their basic chemical characteristics: (i) N-acyl homoserine lactones (AHLs) are produced in Gram-negative bacteria; (ii) autoinducing peptides (AIPs) are generally used by Gram-positive bacteria; (iii) a family of AIs termed AI-2s, used by both Gram-positive and Gram-negative species and (iv) a relatively newly discovered group, the α-hydroxyketones, which are mainly produced by aquatic Gram-negative bacteria including Legionella and Vibrio species. The most extensively studied AIs are the homoserine lactones, and in particular N-(3-oxo-dodecanoyl)-L-homoserine lactone (termed 3-oxo-C12-HSL) from P. aeruginosa. This highly abundant organism is a Gram-negative opportunistic human pathogen that causes serious infections in immunocompromised individuals, such as cystic fibrosis (CF) patients, AIDS patients, burn victims and those requiring long-term hospitalization.
At least two major QS systems – las and rhl – regulate group behavior in these bacteria, in addition to other regulators such as qsc. The las system consists of the synthase LasI, which produces 3-oxo-C12-HSL, and the transcriptional regulator LasR, while the rhl system similarly consists of the pair RhlI/RhlR, which responds to N-butyrylhomoserine lactone (C4-HSL).19,20
Based on structural and characteristic similarities between sulforaphane and certain QS molecules, such as its low molecular weight, relative polarity and hydrophobicity, we decided to examine the effects of sulforaphane and its deoxy analog erucin on QS in P. aeruginosa. In addition, as we recently synthesized isothiocyanate containing AHL analogs that inhibit P. aeruginosa QS through covalent binding of its transcriptional regulator LasR,17 we hypothesized that sulforaphane and erucin might react with LasR as well, should they possess reasonable affinities for its binding pocket. In the course of our investigations Givskov and coworkers reported two important studies in which sulfur-rich natural products were shown to inhibit bacterial QS, thereby strengthening our hypothesis. In the first study they showed that the sulforaphane analog iberin, from horseradish, as well as several analogs (including sulforaphane itself) show potency as QS inhibitors in P. aeruginosa,21 while the second study focused on ajoene and derivatives that are found in garlic, and here too potent QS inhibitory activity was observed.22
Results and discussion
Sulforaphane was first isolated and identified by Zhang et al. in 1992. Sulforaphane isolated from broccoli is chiral, possessing the R configuration, but both R-sulforaphane and the synthetic (R,S) mixture show identical activities in reported assays.4,23In order to obtain large amounts of test compounds, we synthesized racemic 4-methylsulfinylbutyl isothiocyanate (sulforaphane) and its precursor 4-methyl thiobutyl isothiocyanate (erucin), following several modifications of synthetic strategies reported by Ding et al. and Vermeulen et al.24,25 Briefly, 1,4-bromobutanol was reacted with sodium azide to afford compound 3 (Scheme 1). Treatment of 3 with tosylchloride gave the tosylated compound 4, followed by reaction with NaSMe in dry THF (instead of DMF) with mild heating for 4 hours to give the highly volatile azido thioether 5. This compound was then reacted with triphenylphosphine (PPh3) and the triphenyl phosphoramidite intermediate was in situ reacted with carbon disulphide,26 affording erucin (2). Finally, 1 was obtained after oxidizing the sulfide moiety with mCPBA (m-chloroperbenzoic acid) in good yield.25
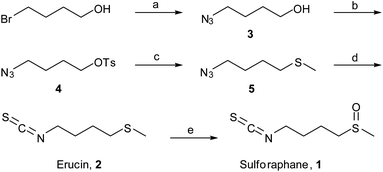 | ||
| Scheme 1 Synthesis of (S,R)-sulforaphane and erucin. Reagents and conditions: (a) NaN3, DMF, 70 °C, 24 h, 83%; (b) TsCl, Et3N, 4-DMAP, 0 °C CH2Cl2, 52%; (c) MeSNa, THF, 4 h, 55 °C, 39%; (d) PPh3, diethyl ether, CS2, 81%; (e) mCPBA, CH2Cl2, 77%. | ||
Both sulforaphane and its precursor erucin were evaluated for their ability to activate or interfere with the QS systems las and rhl of P. aeruginosa. The activities of the compounds were evaluated using several reporter strains of P. aeruginosa: the luminescent PAO1-luxCDABE, PAO-JP2-luxCDABE, and PAO-JP2-(Pkd-rhlA) strains. PAO1-luxCDABE is a P. aeruginosa wild-type strain modified with a luminescent reporter gene luxCDABE cloned downstream of the synthase lasI, and PAO-JP2-luxCDABE is a similar strain, but it lacks the 3-oxo-C12-HSL and C4-HSL synthetases and these strains are used in order to assess whether the probes could specifically mimic or antagonize 3-oxo-C12-HSL. The PAO-JP2 (Pkd-rhlA) strain is used to evaluate specific activation of the rhl system (i.e. response to C4-HSL). An E. coli DH5α strain harboring the LasR expression vector, pJN105L, and a plasmid-borne PlasI–lacZ fusion (pSC11) were used to quantify lasR activation by measuring expression levels of β-galactosidase.27
We measured the effects of sulforaphane on 3-oxo-C12-HSL induced QS activation in the presence or absence of the secondary QS signal in P. aeruginosa, the N-butyryl-L-homoserine lactone (C4-HSL). Since a combination of sulforaphane and C4-HSL could, in theory, occupy the LasR binding pocket well, we examined whether addition of the HSL moiety would increase the activity of sulforaphane.
Experiments with PAO1-luxCDABE revealed inhibition of LasR activation at sulforaphane concentrations of 50 μM and above (Fig. 1A), with strong inhibition seen at 200 μM – although it should be noted that we observed some variation in sensitivity between different experiments. In order to assess whether sulforaphane could specifically mimic or antagonize 3-oxo-C12-HSL, experiments were repeated with PAO-JP2-luxCDABE (a lasI/rhlI knockout strain). In the absence of 3-oxo-C12-HSL no activation of LasR was observed. However, in the presence of its cognate autoinducer (100 nM) clear inhibition of LasR activation was observed for sulforaphane concentrations of 50 μM and above (Fig. 1B). Furthermore, at concentrations up to 100 μM of sulforaphane we did not observe significant differences in both strains upon inclusion or omission of C4-HSL, indicating that sulphoraphane by itself may bind its presumed target (LasR) sufficiently strong. At concentrations above 100 μM we observed growth-inhibitory effects for sulforaphane (Fig. S2†).
Next, in order to examine the role of the sulfoxide moiety we compared the activities of sulforaphane and its deoxy precursor erucin – also present in large amounts in broccoli, as well as in rocket (Eruca sativa)28 – in assays with PAO1-luxCDABE and PAO-JP2-luxCDABE (Fig. 1C and D). In both strains the same attenuation was observed as in the previous experiments, showing slightly stronger QS inhibitory effects for ER.
In addition to studies relying on P. aeruginosa strains, we performed experiments using an E. coli DH5 (pJN105L) (pSC11) reporter strain. This heterologous strain harbours LasR from P. aeruginosa, it does not produce 3-oxo-C12-HSL, and reports binding of this QS molecule upon LasR binding through β-galactosidase production. While QS activation in P. aeruginosa is known to be quite sensitive to changes in environmental conditions, the E. coli reporter assay is considered to be more robust and highly specific for activation by 3-oxo-C12-HSL.29 In order to verify whether both sulforaphane and erucin showed specific 3-oxo-C12-HSL antagonist activity, the reporter gene expression was measured following the addition of sulforaphane and erucin in the presence of 100 nM 3-oxo-C12-HSL. In agreement with the P. aeruginosa reporter assays a similar trend of concentration dependent inhibition was observed (Fig. 2), though in the E. coli assay sulforaphane appears to be a more effective LasR inhibitor than erucin.
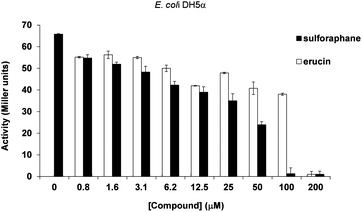 | ||
| Fig. 2 Inhibition of 3-oxo-C12-HSL (100 nM) mediated LasR activation in E. coli-DH5a-lacZ by increasing concentrations of sulforaphane and erucin. Activation was measured through β-galactosidase production (Miller units). | ||
We also examined the effects of sulforaphane and erucin on activation of RhlR. The autoinducer of this system, C4-HSL (10 μM), was added to a rhlA promoter-based P. aeruginosa strain, PAO-JP2 (pKD-rhlA), and attenuation of RhlR activation was measured. We observed similar behavior in other strains, with significant inhibition of QS activity starting from 25 μM of sulphoraphane and erucin. No significant inhibition of growth was observed for erucin, while for sulforaphane slight inhibition of growth was observed at high concentrations (>100 μM) (Fig. S2†).
We further examined the mechanism of action behind the concentration dependent inhibition of QS in P. aeruginosa for both compounds by liquid chromatography/mass spectrometry (LC/MS). As our hypothesis from the start of these investigations had been that the isothiocyanate moieties of sulforaphane and erucin might be able to covalently bind to LasR, due to the presence of the nucleophilic cysteine residue in its binding pocket17 we expressed the ligand binding domain of this protein in the presence of the probes (at concentrations of 50 and 100 μM). Only minute amounts of soluble LasR were observed by gel electrophoresis, indicating that the nascent protein does not fold around sulforaphane or erucin properly, preventing correct folding and resulting in precipitation of LasR. In our expression assays LasR was mainly observed in the pellet, and upon solubilization of the pellet and purification of LasR we analyzed the protein by LC/MS. While we did observe the unmodified LasR (Mw 22![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 430 Da), we did not observe a significant amount of LasR with covalently bound sulforaphane or erucin (expected Mw 22
430 Da), we did not observe a significant amount of LasR with covalently bound sulforaphane or erucin (expected Mw 22![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 606, 22
606, 22![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 590 Da respectively). We did, however, note a number of other molar masses in close proximity, indicating possible further modifications that may have occurred either upon binding of the ligand or in the course of the purification process. Given the difficulties that we encountered in the process of solubilization of the pellet, we cannot rule out that (at least part of) LasR does react with the isothiocyanate moiety of sulforaphane or erucin.
590 Da respectively). We did, however, note a number of other molar masses in close proximity, indicating possible further modifications that may have occurred either upon binding of the ligand or in the course of the purification process. Given the difficulties that we encountered in the process of solubilization of the pellet, we cannot rule out that (at least part of) LasR does react with the isothiocyanate moiety of sulforaphane or erucin.
Next, we set out to examine the effects of sulforaphane and erucin on important quorum sensing controlled phenotypes, such as biofilm formation and pyocyanin (a major virulence factor of P. aeruginosa) production in wild type P. aeruginosa strain PAO1. We observed a strong and reproducible reduction of biofilm formation at sulforaphane concentrations of 12 μM and above (Fig. 3A), indicating anti-biofilm activity that is even slightly stronger than that of the known effective anti-QS and anti-biofilm natural product bromofuranone C30, which we included in our assays as a positive control. Surprisingly, the inhibitory activity of erucin on biofilm formation appeared markedly lower than that of sulforaphane (while in the P. aeruginosa QS assays the compounds display similar activities), and this difference may be attributed to the small difference in overall hydrophobicity between the two compounds, resulting in differences in permeation and distribution in the growing biofilm – while in planktonic culture these small differences are less relevant. Alternatively, the difference may be explained by a slower oxidation of erucin to sulforaphane in the biofilm environment. Upon examination of the effects of both compounds on pyocyanin production we did not observe significant differences between both compounds, as both erucin and sulforaphane showed strong activity. Wild type P. aeruginosa strain PAO1 was grown in the presence of 100 μM sulforaphane, erucin or 0.1% DMSO for 20 hours, and more than 70% inhibition of pyocyanin production was observed for both isothiocyanates (Fig. 3B).
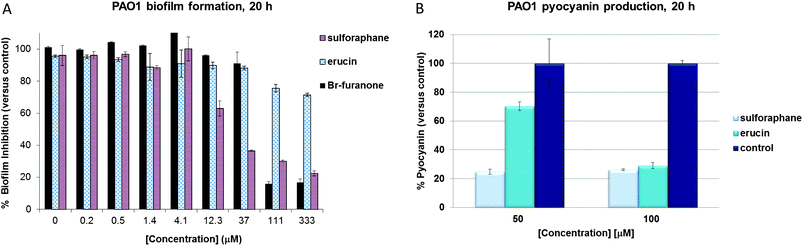 | ||
| Fig. 3 Inhibition of biofilm formation (A) and pyocyanin production (B) in the presence of sulforaphane and erucin (50 and 100 μM for the pyocyanin experiment) in P. aeruginosa wild type strain PAO1 (after 20 h growth). | ||
Isolation of sulforaphane from fresh broccoli
In order to assess the biological relevance of the observed QS-inhibitory effects we next set out to quantify the actual amount of sulforaphane in commercial broccoli. Over the last two decades many different methods for the extraction and quantification of natural sulforaphane from various sources have been published,4,30,31 and following some technical difficulties we decided to follow the protocol reported by Chiang et al.32 The average amount of sulforaphane from three separate broccoli extractions, as quantified by GC-MS, was 450 ± 26 μM (177 μg sulforaphane per gram broccoli), indicating that consumption of broccoli can, at least locally, lead to peak levels that are well above the observed concentrations needed to inhibit QS in P. aeruginosa. This, of course, does not mean that the broccoli plant produces sulforaphane and erucin in order to combat P. aeruginosa. In fact, this opportunistic pathogen is generally not a threat to broccoli plants. P. fluorescens, however, is a known pathogen of these plants, and its primary autoinducer is 3-hydroxyoctanoyl-HSL. Given the observed effective inhibition of QS in P. aeruginosa by sulforaphane and erucin, it is quite likely that these isothiocyanates are also potent QS inhibitors in P. fluorescens.33Conclusions
In summary, we investigated the effects of sulforaphane and erucin on quorum sensing and virulence in P. aeruginosa, and we examined the mechanism behind the strong inhibition of these important phenotypes. The combined reporter assays (in P. aeruginosa and in E. coli) strongly indicate that sulforaphane and erucin effectively bind LasR, resulting in inhibition of QS activation at concentrations that can be found in broccoli. We are currently further investigating the mechanism of action of these isothiocyanates in P. aeruginosa and P. fluorescens, including a hypothetical mechanistic relationship to other sulfur containing natural quorum sensing inhibitors, such as ajoene from garlic.22Acknowledgements
We thank professors E. P. Greenberg (Univ. of Washington), E. Banin (Bar-Ilan Univ.) and M. G. Surette (Univ. of Calgary) for generously providing bacterial strains. We also thank Ilana Berger for technical assistance. This research was supported by the European Research Council (Starting Grant 240356, M.M.M.), the Israel Science Foundation (Grant 749/09) and The Edmond J. Safra Center for the Design and Engineering of Functional Biopolymers in the Negev.Notes and references
- J. W. Fahey, A. T. Zalcmann and P. Talalay, Phytochemistry, 2001, 56, 5–51 CrossRef CAS.
- A. Steinbrecher, K. Nimptsch, A. Hüsing, S. Rohrmann and J. Linseisen, Int. J. Cancer, 2009, 125, 2179–2186 CrossRef CAS.
- L. Tang, G. Zirpoli, V. Jayaprakash, M. Reid, S. McCann, C. Nwogu, Y. Zhang, C. Ambrosone and K. Moysich, BMC Cancer, 2010, 10, 162 CrossRef.
- Y. Zhang, P. Talalay, C. G. Cho and G. H. Posner, Proc. Natl. Acad. Sci. U. S. A., 1992, 89, 2399–2403.
- T. K. Smith, E. K. Lund and I. T. Johnson, Carcinogenesis, 1998, 19, 267–273 CrossRef CAS.
- J. W. Fahey, X. Haristoy, P. M. Dolan, T. W. Kensler, I. Scholtus, K. K. Stephenson, P. Talalay and A. Lozniewski, Proc. Natl. Acad. Sci. U. S. A., 2002, 99, 7610–7615 CrossRef CAS.
- N. V. Matusheski, M. A. Wallig, J. A. Juvik, B. P. Klein, M. M. Kushad and E. H. Jeffery, J. Agric. Food Chem., 2001, 49, 1867–1872 CrossRef CAS.
- A. Aires, V. R. Mota, M. J. Saavedra, A. A. Monteiro, M. Simões, E. A. S. Rosa and R. N. Bennett, J. Appl. Microbiol., 2009, 106, 2096–2105 CrossRef CAS.
- G. R. Fenwick, R. K. Heaney, W. J. Mullin and C. H. VanEtten, CRC Crit. Rev. Food Sci. Nutr., 1982, 18, 123–201.
- P. Talalay, J. W. Fahey, W. D. Holtzclaw, T. Prestera and Y. Zhang, Toxicol. Lett., 1995, 82–83, 173–179 CrossRef CAS.
- E. Heiss, C. Herhaus, K. Klimo, H. Bartsch and C. Gerhauser, J. Biol. Chem., 2001, 276, 32008–32015 CrossRef CAS.
- G. H. Talbot, J. Bradley, J. E. Edwards, D. Gilbert, M. Scheld and J. G. Bartlett, Clin. Infect. Dis., 2006, 42, 657–668 CrossRef.
- G. D. Wright and A. D. Sutherland, Trends Mol. Med., 2007, 13, 260–267 CrossRef CAS.
- M. B. Miller and B. L. Bassler, Annu. Rev. Microbiol., 2001, 55, 165–199 CrossRef CAS.
- B. L. Bassler, Cell, 2002, 109, 421–424 CrossRef CAS.
- U. Müh, B. J. Hare, B. A. Duerkop, M. Schuster, B. L. Hanzelka, R. Heim, E. R. Olson and E. P. Greenberg, Proc. Natl. Acad. Sci. U. S. A., 2006, 103, 16948–16952 CrossRef.
- N. Amara, R. Mashiach, D. Amar, P. Krief, S. A. H. Spieser, M. J. Bottomley, A. Aharoni and M. M. Meijler, J. Am. Chem. Soc., 2009, 131, 10610–10619 CrossRef CAS.
- J. A. Gutierrez, T. Crowder, A. Rinaldo-Matthis, M.-C. Ho, S. C. Almo and V. L. Schramm, Nat. Chem. Biol., 2009, 5, 251–257 CrossRef CAS.
- M. Whiteley, K. M. Lee and E. P. Greenberg, Proc. Natl. Acad. Sci. U. S. A., 1999, 96, 13904–13909 CrossRef CAS.
- M. Hentzer, H. Wu, J. B. Andersen, K. Riedel, T. B. Rasmussen, N. Bagge, N. Kumar, M. A. Schembri, Z. Song, P. Kristoffersen, M. Manefield, J. W. Costerton, S. Molin, L. Eberl, P. Steinberg, S. Kjelleberg, N. Hoiby and M. Givskov, EMBO J., 2003, 22, 3803–3815 CrossRef CAS.
- T. H. Jakobsen, S. K. Bragason, R. K. Phipps, L. D. Christensen, M. van Gennip, M. Alhede, M. Skindersoe, T. O. Larsen, N. Hoiby, T. Bjarnsholt and M. Givskov, Appl. Environ. Microbiol., 2012, 78, 2410–2421 CrossRef CAS.
- T. H. Jakobsen, M. van Gennip, R. K. Phipps, M. S. Shanmugham, L. D. Christensen, M. Alhede, M. E. Skindersoe, T. B. Rasmussen, K. Friedrich, F. Uthe, P. O. Jensen, C. Moser, K. F. Nielsen, L. Eberl, T. O. Larsen, D. Tanner, N. Hoiby, T. Bjarnsholt and M. Givskov, Antimicrob. Agents Chemother., 2012, 56, 2314–2325 CrossRef CAS.
- Y. Zhang and L. Tang, Acta Pharmacol. Sin., 2007, 28, 1343–1354 Search PubMed.
- T. J. Ding, L. Zhou and X. P. Cao, Chin. Chem. Lett., 2006, 17, 2.
- M. Vermeulen, B. Zwanenburg, G. J. F. Chittenden and H. Verhagen, Eur. J. Med. Chem., 2003, 38, 729–737 CrossRef CAS.
- J. K. Whitesell and M.-S. Wong, J. Org. Chem., 1994, 59, 597–601 CrossRef CAS.
- H. Ganin, Y. Danin-Poleg, Y. Kashi and M. M. Meijler, ACS Chem. Biol., 2012, 7, 659–665 CrossRef CAS.
- L. Ye, A. T. Dinkova-Kostova, K. L. Wade, Y. Zhang, T. A. Shapiro and P. Talalay, Clin. Chim. Acta, 2002, 316, 43–53 CrossRef CAS.
- G. D. Geske, J. C. O'Neill, D. M. Miller, R. J. Wezeman, M. E. Mattmann, Q. Lin and H. E. Blackwell, ChemBioChem, 2008, 9, 389–400 CrossRef CAS.
- D. Bertelli, M. Plessi, D. Braghiroli and A. Monzani, Food Chem., 1998, 63, 417–421 CrossRef CAS.
- K. Nakagawa, T. Umeda, O. Higuchi, T. Tsuzuki, T. Suzuki and T. Miyazawa, J. Agric. Food Chem., 2006, 54, 2479–2483 CrossRef CAS.
- W. C. K. Chiang, D. J. Pusateri and R. E. A. Leitz, J. Agric. Food Chem., 1998, 46, 1018–1021 CrossRef CAS.
- X. Cui, R. Harling, P. Mutch and D. Darling, Eur. J. Plant Pathol., 2005, 111, 297–308 CrossRef CAS.
Footnote |
| † Electronic supplementary information (ESI) available. See DOI: 10.1039/c2md20196h |
| This journal is © The Royal Society of Chemistry 2013 |
