Hot off the press
Hot off the Press highlights recently published work for the benefit of our readers. Our contributors this month have focused on peptide beacons for targeting antibodies, peptide ligation using gold nanoparticles and a new method for fabricating probe-bearing particles. New contributors are always welcome. If you are interested please contact http://molbiosyst@rsc.org for more information, we'd like to hear from you.
Peptide beacons—peptide based optical biosensors
Molecular beacons (MB), stem—loop DNA with fluorophore and quencher moieties attached to the both ends of the stem, have been used for more than a decade in detection of minute amounts of DNA. The principle based on FRET was applied to different designs and it proved to be sensitive and convenient, but applicable only to DNA detection. To extend the utility of this approach also to other analytes such as antibodies, other the changes in design are needed. Now, the researchers at the University of California have come up with the possible solution. They have designed a novel class of sensors termed peptide beacons (PB) which can be used in detection of picomolar amounts of target antibodies. The principle of PBs makes used of the peptide’s property to be highly dynamic while unbound and rigid upon binding to a macromolecular target. PB architecture was based on six residue epitope from HIV protein p17. The important factor for a choice of this particular polypeptide was that two ends of the unbound epitope collide on the time scale <100 ns, but when bound to the target the epitope adopts a rigid conformation. Ruthenium(II) bisbipyridine-phenanthroline was used as a fluorophore exciting in the visible region and methyl viologen as a quencher. Both were attached to the end of the peptide via cysteine linkers. When the target antibody was added to PB solution, the fluorophore emission was increased 6-fold and the change was observed only in the presence of the right target indicating excellent selectivity.Such PB sensing strategy could be used in many different applications, the only requirement being that fluorophore and quencher do not disrupt binding and that the bound state of peptide leads to at least few angstroms of separation between molecules at the ends. This indicates the general applicability for many different targets. The only disadvantage at the moment, as the authors point out, is relatively low sensitivity compared to ELISA or Western Blots which go down to femtomolar range. However, PB approach does not require such extensive batch processing and the use of additional equipment. Therefore it might be expected that PBs will find additional uses as a selective sensors that can be applied in complex samples such as saliva and blood.
K.J. Oh, K. J. Cash, V.Hugenberg, K. W. Plaxco, Bioconj. Chem., 2007, 18(3), 607–609.
Reviewed by: Ljiljana Fruk, University of Dortmund, Germany.Catalysis of peptide ligation by gold nanoparticles
Gold nanoparticles (GNPs) have induced a little scientific revolution and largely contributed to the development of nanotechnology. Till now they have been used as versatile scaffolds for building of the nanoarchitectures or in the design of elaborate optical or electrochemical sensors. Some recent studies also focused on specially designed GNPs which can be used as catalysts in reactions involving the bonds cleavage. Now the group of scientists from the University of Massachusetts employed GNPs to bring together the peptide fragments and assist in peptide coupling reaction (Fig. 1). This is based on the ability of cationic GNPs to bind the negatively charged peptide fragments which should be coupled and therefore acting as a template for their ligation into larger peptide. The ligation described is achieved by using one peptide fragment containing a thioester at C-terminus and the other cysteine at N-terminus in reducing environment (Kent's native chemical ligation). 250 µM solution of the fragments was used in presence of up to 75 µM of GNPs (6 nm) for the ligation. The product formation was followed by HPLC and showed the clear increase in the amount of ligated peptide with addition of cationic GNPs. In the control reaction with negatively charged GNP no product was formed as well as in the case when binding of the fragments was hindered by addition of monomeric cations. This indicated that the electrostatic interactions between GNPs and the fragments are the main step in the formation of appropriate templates and subsequent ligation. In future this approach could be extended to larger scaffolds and catalysis of different supramolecular coupling reactions. Finally, as the authors state, this system could also be used as a model to study the role of NPs in the polymerization of early biopolymers in prebiotic conditions.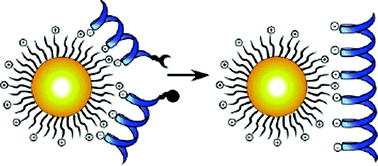 | ||
| Fig. 1 The design of functionalized, cationic GNPs as a template for peptide ligation. Reprinted with permission from Y. Fillon, A. Verma, P. Ghosh, D. Ernenwein, V. M. Rotello, J. Chmielewski, J. Am. Chem. Soc, 2007, 129, 6676–6677. Copyright 2007 American Chemical Society. | ||
Y. Fillon, A. Verma, P. Ghosh, D. Ernenwein, V. M. Rotello, J. Chmielewski, J. Am. Chem. Soc, 2007, 129, 6676–6677.
Reviewed by: Ljiljana Fruk, University of Dortmund, Germany.More from p53
p53 has a wealth of research detailing its many functions. However this database of information provides no clues as to why this gene is mutated so frequently in various cancers. Analysis of mutations found within gastric cancer and non-cancerous gastritis has shown that the most commonly mutated gene is p53. In addition, Helicobacter pylori is identified as a class one carcinogen by the World Health Organization because of the role this strain plays in chronic gastric inflammation.A paper in Nature Medicine by Matsumoto, et al. investigates the putative connection between H. pylori infections and p53 mutations. The group shows that cagPAI-positive H. pylori infections correlate with higher frequencies of p53 mutations. p53 was highly expressed in gastric epithelial cells while c-Myc was expressed at low levels and showed no mutations from the analyzed clones. The data reported suggests that somatic mutations depend on the gene level of transcription. These somatic mutations also coincide with the upregulation of activation-induced cytidine deaminase (AID) expression through the IKK-dependent NF-κB pathway. In the end, this group has demonstrated a strong link between p53 mutations in gastric epithelial cells and cagPAI-positive H. pylori infection. Despite the incongruent evidence in vivo, this paper opens the field to a mechanism controlling another aspect of the ever-evolving p53 story.
Matsumoto, et al., Nature Medicine, 2007, 13(4), 470–6.
Reviewed by: Melissa O'Neal, UT Southwestern Medical Center, USA.Molecular probes that wear nametags
Highly parallel binding assays have emerged as crucial tools for genetic and epigenetic analysis, transcript profiling, proteomics, and combinatorial library screening (1–3). These methods harness molecular recognition to rapidly and simultaneously characterize many components in complex mixtures. They use large collections (hundreds to millions) of potential ligands, or probes, to capture and quantitate specific biomolecules. Knowledge of the probes involved in the observed binding interactions is usually critical to interpretation of the assay. One way to keep track of the identity of each probe is to immobilize all probes on a single solid support, a microarray, with a defined position for each. An alternative format presents the array of probes on a population of microscopic particles, such as beads or rods, with a unique probe molecule on each particle. These particle-based formats can have some advantages over planar array formats in throughput, binding kinetics, and array quality control (2). Because the identity of the probe on a particle is no longer encoded in its position on a surface, many schemes have been devised that allow each particle to carry an easily read signature identifying the probe molecule it displays (4, 5). However, available encoding methods have significant limitations in the number of different probes that can be encoded, the ease and robustness of particle fabrication, and interference between the encoding signal and the signal for analyte measurement.A recent report (6) describes a new method for fabricating probe-bearing particles that alleviates some of these drawbacks. In this approach, particles are created with distinct regions dedicated to displaying probes and displaying encoding patterns. The particles are photopolymerized from adjacent streams of photoreactive monomers (Fig. 2). Monomer streams are joined in a microfluidic channel where they flow parallel to each other with little mixing between them. Thus, when polymerized by a pattern of light that spans the streams, each stream gives rise to a distinct region of the nascent polymer particle. A stream containing probe molecules that are modified for incorporation into the polymer gives rise to a defined region of the particle for specific binding and detection of an analyte. A separate stream that includes a fluorescent monomer creates a fluorescent region of the particle, which can be formed with a coding pattern of holes that identifies the particle and defines its orientation in space (Fig. 2, inset). The use of a pattern of holes on a fluorescent background as the encoding scheme allows creation of at least one million distinguishable particles, far more than the ∼100 attainable with standard fluorescent barcodes. Because the encoding region and probe regions are distinct, their signals do not interfere with each other, and the same fluorophore can be used in the encoding region and for analyte detection. By using even more streams, multiple different probe regions and internal negative control regions can be created in a single particle. Similarly, gradients of probe density can be created to expand the dynamic range of analyte detection. Analyte detection and code read-out can be done with a device that aligns the particles as they flow over a detector.
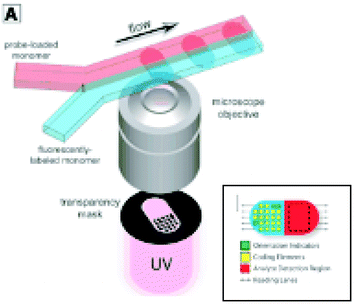 | ||
| Fig. 2 Schematic diagram of system for fabrication of multifunctional polymer particles by photopolymerization across two adjacent laminar streams. Inset: Diagram of a multifunctional particle showing a dot-encoded region and a probe-bearing analyte detection region. From D. C. Pregibon et al., Science, 315,1393–1396 (9 March 2007).Reprinted with permission from AAAS. | ||
The demonstrated and potential power of highly parallel assays has led to a profusion of multiplex binding assay formats and probe array fabrication methods, each with its own particular strengths and target applications. Time will tell which applications will be best enabled by this novel fabrication methodology. With its many unique and useful features, it may make possible high throughput assays that have yet to be considered.
1. Hoheisel, J. D., Nat. Rev. Genet., 2006, 7, 200–210.
2. Nolan, J. P., Sklar, L. A., Trends Biotechnol., 2002, 20, 9–12.
3. Kodadek T. Bachhawat-Sikder K. Mol. BioSyst., 2006, 2, 25–35.
4. Finkel, N. H., Lou, X., Wang, C., He, L., Anal. Chem., 2004, 76, 352A–359A.
5. Braeckmans, K., Smedt, S. C. D., Leblans, M., Pauwels, R., Demeester, J., Nat. Rev. Drug Discov., 2002, 1, 447–456.
6. Pregibon, D. C., Toner, M., Doyle, P. S., Science, 2007, 315, 1393–1396.
Hot off the RSC press
Illuminating infection
To detect a parvovirus you just need to lighten up, say Italian scientists.Aldo Roda and co-workers at the University of Bologna have developed an ultrasensitive way of detecting parvovirus B19 infections. The virus is responsible for a range of clinical syndromes in humans, including aplastic crisis. Aplastic crisis causes suppression of red blood cell production and is a particular threat to children with sickle cell disease, for whom infection can result in severe anaemia.
By exploiting three detection techniques in combination, Roda’s team has engineered a system that they claim is more sensitive than any previously developed in situ assays. The method is sensitive enough to detect the virus within cells and at early stages of infection.
In Roda’s assay, infected cells are exposed to short strands of a synthetic version of DNA: peptide nucleic acid, or PNA. PNA has a neutral backbone rather than the negatively charged one of DNA and RNA, eliminating electrostatic repulsion effects and allowing fast, strong binding to DNA and RNA. The PNA strands bind to the viral nucleic acids and are then labelled with an enzyme (alkaline phosphatase) using a linker molecule already on the PNA.
The key to the new system is chemiluminescence, light produced by a chemical reaction. Used in detection, it is extremely sensitive and the signal can be quantified. By adding a substrate that chemiluminesces on binding to the alkaline phosphatase label, the PNA and hence viral nucleic acids can be detected (Fig. 3).
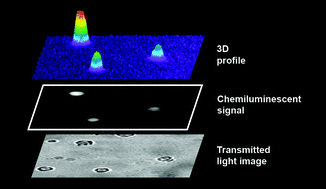 | ||
| Fig. 3 Chemiluminescence is used to detect cells infected with parvovirus B19. Figure used with permission, Royal Society of Chemistry, 2007. | ||
Importantly, Roda’s method does not require amplification or even extraction of the viral DNA and RNA from the cells, reducing sample handling and potentially shortening assay times. According to the team, the system could eventually be used to monitor persistent low-level viral replication in chronic infections and to study virus replication kinetics and the effects of antiviral drugs.
F Bonvicini et al., Analyst, 2007, DOI: 10.1039/b701664f
Reviewed by: Freya Mearns, Royal Society of Chemistry, Cambridge, UK.Taking a protein’s fingerprint
Researchers in Singapore have used microarray technology to study the enzyme family linked to diseases such as SARS.3CL protease is an important enzyme for the SARS virus, the cause of severe acute respiratory syndrome. It is a cysteine protease, an enzyme that hydrolyses peptide bonds using a cysteine residue in its active site. Now Shao Yao and colleagues at the National University of Singapore have made an array to identify cysteine protease activity and used it to study protease inhibitors; a method that could eventually be used to find drugs to target 3CL protease.
The group made a series of fluorescent probes, each containing a vinyl sulfone group, a group known to inhibit cysteine proteases. They varied the structures by changing the amino acid to be recognised by the protease in each compound. As a general test for cysteine proteases, a cocktail of these probes was used to screen an array of enzymes. Any cysteine proteases could be identified because they were the only enzymes that the probes bound to and so labelled, giving a fluorescent signal (Fig. 4).
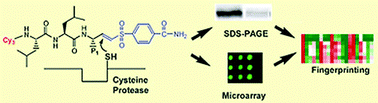 | ||
| Fig. 4 A cysteine protease recognises vinyl sulfone probes to give its own protein fingerprint. Figure used with permission, Royal Society of Chemistry, 2007. | ||
Another microarray was used to screen each of the identified cysteine proteases with each of the probes, separately; this gave a fingerprint for each enzyme. Some of the proteases act only on substrates with specific amino acids in the peptide bond to be hydrolysed; these enzymes showed the strongest fluorescent signal with probes containing those amino acids. Others are more general and were labelled equally well by many of the probes. ‘The resulting fingerprints reveal unique signatures for individual proteins, providing biological insights into their functional roles,’ Yao said.
The group used a third array to screen small molecule inhibitors against the identified proteases to find inhibitor fingerprints. They added solutions of varying inhibitor concentration to each protease, followed by a vinyl sulfone probe. Increased inhibition was shown by lower probe fluorescence as the inhibitor blocked the probe from the enzyme.
Yao explained that future challenges for the field would include developing microarrays with detection methods that could be used to follow an enzyme as it reacts. His group hopes to extend its research to other kinds of enzymes, such as kinases and phosphatases.
M Uttamchandani et al., Chem. Commun., 2007, 1518
Reviewed by: Rachael Warfield, Royal Society of Chemistry, Cambridge, UK.Hot-wiring enzymes for fuel cells
Stable enzyme-modified graphite electrodes could be used in fuel cells, according to researchers in the UKLaccases are copper-containing enzymes that use electrons to reduce oxygen to water efficiently, making them of interest as electrocatalysts at the cathodes of hydrogen–oxygen fuel cells. In this type of fuel cell, hydrogen is oxidised to hydrogen ions and electrons, which flow to the cathode (supplying power) where they reduce oxygen.
Fraser Armstrong and colleagues at the University of Oxford have chemically attached anthracene to graphite electrodes to form a stable attachment between the electrode and laccase. Armstrong described anthracene as ‘a long hydrophobic molecule, similar to laccase’s own substrate molecules, which can penetrate the active site pocket and approach closely to one of the copper atoms’ where the oxygen reduction takes place. The anthracene not only binds the laccase to the electrode, but, as it can conduct electrons, it also delivers them deep into the enzyme close to the site for oxygen reduction.
The anthracene ‘plug’ modification of the electrode is essential to provide the high and long-term oxygen-reduction activity. Without it, the electrons are not delivered efficiently into the enzyme and the binding of the laccase to the electrode is not stable, both of which leads to reduced activity. The ability to use graphite is also an advantage; it is relatively cheap compared to precious metals, such as gold, which have been previously used as electrodes for laccases.
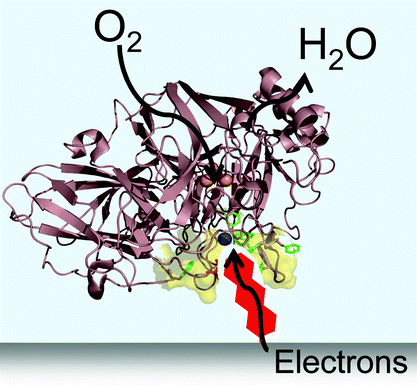 | ||
| Fig. 5 Figure used with permission, Royal Society of Chemistry, 2007. | ||
To complete the fuel cell, Armstrong said that in future research they would like to ‘identify rational attachment strategies for other enzymes, particularly hydrogenases, which can be incorporated with laccases to produce small fuel cells’.
C F Blanford, R S Heath and F A Armstrong, Chem. Commun., 2007, DOI: 10.1039/b703114a
Reviewed by: Nicola Convine, Royal Society of Chemistry, Cambridge, UK.| This journal is © The Royal Society of Chemistry 2007 |
