Rapid assembly of potent type II dehydroquinase inhibitorsvia “Click” chemistry†
Anh Thu
Tran
a,
Katie M.
Cergol
a,
Warwick J.
Britton
b,
Syed Ali
Imran Bokhari
c,
Musadiq
Ibrahim
c,
Adrian J.
Lapthorn
c and
Richard J.
Payne
*a
aSchool of Chemistry, The University of Sydney, Building F11, Sydney, Australia. E-mail: richard.payne@sydney.edu.au; Fax: +61 2 9351 3329; Tel: +61 2 9351 5877
bFaculty of Medicine, Blackburn Building, The University of Sydney, NSW 2006, Australia and Mycobacterial Research Program, Centenary Institute, NSW, Australia
cDepartment of Chemistry and Division of Biochemistry and Life Science, University of Glasgow, UK G12 8QQ
First published on 23rd August 2010
Abstract
The rapid synthesis of a library of potent type II dehydroquinase inhibitors is described. Inhibitors were prepared via a key quinate-derived ene-yne intermediate using Cu(I)-catalysed azide-alkynecycloaddition (CuAAC) chemistry with a variety of aryl- and heteroaryl-azides.
Introduction
The shikimate pathway is responsible for the conversion of erythrose-4-phosphate and phosphoenol pyruvate into chorismate over seven enzyme-catalysed steps.1–3 The end product of the pathway, chorismate, serves as the key branchpoint for the biosynthesis of a host of aromatic secondary metabolites in plants, bacteria, fungi and apicomplexan parasites in which the pathway is known to operate.2,4 These include folate, the aromatic amino acids tyrosine, tryptophan and phenylalanine, and essential vitamins including the folatecoenzymes, benzoid and naphthenoid quinones. The absence of the shikimate pathway in mammals, along with the essential role of this pathway in the viability of plants, bacteria, fungi and apicomplexan parasites,5,6 makes the pathway an excellent target for the development of broad spectrumherbicides, antibacterials, antifungals and antimalarial agents.2,7The third step of the shikimate pathway, the dehydration of 3-dehydroquinate (1) to 3-dehydroshikimate (2), is catalysed by the enzyme dehydroquinase (3-dehydroquinate dehydratase, EC 4.2.1.10). Interestingly, two structurally and mechanistically different enzymes, type I and type II, have evolved to carry out this transformation.8,9 Type I enzymes are dimeric proteins that catalyse the syn-elimination of waterviaSchiff base formation with a conserved active site lysine residue. In contrast, type II dehydroquinases are dodecameric proteins which catalyse the dehydration reactionvia an E1CB mechanism (Scheme 1).10,11 The dehydration is initiated by abstraction of the pro-S hydrogen by a conserved tyrosine (Tyr) residue to form an enol intermediate. Anti-elimination of water then occurs, facilitated by a water molecule and a conserved asparagine and histidine residue (Scheme 1).
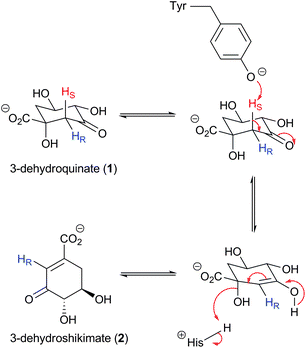 | ||
| Scheme 1 Proposed mechanism of the type II dehydroquinase-catalysed dehydration of 3-dehydroquinate (1) to 3-dehydroshikimate (2). | ||
In the past decade there has been significant interest in the design and synthesis of type II dehydroquinase inhibitors.12 This is due, in major part, to the operation of this enzyme in a number of pathogenic bacteria including Mycobacterium tuberculosis13 and Helicobacter pylori,14 the etiological agents of tuberculosis, and gastric ulcers and gastric cancer, respectively. It is therefore viewed as a potential target for the development of broad spectrumantibacterial agents.12 Previous work by Abell and coworkers15–17 and González-Bello and coworkers18–21 has revealed a number of key features deemed essential for selective and potent inhibition of the type II dehydroquinases. Specifically, both groups have reported that compounds possessing an anhydroquinate core with a terminal aromatic moiety extending from C-3 (with or without a bridging linker) are potent inhibitors of these enzymes (Fig. 1). Recent crystallographic evidence suggests that potent inhibition of the enzyme by these compounds is facilitated by a combination of the anhydroquinate core forming a number of hydrogen bonds with both active site residues and a conserved water molecule, and secondly the terminal aromatic moiety partaking in a π-stacking interaction (face to face or edge to face) with the conserved active site Tyr residue (which serves as the base in the proposed mechanism, Scheme 1).16,21
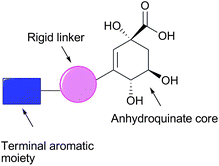 | ||
| Fig. 1 Schematic representation of potent type II dehydroquinase inhibitors previously reported.15–21 | ||
In our present investigation, we envisioned the use of reliable and robust chemistry to rapidly and efficiently assemble a library of inhibitors, which contain these key inhibitory features, from ene-yne intermediate 3 (Scheme 2). We initially proposed an inhibitor library bearing a triazole functionality as a rigid linker between the anhydroquinate core and a range of terminal aryl or heteroaryl moieties at C-3, which we proposed to install via a Cu(I)-catalysed alkyne-azidecycloaddition (CuAAC) reaction.22,23 In order to maximise diversity in our inhibitor library, we chose to incorporate terminal aryl rings with a variety of electronic properties (4a–4f, Scheme 2). Incorporation of a flexible methylene linker between the triazole and the electron-rich thiophene and furan rings in inhibitors5a and 5b was also proposed to probe the effect of conformational freedom on inhibition of the type II dehydroquinases (Scheme 2).
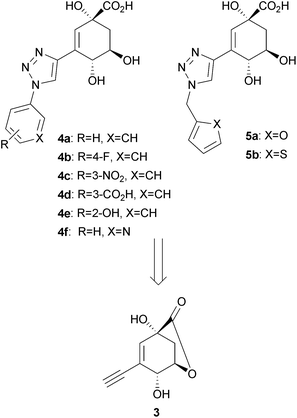 | ||
| Scheme 2 Retrosynthesis of triazole-based inhibitors4a–4f, 5a and 5b | ||
Results and discussion
Before embarking on the synthesis of the proposed inhibitor library, triazole-based inhibitors4a–4f, 5a and 5b were docked into the active site of the type II dehydroquinases from S. coelicolor and H. pylori (PDB code: 1GU1 and 2WKS respectively) using Glide® (Schrödinger Inc.).24 The majority of the inhibitors showed very similar binding modes (see ESI†). Representative dockings of compound 4a into the active sites of S. coelicolor and H. pylori type II dehydroquinases are shown in Fig. 2. The typical docking mode involved the anhydroquinate core binding into the active site where it forms a number of favourable hydrogen bonding interactions with active site residues and a conserved water molecule (see Fig. 2 and ESI†). In addition, the terminal aryl and heteroaryl moieties were positioned in close proximity to the essential Tyr residue (Tyr28 in S. coelicolor) in the enzyme and, as such, were predicted to participate in a π-stacking interaction with this conserved residue, consistent with our design strategy.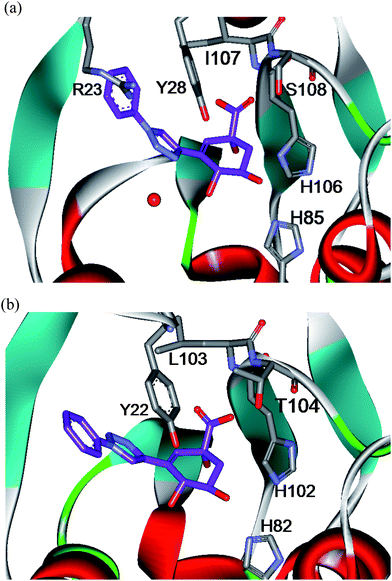 | ||
| Fig. 2 Docking of inhibitor4a into the active site of (a) S. coelicolor type II dehydroquinase and (b) H. pylori type II dehydroquinase. | ||
Having established the suitability of the proposed inhibitorsvia docking studies, we next embarked on the synthesis of key ene-yne intermediate 3 (Scheme 3). The synthesis began with 3-silylated lactone 6, obtained in 2 steps from commercially available (−)-quinic acid using an established literature procedure.25 Treatment of 6 with excess dimethoxymethane and phosphorus pentoxide gave the fully protected lactone7 in 78% yield. Desilylation at C-3 using tetrabutylammonium fluoride (TBAF) furnished alcohol 8 in 87% yield and was followed by oxidation with Dess–Martin periodinane (DMP) to afford the corresponding ketone9 in 90% yield. Ketone9 was subsequently converted to vinyl triflate10via deprotonation with potassium hexamethyldisilazane, followed by reaction of the resulting enolate with Comin's triflating reagent to afford 10 in 67% yield. Vinyl triflate10 was next subjected to palladium-catalysed Sonogashira cross-coupling conditions with TMS-acetylene to afford protected ene-yne11 in 75% yield. Subsequent silyl deprotection with TBAF proceeded smoothly to afford 12, which was treated with 90% trifluoroacetic acid (TFA) to give the key ene-yne intermediate 3 in 64% yield over two steps.
![Reagents and conditions: a) (OCH3)2CH2, P2O5, CH2Cl2, rt, 78%; b) TBAF, THF, 0 °C, 87%; c) DMP, CH2Cl2, rt, 90%; d) (i) KHMDS, DMF, PhCH3, −78 °C, (ii) Comin's reagent, −78 °C to rt, 67%; e) TMS-acetylene, [Pd(PPh3)4] (15 mol%), CuI (20 mol%), piperidine, THF, 40 °C, 75%; f) Cat. TBAF, THF, rt.; g) 90% aq. TFA, rt, 64% over 2 steps.](/image/article/2010/MD/c0md00097c/c0md00097c-s3.gif) | ||
| Scheme 3 Reagents and conditions: a) (OCH3)2CH2, P2O5, CH2Cl2, rt, 78%; b) TBAF, THF, 0 °C, 87%; c) DMP, CH2Cl2, rt, 90%; d) (i) KHMDS, DMF, PhCH3, −78 °C, (ii) Comin's reagent, −78 °C to rt, 67%; e) TMS-acetylene, [Pd(PPh3)4] (15 mol%), CuI (20 mol%), piperidine, THF, 40 °C, 75%; f) Cat. TBAF, THF, rt.; g) 90% aq. TFA, rt, 64% over 2 steps. | ||
With the key ene-yne intermediate 3 in hand, we next focussed on the preparation of the proposed triazole-based inhibitors. To this end, ene-yne3 was reacted with a library of aryl- and heteroaryl-azides (see ESI†) using the robust CuAAC reaction conditions22 (0.1 eq. CuSO4, 1 eq. sodium ascorbate, 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 v/v water/tert-butanol) to furnish the corresponding triazole lactones (Scheme 4). Saponification of the resulting lactones followed by acidification afforded the desired triazole-based inhibitors4a–4f, 5a and 5b in moderate to good yields (27–69%) over the two steps.
1 v/v water/tert-butanol) to furnish the corresponding triazole lactones (Scheme 4). Saponification of the resulting lactones followed by acidification afforded the desired triazole-based inhibitors4a–4f, 5a and 5b in moderate to good yields (27–69%) over the two steps.
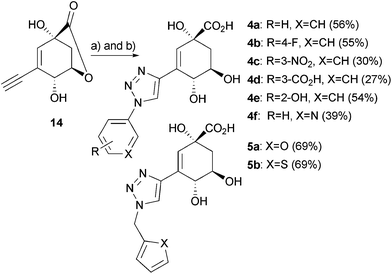 | ||
Scheme 4 Reagents and conditions: a) sodium ascorbate (1 eq.), CuSO4·5H2O (0.1 eq.), R-N3, tert-butanol/water (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 v/v), rt; b) (i) LiOH, THF, rt, (ii) Amberlite IR-120 (H+ form). 1 v/v), rt; b) (i) LiOH, THF, rt, (ii) Amberlite IR-120 (H+ form). | ||
The biological activity of triazole-based inhibitors4a–4f, 5a and 5b against type II dehydroquinase from three different organisms, namely Streptomyces coelicolor, H. pylori and M. tuberculosis, was determined using a UV spectrophotometric assay (see ESI†).17 This entailed measuring the initial rate of product (3-dehydroshikimate, 2) formation from 3-dehydroquinate (1)26 by detection of the enone-carboxylatechromophore at λ = 234 nm. All the compounds proved to be competitive, reversible inhibitors of the enzyme as shown by a least squares fitting to a competitive model.
Gratifyingly, the majority of the triazole-based compounds proved to be potent inhibitors of all three of the type II deydroquinases, with inhibition constants in the low- to mid-nanomolar range (Table 1). Inhibitors4a and 4f, bearing terminal phenyl and 3-pyridyl rings respectively, proved to be particularly potent against all three enzymes. Both compounds exhibited an inhibition constant of 5.7 nM against S. coelicolor type II dehydroquinase and, as such, represent two of the most potent inhibitors of any type II dehydroquinase synthesised to date. These two compounds also exhibited potent inhibition of H. pylori type II dehydroquinase (4a: KI = 225 nM, 4f: KI = 375 nM) and of the M. tuberculosisenzyme (4a: KI = 39 nM, 4f: KI = 136 nM). Inspection of the docking results reveals a number of key features which are postulated to contribute to the potency of these compounds. Firstly, the anhydroquinate core appeared to bind to the active site of the enzymes in a very similar mode to that observed in reported co-crystal structures of other inhibitors.11,16,21 Specifically, the C-1carboxylate is held in place by two backbone amides of the protein chain, and the C-1hydroxyl by the side chain imidazole of a conserved histidine residue. There also appears to be a hydrogen bonding interaction between the C-5hydroxylgroup and the side chain of another histidine residue. The triazole-linked aryl rings appear to be involved in a π-stacking interaction with the conserved Tyr residue on the flexible loop and in the S. coelicolor type II dehydroquinase dockings there is a potential cation-π interaction with an arginine residue. A positive entropic effect resulting from the displacement of water molecules from the region under the flexible loop into the bulk solvent is also proposed to contribute to the potency of these compounds. It is also important to note that 4a, which has a terminal aromatic moiety directly attached to the triazole linker, is significantly more potent compared to an analogous compound bearing a flexible methylene linker between the phenyl ring and the triazole reported previously (KI = 30 nM against S. coelicolor type II dehydroquinase and KI = 3.75 μM against M. tuberculosis type II dehydroquinase).19 This suggests that rigid positioning of the terminal aromatic moiety may be an important feature for a favourable π-stacking interaction with the conserved Tyr residue, thus leading to more potent inhibition. This hypothesis is supported by the significantly weaker inhibition exhibited by 5a and 5b, containing flexible methylene linked furan and thiophenegroups respectively. Whilst these compounds still exhibited nanomolar inhibition constants against S. coelicolor type II dehydroquinase, they were significantly less potent against the H. pylori and M. tuberculosis enzymes.
| Inhibitor | S. coelicolor type II DHQaseKI/nM | H . pylori type II DHQaseKI/nM | M. tuberculosis type II DHQaseKI/nM |
|---|---|---|---|
| a Kinetic parameters of the type II dehydroquinase enzymes: S. coelicolor: KM = 679 ± 68 μM, kcat = 84 s−1, 50 mM Tris.HCl, pH 7.0, 25 °C; H. pylori: KM = 228 ± 33 μM, kcat = 1.4 s−1, 50 mM Tris·HOAc, pH 7.0, 25 °C; M. tuberculosis: KM = 52 ± 8 μM, kcat = 6.3 s−1, 50 mM Tris.HCl, pH 7.0, 25 °C. | |||
| 4a | 5.7 ± 0.9 | 225 ± 51 | 39 ± 5 |
| 4b | 29 ± 3 | 561 ± 75 | 154 ± 16 |
| 4c | 78 ± 8 | 538 ± 83 | 230 ± 20 |
| 4d | 1720 ± 120 | 5780 ± 540 | 3126 ± 315 |
| 4e | 48 ± 4 | 157 ± 14 | 330 ± 24 |
| 4f | 5.7 ± 0.8 | 375 ± 53 | 136 ± 12 |
| 5a | 217 ± 17 | 2200 ± 300 | 2122 ± 193 |
| 5b | 73 ± 10 | 925 ± 120 | 1579 ± 149 |
Incorporation of a terminal p-fluorophenyl substituent in 4b and an o-hydroxyphenyl moiety in 4e also led to potent inhibition of all three type II dehydroquinases (KI = 29–330 nM). However, these compounds were slightly less potent against all three enzymes when compared to 4a, suggesting that small changes in the electronics of the aryl moiety can modulate potency of these compounds.
As part of our original design strategy we anticipated that the electron-withdrawing m-nitro and m-carboxylate substituents, in 4c and 4d respectively, would result in more favourable π–π interactions with the conserved active site Tyr residue of the type II dehydroquinases, leading to more potent inhibitors. Unfortunately this was not the case as 4c proved to be a less potent inhibitor than compound 4a which bears only a phenyl substituent. Furthermore, 4d exhibited the weakest inhibition of any compound in this series, with inhibition constants in the low micromolar range against all three enzymes. Inspection of the S. coelicolor type II dehydroquinase docking results allows for some speculative insight into the drop in potency of 4d. In contrast to every other member of the triazole-based inhibitor library, the anhydroquinate core in compound 4d docked into the active site in a different binding mode (see ESI†). In this mode 4d is no longer able to make a network of hydrogen bonding interactions with the active site of the enzyme which may explain the poor inhibitory properties of this compound.
Conclusions
In summary, we have designed and synthesised a novel series of triazole-based type II dehydroquinase inhibitors using CuAAC reactions between a key quinate-derived ene-yne and a variety of aryl- and heteroaryl-azides. These were screened against S. coelicolor, H. pylori and M. tuberculosis type II dehydroquinases. The majority of the compounds proved to be potent inhibitors of all three type II dehydroquinases with inhibition constants in the low to mid-nanomolar range. In particular, 4a and 4f bearing terminal phenyl and 3-pyridyl rings directly linked to the triazole moiety, represent some of the most potent inhibitors of the type II dehydroquinases from S.coelicolor, H.pylori and M. tuberculosis ever synthesised. Future work in our laboratories will involve co-crystallisation of the inhibitors in this series with the type II dehydroquinases and evaluation of their antibacterial activity against M. tuberculosis and H. pylori in vitro.Acknowledgements
We would like to thank the Australian National Health and Medical Research Council for funding (Project grant 632769) and the Australian Postgraduate Award for PhD funding (AT). We would also like to thank Associate Professor Dai Hibbs for assistance with docking studies and Dr Margaret Sunde and Dr David Gell from the School of Molecular Bioscience, The University of Sydney for use of a UV spectrophotometer. Finally, we would like to acknowledge the technical support of Dr Kelvin Picker, Dr Ian Luck and Dr Keith Fisher.Notes and references
- E. Haslam, Shikimic Acid: Metabolism and Metabolites, John Wiley & Sons Ltd, Chicester, UK, 1993, pp. 1–400 Search PubMed.
- C. Abell, in Comprehensive Natural Products Chemistry, ed. U. Sankawa, Elsevier, Amsterdam, 1999, pp. 573–607 Search PubMed.
- R. Bentley, Crit. Rev. Biochem. Mol. Biol., 1990, 25, 307–384 CrossRef CAS.
- U. Weiss and J. M. Edwards, in The Biosynthesis of Aromatic Compounds, Wiley: New York, 1980, pp. 1–728 Search PubMed.
- F. Roberts, C. W. Roberts, J. J. Johnson, D. E. Kyle, T. Krell, J. R. Coggins, G. H. Coombs, W. K. Milhous, S. Tzipori, D. J. P. Ferguson, D. Chakrabarti and R. McLeod, Nature, 1998, 393, 358–805 CrossRef.
- P. J. Keeling, J. D. Palmer, R. G. K. Donald, D. S. Roos, R. F. Waller and G. I. McFadden, Nature, 1999, 397, 219–220 CrossRef CAS.
- G. A. McConkey, Antimicrob. Agents Chemother., 1999, 43, 175–177 CAS.
- C. Kleanthous, R. Deka, K. Davis, S. M. Kelly, A. Cooper, S. E. Harding, N. C. Price, A. R. Hawkins and J. R. Coggins, Biochem. J., 1992, 282, 687–695 CAS.
- J. Harris, C. Kleanthous, J. R. Coggins, A. R. Hawkins and C. Abell, J. Chem. Soc., Chem. Commun., 1993, 1080–1081 RSC.
- J. M. Harris, C. González-Bello, C. Kleanthous, A. R. Hawkins, J. R. Coggins and C. Abell, Biochem. J., 1996, 319, 333–336 CAS.
- A. W. Roszak, D. A. Robinson, T. Krell, I. S. Hunter, M. Fredrickson, C. Abell, J. R. Coggins and A. J. Lapthorn, Structure, 2002, 10, 493–503 CrossRef.
- C. González-Bello and L. Castedo, Med. Res. Rev., 2007, 27, 177–208 CrossRef CAS.
- T. Garbe, S. Selvos, A. Hawkins, G. Dimitriadis, D. Young, G. Dougan and I. Charles, MGG, Mol. Gen. Genet., 1991, 228 Search PubMed.
- J. R. Bottomley, C. L. Clayton, P. A. Chalk and C. Kleanthous, Biochem. J., 1996, 319, 559–565 CAS.
- M. D. Toscano, R. J. Payne, A. Chiba, O. Kerbarh and C. Abell, ChemMedChem, 2007, 2, 101–112 CrossRef CAS.
- R. J. Payne, A. Riboldi-Tunnicliffe, O. Kerbarh, A. D. Abell, A. J. Lapthorn and C. Abell, ChemMedChem, 2007, 2, 1010–1013 CrossRef CAS.
- R. J. Payne, F. Peyrot, O. Kerbarh, A. D. Abell and C. Abell, ChemMedChem, 2007, 2, 1015–1029 CrossRef CAS.
- C. Sánchez-Sixto, V. F. V. Prazeres, L. Castedo, H. Lamb, A. R. Hawkins and C. González-Bello, J. Med. Chem., 2005, 48, 4871–4881 CrossRef CAS.
- V. F. V. Prazeres, C. Sánchez-Sixto, L. Castedo, H. Lamb, A. R. Hawkins, A. Riboldi-Tunnicliffe, J. R. Coggins, A. J. Lapthorn and C. González-Bello, ChemMedChem, 2007, 2, 194–207 CrossRef CAS.
- C. Sánchez-Sixto, V. F. V. Prazeres, L. Castedo, S. W. Suh, H. Lamb, A. R. Hawkins, F. J. Cañada, J. Jiménez-Barbero and C. González-Bello, ChemMedChem, 2008, 3, 756–770 CrossRef CAS.
- V. F. V. Prazeres, L. Tizón, J. M. Otero, P. Guardado-Calvo, A. L. Llamas-Saiz, M. J. van Raaij, L. Castedo, H. Lamb, A. R. Hawkins and C. González-Bello, J. Med. Chem., 2010, 53, 191–200 CrossRef CAS.
- V. V. Rostovtsev, L. G. Green, V. V. Fokin and K. B. Sharpless, Angew. Chem., Int. Ed., 2002, 41, 2596–2599 CrossRef CAS.
- C. W. Tornøe, C. Christensen and M. Meldal, J. Org. Chem., 2002, 67, 3057–3064 CrossRef CAS.
- R. A. Friesner, R. B. Murphy, M. P. Repasky, L. L. Frye, J. R. Greenwood, T. A. Halgren, P. C. Sanschagrin and D. T. Mainz, J. Med. Chem., 2006, 49, 6177–6196 CrossRef CAS.
- L. Sánchez-Abella, S. Fernández, N. Armesto, M. Ferrero and V. Gotor, J. Org. Chem., 2006, 71, 5396–5399 CrossRef CAS.
- C. Le Sann, C. Abell and A. D. Abell, Synth. Commun., 2003, 33, 527–533 CrossRef CAS.
Footnote |
| † Electronic supplementary information (ESI) available: Experimental procedures and full characterisation for all new compounds. 1H and 13C NMR data for all new compounds and docking results. See DOI: 10.1039/c0md00097c |
| This journal is © The Royal Society of Chemistry 2010 |
