Editorial: Vitamins and cofactors—chemistry, biochemistry and biology
This themed issue of Natural Product Reports has as its focus vitamins and cofactors. Vitamins are organic substances that are required in small amounts in the human diet, i.e. they do not include essential amino acids and fatty acids nor essential elements. One of the first clues that lack of certain substances in the diet can cause disease, and even death, dates back to 1753 when James Lind discovered that citrus fruits can prevent scurvy. However, it was not until the end of the 19th and beginning of the 20th century that more careful experimentation showed that there were substances other than protein, fats, sugars and minerals that were present in very small quantities in a healthy diet and that were essential for life. Christiaan Eijkman, after a decade working on polyneuritis in chickens, showed in 1897 that beri-beri, the equivalent disease in humans, could be prevented in rice-eating populations of south-east Asia by not removing the bran from the rice. Meanwhile, Frederick Hopkins, working on mice, showed that some foods contained “accessory factors” that were necessary for human health. Eijkman and Hopkins were awarded the 1929 Nobel Prize in Medicine for their discoveries. The word “vitamine”, however, was coined by Casimir Funk in 1912. Funk had attempted to isolate the substance from rice-bran that prevented polyneuritis and beri-beri and thought the vital substance was an amine, hence the name “vitamine”. In fact, he probably had not isolated the right substance, but nevertheless the name stuck. Subsequently, when it was realised that many vitamins were not amines, the final “e” was dropped.Initially, vitamins were divided into two fractions—fat-soluble (A) and water-soluble (B). Over time, as individual vitamins were identified, the B vitamins were given subscripts (B1, B2, etc., the exception being the water-soluble vitamin C), and the fat-soluble ones were given different letters (A, D, E, K). Many compounds were originally claimed to be vitamins and given numbers or letters but were subsequently found not to be essential—hence the gaps in the numbering and lettering.
Inevitably the reviews in this issue cannot cover the whole field of vitamins and as a result they concentrate on the B vitamins. One reason for this is that it is now known that the role of all the B vitamins is to provide cofactors for enzymic reactions (coenzymes). For example, vitamin B1 (thiamine) is required for the formation of thiamine pyrophosphate (TPP), the cofactor of several enzymes in energy production, sugar metabolism and other essential pathways. Fig. 1 gives a complete list of the B vitamins, why they are required, and the name of their deficiency disease. As coenzymes are only required in relatively small amounts by each cell, and as they are relatively ubiquitous in Nature and therefore available in most diets, animals have, over the course of evolution, lost the ability to biosynthesise these compounds, and have instead developed mechanisms for the uptake and absorption of the corresponding vitamins and transport to the site of their utilisation.
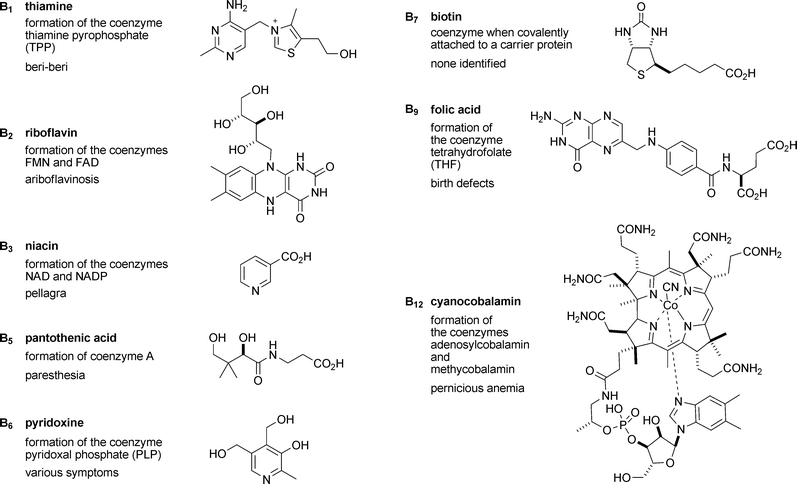 | ||
| Fig. 1 Structures of the B vitamins along with their role in cells and the disease caused by their deficiency. It should be noted that in many cases closely related structures should also be included. Thus, for example, in the case of vitamin B6 (pyridoxol), pyridoxal, pyridoxamine and their respective 5′-phosphates all have vitamin activity. | ||
Not all coenzymes are derived from vitamins, however. A few can be made by human cells, in particular the haems (a, b, and c), molybdenum cofactor (Moco), lipoic acid and coenzyme Q10 (ubiquinone). Structures of these coenzymes are shown in Fig. 2.
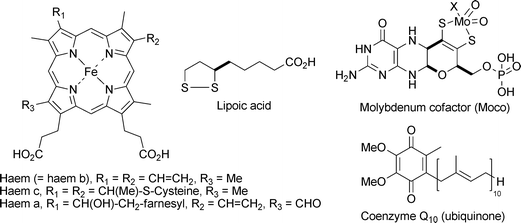 | ||
| Fig. 2 Coenzymes that can be made by human cells and so are not derived from vitamins. | ||
Many reviews are available in the literature that cover the chemistry and biology of one cofactor. However, in this issue the authors have taken a different approach. They have chosen to take different aspects of research into the chemistry, biochemistry and biology of vitamins and cofactors and show how, for each aspect, there are common themes for many of the pathways. Why should there be common themes for vitamins and cofactors that are not just as relevant to all other natural products?
Well, firstly there is the common factor that they are essential to humans and, for the vitamins at least, have to be present in the diet. Therefore they have in common (i) that they participate in essential metabolic processes, (ii) that deficiency of the vitamins (or genetic mutations that interfere with the biosynthesis of the other cofactors) causes diseases, whose physiology is sometimes difficult to relate to their metabolic function, and (iii) that, for the vitamins, nutritional aspects (e.g. quantity, availability, stability) are of great relevance. These are the aspects that are the subject of the review by Rébeillé et al. (DOI: 10.1039/b703104c).
A second common theme is that many of their biosynthetic pathways have proved to be unusually difficult to elucidate. There are several reasons why this is: (i) they are made in very small quantities; (ii) many of the intermediates are unstable; (iii) nearly all the intermediates are water-soluble and many are phosphorylated (often when this was not predicted a priori); and (iv) it has turned out that for some cofactors two or more distinct biosynthetic routes exist in different organisms, and until this was realised it caused considerable confusion. These factors, and more, are the subject of the review by Webb et al. (DOI: 10.1039/b703105j) on the elucidation of the biosynthetic pathways. The focus of this review is to look across the various biosynthetic pathways, at features that recur in several of them, rather than to follow each pathway from beginning to end. However, overviews of most of the pathways to cofactors are included either in this review or one of the others. For the benefit of readers who are particularly interested in biosynthetic pathways, Table 1 shows which review(s) and which scheme or figure gives the best overview of each pathway.
| Vitamin | Name | Review | Scheme or Figure |
|---|---|---|---|
| B1 | Thiamine | Webb et al. | Scheme 6 |
| B2 | Riboflavin | Webb et al. | Scheme 5 |
| Scott et al. | Scheme 2 | ||
| B3 | Niacin | Not covered | — |
| B5 | Pantothenate | Webb et al. | Scheme 2 |
| B6 (E. coli) | Pyridoxol | Webb et al. | Scheme 3 |
| Scott et al. | Scheme 10 | ||
| B6 (Yeast and B. subtilis) | Pyridoxal | Webb et al. | Scheme 10 |
| Scott et al. | Scheme 12 | ||
| B7 | Biotin | Webb et al. | Schemes 1 and 11 |
| Marquet et al. | Scheme 6 | ||
| B9 | Folate | Webb et al. | Scheme 8 |
| B12 | Cobalamin | Holliday et al. | Fig. 8 |
| Haem | Holliday et al. | Fig. 7 | |
| Marquet et al. | Scheme 11 | ||
| Moco | Webb et al. | Scheme 7 | |
| Mendel et al. | Fig. 3 | ||
| Lipoic acid | Marquet et al. | Scheme 5 |
Another common factor in vitamin biosynthesis is that it does not occur in humans but does in other organisms, where it is often essential. This makes the biosynthetic pathways of particular interest as targets for the development of antimicrobial drugs and agrochemical agents. In order to develop such compounds, it is important to have detailed knowledge of the biosynthetic pathways at the enzymic level, preferably with a good understanding of the reaction mechanisms and high resolution crystal structures. The review by Scott et al. (DOI: 10.1039/b703108b) takes a few selected examples of enzymes from vitamin biosynthesis and shows how the combination of crystal structures, mechanistic studies and inhibition studies can give an in-depth understanding of the enzymic process, thus laying the foundations for pharmaceutical or agrochemical applications.
Vitamins and cofactors are thought to be very early products of evolution, some of them probably dating from the “RNA world”. A study of the evolutionary relationships between the enzymes of their biosynthesis, as well as between the enzymes that use them, can give a fascinating glimpse into the way life evolved. This is the topic reviewed by Holliday et al. (DOI: 10.1039/b703107f). With ever more genomes being sequenced and enzyme structures being solved, there is a wealth of information that, with careful interpretation, could eventually tell us what evolved from what, step by step back to the earliest days of life on earth. Particularly fascinating are the instances where different biosynthetic pathways lead to the identical final cofactor (as for vitamins B1, B6 and B12). Did the different pathways arise independently to make a prebiotic compound that had already found use as a cofactor, or did they evolve divergently from a single primitive biosynthetic pathway that was already making the cofactor?
Probably the oldest cofactors of all are the iron-sulfur clusters, which, despite their sensitivity to oxygen, remain abundant in all living organisms. Studies of cofactor biosynthesis, however, have revealed new roles for iron-sulfur clusters beyond their well known electron transfer properties. In at least two of the cofactors (biotin and lipoic acid) it appears that the cluster also donates the sulfur atom(s) that get incorporated into the product. The other emerging role for iron-sulfur clusters is in the reductive cleavage of S-adenosylmethionine (SAM or AdoMet) to give the 5′-deoxyadenosyl radical. This radical can then, via hydrogen atom abstraction, initiate all sorts of reactions, including complicated rearrangements, as found in Moco biosynthesis. Bioinformatics studies predict that there are large numbers of such “radical SAM” enzymes, the vast majority of which are of unknown function. The article by Marquet et al. (DOI: 10.1039/b703109m) reviews these aspects of iron-sulfur clusters, not just in cofactor biosynthesis but also in other metabolic pathways. Also covered are a class of “chemically difficult” enzymic reactions which are thought to be facilitated by electron donation from high potential iron-sulfur clusters into π*-orbitals of the substrate.
Finally, once the organic part of a cofactor has been synthesised, it may need to be combined with the appropriate metal ion, and all must be assembled into their cognate apoproteins. In many cases this is not just a simple self-assembly process but needs further enzymes, chaperones and transport proteins. For example the assembly of cytochrome c from haem and apocytochrome c requires, in some organisms, twelve different proteins. The article by Mendel et al. (DOI: 10.1039/b703112m) reviews the current knowledge about these assembly systems, with particular emphasis on Moco but also covering iron-sulfur clusters and haems, cobalt and B12, and copper-containing enzymes. It is clear there is much still to learn about these complicated systems.
We hope that this series of reviews will give the reader a good insight into the processes of biosynthesis and assembly and the physiological role of vitamins and cofactors. In particular we hope that, by looking at certain aspects across the range rather than focussing on individual cofactors, it will give a different insight from other reviews in the area. There is, of course, a lot that has not been considered. The fat-soluble vitamins and cofactors (e.g. vitamin K and coenzyme Q10) have not been covered. Cofactors not found in mammals have hardly been mentioned. Methanogenic bacteria, in particular, have a number of cofactors involved in methane production which are unique.
But most of all we have not covered the enzymic reactions in which these cofactors participate (except in so far as a cofactor participates in its own biosynthesis or that of another cofactor). So to fill that hole, we illustrate here the chemical mechanism of just one enzyme, but a remarkable one in many respects. The pyruvate dehydrogenase (PDH) complex catalyses the reaction at the centre of any metabolic chart. It is the last step of glycolysis, which produces acetyl CoA, the central metabolite that feeds the citric acid cycle as well as being the basic precursor of fatty acids, terpenes, polyketides and much else. PDH has a remarkable architecture: it consists of three different types of subunit (E1, E2 and E3), which together form a huge multimeric complex with a molecular weight of up to 107 Da. But, from our point of view, the most remarkable feature is the mechanism, because it involves no fewer than five of the cofactors that have been the subject of these reviews, as shown in Fig. 3.
Finian J. Leeper and Alison G. Smith
University of Cambridge
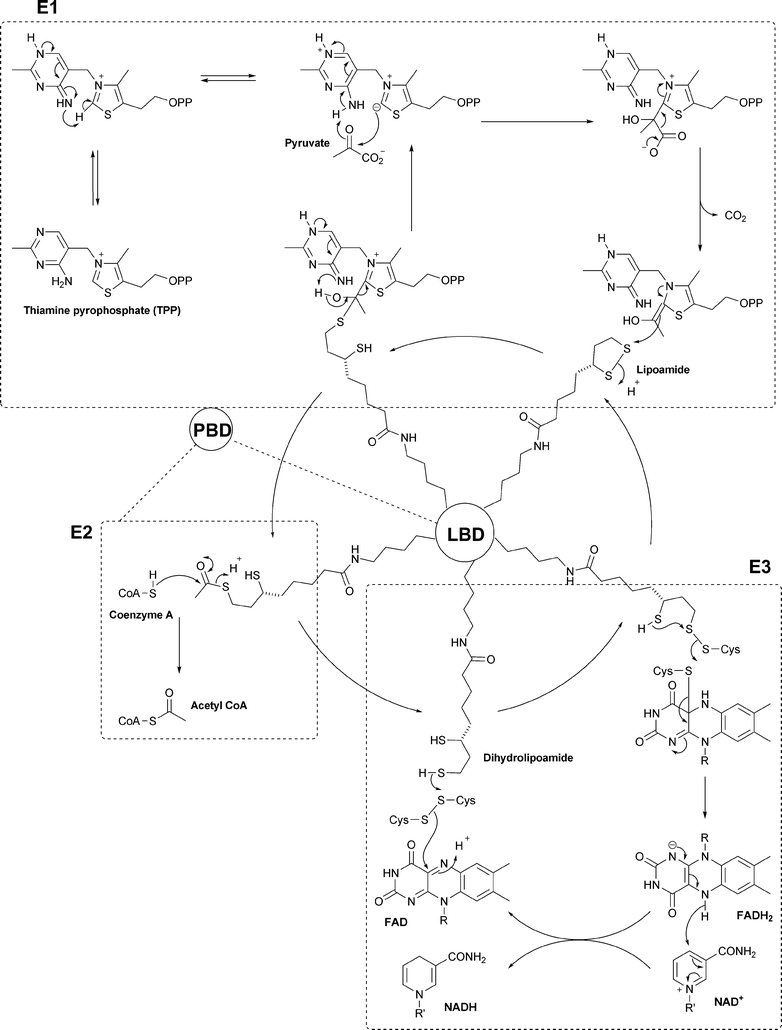 | ||
| Fig. 3 Schematic mechanism for the pyruvate dehydrogenase complex. The reactions in the three dashed boxes occur in the active sites of the E1, E2 and E3 subunits as shown. The E2 subunits (24 or 60 of them depending on the organism) form the core of the complex. Each E2 has a flexible arm containing a peripheral binding domain (PBD) and then one or more lipoyl binding domains (LBD). Each PBD binds to an E1 or an E3 subunit which form a hollow sphere. In the space between the core and the outer sphere the lipoyl binding domain is free to move around visiting the different active sites to effect the reactions shown. | ||
| This journal is © The Royal Society of Chemistry 2007 |
