Locating the Band III protein in quasi-native cell membranes
Yuping
Shan†
ad,
Zhiyong
Wang†
b,
Xian
Hao
ad,
Xin
Shang
a,
Mingjun
Cai
a,
Junguang
Jiang
a,
Xuexun
Fang
b,
Hongda
Wang
*a and
Zhiyong
Tang
*c
aState Key Laboratory of Electroanalytical Chemistry, Changchun Institute of Applied Chemistry, Chinese Academy of Sciences, Changchun, Jilin 130022, P.R. China. E-mail: hdwang@ciac.jl.cn
bKey Laboratory for Molecular Enzymology & Engineering, the Ministry of Education, Jilin University, Changchun, Jilin 130021, P.R.China
cNational Center for Nanoscience and Technology, Beijing, 100190, P.R.China. E-mail: zytang@nanoctr.cn
dGraduate School of Chinese Academy of Sciences, Beijing, 100049, P.R. China
First published on 7th June 2010
Abstract
Band III is a key protein for the structure and function of red blood cell membranes. To date, the distribution and morphology of Band III in cell membranes is still unclear because of limited approaches. We applied Topography and RECognition imaging microscopy (TREC), which extends the application of atomic force microscopy (AFM) to recognize a single molecule in a biological complex, to visually locate a single Band III protein in quasi-native cell membranes by anti-Band III-functionalized AFM tips under physiological conditions. The Band III proteins are well distributed in the inner leaflet of cell membranes. The height of the whole Band III protein in cell membranes is in the range of 9–13 nm. The unbinding force between Band III in the membrane and anti-Band III on the AFM tip is about 70 pN with the loading rate at 40 nN/s. Our result is significant in revealing the location and morphology of Band III in the inner cell membrane at the molecular level.
Introduction
Band III protein, one of the most abundant proteins in human red blood cell (hRBC) membranes, has been widely studied for decades due to its important role in cell membranes.1,2 Band III is the major integral membrane protein that facilitates the one-for-one exchange of Cl− and HCO3− in hRBC,3 so that Band III serves as a model for investigating the structure and function of integral membrane proteins.4 Furthermore, researchers revealed that Band III is related to control of the cell shape and support of the membrane skeleton.5,6 The crystal structures of membrane domain and cytoplasmic domain of Band III have been reported by X-ray diffraction analysis,7–9 respectively. However, the distribution of Band III in cell membranes and the relationship between Band III and other proteins that may be significant to reveal how the membrane skeleton is built, are still an unknown topic because of limited approaches. Fluorescent microscopy has provided the direct evidence for observing objects in cell membranes. However, its relatively low lateral resolution (about 200 nm for confocal fluorescence microscopy) limits it to mapping a single protein at a few nanometer resolution.10 Although the electron microscope shows a high resolution, it could not be used to map a single molecule under in situ conditions. Therefore, a higher resolution and in situ technique is required to locate the Band III protein in cell membranes.Atomic Force Microscope (AFM) is well known as a very useful tool to observe single molecules due to its high-resolution and ability of imaging in variable environments.11,12 Especially, Topography and RECognition imaging microscopy (TREC) with several nanometers resolution has been recently developed to expand AFM's application for specifically recognizing biological molecules.13–15 TREC scans a sample with an antibody-functionalized AFM tip. A topographic image is acquired simultaneously with a recognition image that locates the sites of antigen-antibody binding events, leading to accurately locate the specific individual molecules within the topographic image. The recognition signal induced by the interaction between an antibody on the AFM tip and an antigen on the surface is highly specific and efficient. TREC has been successfully used in the recognition of single molecules in protein complexes and fixed cell membranes.13,15–17 It is known that the inner leaflet of cell membranes are covered by dense and different types of proteins, and in our recent study the positions of the ATPases in the inner leaflet of cell membranes were successfully explored by AFM and TREC.18 Herein, we extend the application of TREC to “observe” and “map” the Band III protein in quasi-native cell membranes under physiological conditions. This study would be significant to reveal in situ the location of Band III in cell membranes and their relationship with other membrane proteins at the molecular level.
Experimental
APTES functionalized mica
A desiccator was purged with argon for 2 min and 30 μL APTES (3-aminopropyltriethoxysilane, 99%, Sigma-Aldrich, St. Louis, MO) was placed into a small container at the bottom of the desiccator.19 10 μL N,N-diisopropylethylamine (99%, distilled, Sigma-Aldrich) was placed into another small container, and the desiccator was purged with argon for a further 2 min. Mica sheets were stripped on one side until smooth and immediately placed into the desiccator. The desiccator was purged for another 3 min and then sealed off, leaving the mica exposed to APTES vapor for 1 h. After this exposure, the APTES was removed, the desiccator purged, and the treated mica (AP-mica) stored in the sealed desiccator until needed.Preparing cell membranes
To prepare the hRBC sample, two drops of fresh human blood from a donor were washed five times in 150 mM PBS buffer (pH 7.5). Subsequently, the diluted hRBC was deposited onto AP-mica for 15 min. With the purpose of obtaining the inner membrane, hRBCs attached to AP-mica were sheared open by a fast stream of 150 mM PBS as described.20 Lastly, the cell membranes were incubated in 0.3 mM PBS at 37 °C for 1 h to remove the skeleton proteins. The prepared cell membranes were imaged in the PBS buffer (pH 7.5) immediately.Conjugation of antibody and ligand to AFM tips
Conjugation of the anti-Band III to AFM-tip (Microlever, Veeco, Santa Barbara, CA, coated for MacMode AFM by Agilent Technologies, Chandler, AZ) by a flexible PEG [poly(ethylene glycol)]-crosslinker (MAL-PEG2000-NHS, JenKem Technology Co., Ltd, Beijing) was performed as described in ref. 13. In brief, anti-Band III (Abcam, catalog no. ab55830) were reacted with N-succinimidyl 3-(acetylthio) propionate (SATP, Sigma inc.) and purified in a PD-10 column (Amersham Pharmacia Biotech). The cantilevers were cleaned in a UV cleaner, vapor treated with APTES and reacted with PEG crosslinker using triethylamine and CHCl3. The SATP-labeled antibodies were then bound to the PEG crosslinkers with NH2OH (Sigma) in a NaCl/phosphate buffer. The tips were then rinsed in PBS buffer and stored at 4 °C until use.AFM recognition imaging
Magnetically coated cantilevers were driven by a small solenoid using a MacMode dynamic 5500 AFM (Agilent Technologies, Chandler, AZ). The tip was oscillated by an alternating magnetic field while being scanned across the surface. Topography and recognition images of hRBC membrane were acquired simultaneously with MacMode AFM at an amplitude setting between 2.0 and 2.5 volts (equivalent to 6–8 nm). The recognition imaging was blocked by injecting 100 µl 1 mg/ml immunogen peptide (CEQGDGGTEGHSPSGILEKI) solution into the AFM liquid cell.Data analysis
Particles were measured using PicoScan 5.3.3 software (Agilent Technologies, Chandler AZ). A maximum height was taken as the peak height relative to the local background. Recognition imaging was analyzed by the software – Recognition Imaging Analysis.21 This program compiles histograms of the pixel intensity distribution from background regions containing no features, and compares them to regions containing visible recognition spots. Recognition events give rise to a second peak in the intensity distribution and these were clearly separated from the background by selecting a cut-off of 75% of the background intensity (recognition spots correspond to a decrease in the signal).Fluorescence image
The mercaptopropionic acid (MPA) capped CdTe nanoparticles were prepared by following the recipe reported in the literature.22 The prepared CdTe nanoparticles have a maximum photoluminescence peak at 640 nm, and average diameters of 4 nm. Conjugation of MPA-CdTe with anti-Band III was prepared as follows: 10 μL of 1-ethyl-3-(3-dimethylaminopropyl) carbodiimide hydrochloride (EDAC) (10 mM) solution and 10 μL of N-hydroxysuccinimide (NHS) (10 mM) solution were added to 100 μL of MPA-capped CdTe (2.98μM) PBS buffer solution. Then an appropriate amount of anti-Band III was added to the reaction mixture. The reaction solution was incubated at room temperature overnight with continuous gentle stirring. The anti-Band III–CdTe conjugation was purified through a PD SpinTrap G-25 filtration column (GE Healthcare) equilibrated in PBS, and stored at 4 °C.To obtain the inner membrane, hRBC attached to AP-mica were sheared open by a fast stream of low salt PBS as described.20 5 μL of anti-Band III-CdTe conjugate solution was added onto the inner membrane for 1 h and washed with PBS buffer 4 times before imaging. Imaging was done on a Leica TCS SP2 Laser scanning confocal microscope.
Results and discussion
In this communication, we used hRBC for the Band III protein study. The hRBCs have the merits of simplification and capacity of abundant Band III.3 To keep the inner cell membrane at near-native conditions,20 the hRBCs were immobilized on 3-aminopropyltriethoxysilane-modified mica (AP-mica) substrates and sheared open by a stream of phosphate buffer solution (150 mM, pH 7.5) without further fixation. The skeleton proteins, such as spectrin, have been mostly washed away by 0.3 mM PBS buffer as described.20 As shown in Fig. 1A, a topographic AFM image of inside-out cell membranes covered by dense particles – membrane proteins were observed. The membrane patches are about 6–8 μm, which are identical to the size of single hRBC.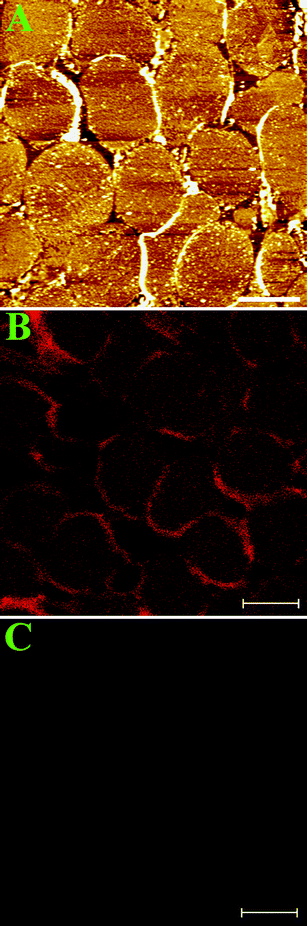 | ||
| Fig. 1 (A) AFM image of inside-out cell membranes at low resolution. Scale bar is 6 μm. (B) Fluorescent image of inside-out hRBC membranes (red patches) labeled with anti-Band III-QDs. The scale bar is 6 μm. (C) Control experiment for the fluorescent image in Figure 1B. The membrane patches were incubated with QDs only. The scale bar is 6 μm. | ||
To verify the existence of Band III protein in cell membranes, we incubated the cell membranes with anti-Band III-QD (quantum dots) and imaged them using a confocal laser scanning fluorescence microscope. Red patches in Fig. 1B indicate that there are abundant Band III proteins in cell membranes. The control experiment in Fig. 1C demonstrates that the non-specific signal from QDs on cell membranes is very low.
In order to map single Band III protein in the inner cell membrane, we functionalized anti-Band III onto the AFM tip and acquired topographic image and recognition image simultaneously (illustrated in Fig. 2A). Fig. 2B shows the topographic image of the inner membrane, in which the proteins are observed clearly. The corresponding recognition image is shown in Fig. 2C. The dark spots, spreading in the inner membrane with few aggregations, represent the protein sites recognized by the anti-Band III. A magnifying typical single recognition site is shown in Fig. 2E. The cross section analysis indicates that the recognition signal dip of tip amplitude is about 3 nm (Fig. 2F). To test the specificity of recognition signal, a blocking reagent (immunogenic peptides) was flowed into the liquid cell during imaging. After the addition of the blocking reagent, the recognition signals in Fig. 2C were abolished by an excess of free peptides in solution (Fig. 2D), demonstrating that the recognition signals came from specific interactions between the anti-Band III on the tip and Band III in the inner membrane. Fig. 2G shows one blocked area corresponding to Fig. 2E. The cross section analysis (Fig. 2H) indicates that the recognition signal was mostly abolished (<0.7 nm).
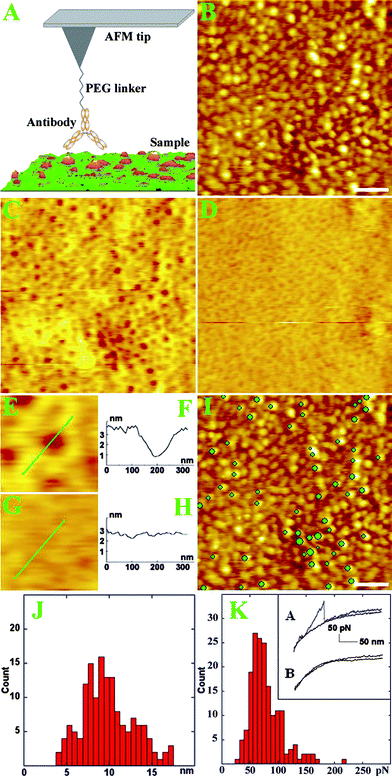 | ||
| Fig. 2 (A) An AFM tip functionalized with an antibody by a PEG linker scans the complex sample. The antibody temporally binds the antigen on the surface during scanning. (B) Topographic image of the inner membrane scanned by anti-Band III AFM tip. Scale bar is 500 nm. (C) Corresponding recognition image by anti-Band III AFM tip. Brown dots indicate the recognized Band III sites. (D) Recognition image of Band III proteins after blocking the recognition signal by free peptides. (E) Magnifying recognition image from Figure 2C. (F) Cross section analysis along the line in Figure 2E. (G) Magnifying recognition image from Figure 2D. (H) Cross section analysis along the line in Figure 2G. (I) Topographic image of the inner cell membrane with recognition spots superimposed by overlaying them (green dots) on the top of the topographic images. Scale bar is 500 nm. (J) The height distribution of proteins in the inner membrane from Figure 2B. (K) Distribution of unbinding force between the Band III in the inner membrane and the anti-Band III on the AFM tip. Inset A shows a typical approach and retraction force curve. Inset B shows a force curve after the addition of the blocking reagent. | ||
To obtain the single molecule location of Band III in the membrane proteins, we marked the recognition signal in recognition image, and superimposed the recognition spot onto the topographic image (Fig. 2I). The green dots on the topographic image represent the Band III sites recognized by the anti-Band III AFM tip. This map indicates that Band III proteins are well distributed in the membrane. Band III is presumed to anchor skeleton proteins (e.g. spectrin) in cell membranes.23 The end-to-end of the spectrin molecule in the skeleton meshwork is about 200 nm as reported.24 We measured that the distances between two adjacent Band III are in the range of 200–600 nm, which is consistent with the skeleton meshwork model.
We measured the size of proteins in the inner membrane. These proteins occupied a broad height distribution between 4 and 17 nm (shown in Fig. 2J), which is supported by the mess spectroscopy analysis that stated there were more than 100 types of proteins in RBC cell membranes.25 There might be some larger particles compared with the previous study because of the difference in sample preparation.18 The height distribution of the membrane proteins is reliable due to less AFM tip effect on the height measurement so that we could compare these proteins by the height measurement. On the basis of X-ray diffraction analysis, the height of the membrane part of Band III is about 8.0 nm,8 while the height of the cytoplasmic domain of Band III is about 4.5 nm.9 Therefore, we estimate that the height of a whole Band III protein in the cell membranes could be about 12.5 nm. From the height measurement in Fig. 2I, we noticed that the heights of particles recognized by the anti-Band III tip are mainly in the range of 9–13 nm. These results further testify that the particles recognized by anti-Band III (spots in Fig. 2C) were the Band III proteins in cell membranes. After counting the green spots on the particles, we obtained about 16% particles recognized by anti-Band III. This value is roughly consistent with the density of Band III in cell membranes as previously reported1,26 and also reasonable because the Band III is a type of main protein for cell functions. We assumed that most of the Band III proteins might be distributed in cell membranes randomly and solely according to the size measurement and distribution percentage; meanwhile, there may be also some aggregations that could be relative to the associated proteins around Band III.
To test the specific interaction and measure the binding force between Band III in the membrane and anti-Band III on the AFM tip, the force spectroscopy was accomplished by engaging the antibody-functionalized tip on the inner membrane. A typical force unbinding event on the cell membranes is shown in Fig. 2K (inset A). The histogram in Fig. 2K includes 250 unbinding force curves. The major peak is about 70 pN with the apparent loading rate at 40 nN/s (cantilever spring constant multiplied by retraction velocity18), which is much larger than the value from non-specific unbinding events (less than 20 pN).13 The inset B in Fig. 2K shows that the unbinding force was blocked after the addition of the blocking reagent. These results further demonstrate that the recognition between anti-Band III on the tip and the Band III in the membrane is specific.
Conclusions
In summary, we have located the Band III in native cell membranes by TREC at the single molecule level. Band III proteins were well distributed in the inner membrane. The height of the whole Band III protein in the inner membrane is in the range of 9–13 nm. The unbinding force between Band III in the membrane and anti-Band III on the tip is about 70 pN with the loading rate at 40 nN/s. Our results are significant in revealing the location and morphology of Band III in the inner leaflet of hRBC membranes. We expect that TREC could open a novel way in studying plasma membranes because of its direct observation and high resolution. The next challenging task will be to locate two types of membrane proteins by TREC simultaneously.Acknowledgements
This work was supported by National Natural Science Foundation of China (NSFC, Grant no. 20975098)References
- L. K. Drickamer, J. Biol. Chem., 1976, 251, 5115–5123 CAS.
- D. G. Williams, R. E. Jenkins and M. J. A. Tanner, Biochem. J., 1979, 181, 477–493 CAS.
- D. Jay and L. Cantley, Annu. Rev. Biochem., 1986, 55, 511–538 CrossRef CAS.
- J. D. Groves, P. Falson, M. leMaire and M. J. A. Tanner, Proc. Natl. Acad. Sci. U. S. A., 1996, 93, 12245–12250 CrossRef CAS.
- D. G. Jay, Cell, 1996, 86, 853–854 CrossRef CAS.
- N. Mohandas and P. G. Gallagher, Blood, 2008, 112, 3939–3948 CrossRef CAS.
- D. N. Wang, W. Kuhlbrandt, V. E. Sarabia and R. A. F. Reithmeier, EMBO J., 1993, 12, 2233–2239 CAS.
- D. N. Wang, V. E. Sarabia, R. A. F. Reithmeier and W. Kuhlbrandt, EMBO J., 1994, 13, 3230–3235 CAS.
- D. C. Zhang, A. Kiyatkin, J. T. Bolin and P. S. Low, Blood, 2000, 96, 2925–2933 CAS.
- G. J. Schutz, G. Kada, V. P. Pastushenko and H. Schindler, EMBO J., 2000, 19, 892–901 CrossRef CAS.
- H. Schillers, Pfluegers Arch., 2008, 456, 163–177 CrossRef CAS.
- W. H. Han, S. M. Lindsay, M. Dlakic and R. E. Harington, Nature, 1997, 386, 563–563 CrossRef CAS.
- C. Stroh, H. Wang, R. Bash, B. Ashcroft, J. Nelson, H. Gruber, D. Lohr, S. M. Lindsay and P. Hinterdorfer, Proc. Natl. Acad. Sci. U. S. A., 2004, 101, 12503–12507 CrossRef CAS.
- P. Hinterdorfer and Y. F. Dufrene, Nat. Methods, 2006, 3, 347–355 CrossRef CAS.
- H. Wang, R. Bash and D. Lohr, Anal. Biochem., 2007, 361, 273–279 CrossRef CAS.
- L. A. Chtcheglova, F. Atalar, U. Ozbek, L. Wildling, A. Ebner and P. Hinterdorfer, Pfluegers Arch., 2008, 456, 247–254 CrossRef CAS.
- A. Ebner, D. Nikova, T. Lange, J. Haberle, S. Falk, A. Dubbers, R. Bruns, P. Hinterdorfer, H. Oberleithner and H. Schillers, Nanotechnology, 2008, 19 Search PubMed.
- J. Jiang, X. Hao, M. Cai, Y. Shan, X. Shang, Z. Tang and H. Wang, Nano Lett., 2009, 9, 4489–4493 CrossRef CAS.
- Y. Lyubchenko, L. Shlyakhtenko, R. Harrington, P. Oden and S. Lindsay, Proc. Natl. Acad. Sci. U. S. A., 1993, 90, 2137–2140 CrossRef CAS.
- T. Lange, P. Jungmann, J. Haberle, S. Falk, A. Duebbers, R. Bruns, A. Ebner, P. Hinterdorfer, H. Oberleithner and H. Schillers, Mol. Membr. Biol., 2006, 23, 317–323 CrossRef CAS.
- L. Lin, H. D. Wang, Y. Liu, H. Yan and S. Lindsay, Biophys. J., 2006, 90, 4236–4238 CrossRef CAS.
- N. Gaponik, D. V. Talapin, A. L. Rogach, K. Hoppe, E. V. Shevchenko, A. Kornowski, A. Eychmuller and H. Weller, J. Phys. Chem. B, 2002, 106, 7177–7185 CrossRef CAS.
- A. Elgsaeter, B. T. Stokke, A. Mikkelsen and D. Branton, Science, 1986, 234, 1217–1223 CAS.
- T. J. Byers and D. Branton, Proc. Natl. Acad. Sci. U. S. A., 1985, 82, 6153–6157 CrossRef CAS.
- E. M. Pasini, M. Kirkegaard, D. Salerno, P. Mortensen, M. Mann and A. W. Thomas, Mol. Cell. Proteomics, 2008, 7, 1317–1330 CrossRef CAS.
- J. Gimsa and C. Ried, Mol. Membr. Biol., 1995, 12, 247–254 CrossRef CAS.
Footnote |
| † Y. Shan and Z. Wang contributed equally to this paper. |
| This journal is © The Royal Society of Chemistry 2010 |
