Oscillations of Min-proteins in micropatterned environments: a three-dimensional particle-based stochastic simulation approach†
Abstract
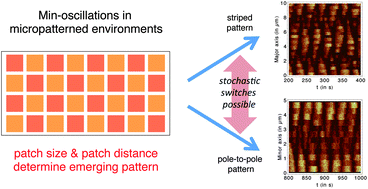
The Min-proteins from E. coli and other bacteria are the best characterized pattern forming system in cells and their spatiotemporal oscillations have been successfully reconstituted in vitro. Different mathematical and computational models have been used to better understand these oscillations. Here we use particle-based stochastic simulations to study Min-oscillations in patterned environments. We simulate a rectangular box of length 10 μm and width 5 μm that is filled with grid or checkerboard patterns of different patch sizes and distances. For this geometry, we find different stable oscillation patterns, typically pole-to-pole oscillations along the minor axis and striped oscillations along the major axis. The Min-oscillations can switch from one pattern to the other, either effected by changes in pattern geometry or stochastically. By automatic analysis of large-scale computer simulations, we show quantitatively how the perturbing effect of increased patch distance can be rescued by increased patch size. We also show that striped oscillations occur robustly in arbitrarily shaped filamentous E. coli cells. Our results highlight the robustness and variability of Min-oscillations, put limits on the effect of putative division sites, and provide a powerful computational framework for future studies of protein self-organization in patterned environments.
- This article is part of the themed collection: Proteins, cells, and tissues in patterned environments

 Please wait while we load your content...
Please wait while we load your content...