Machine learning designs non-hemolytic antimicrobial peptides†
Abstract
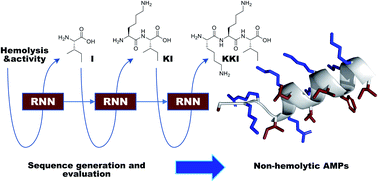
Machine learning (ML) consists of the recognition of patterns from training data and offers the opportunity to exploit large structure–activity databases for drug design. In the area of peptide drugs, ML is mostly being tested to design antimicrobial peptides (AMPs), a class of biomolecules potentially useful to fight multidrug-resistant bacteria. ML models have successfully identified membrane disruptive amphiphilic AMPs, however mostly without addressing the associated toxicity to human red blood cells. Here we trained recurrent neural networks (RNN) with data from DBAASP (Database of Antimicrobial Activity and Structure of Peptides) to design short non-hemolytic AMPs. Synthesis and testing of 28 generated peptides, each at least 5 mutations away from training data, allowed us to identify eight new non-hemolytic AMPs against Pseudomonas aeruginosa, Acinetobacter baumannii, and methicillin-resistant Staphylococcus aureus (MRSA). These results show that machine learning (ML) can be used to design new non-hemolytic AMPs.


 Please wait while we load your content...
Please wait while we load your content...