Facile saccharide-free mimetics that recapitulate key features of glycosaminoglycan sulfation patterns†
Abstract
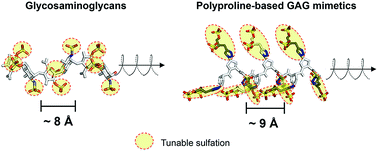
Controlling glycosaminoglycan (GAG) activity to exploit its immense potential in biology ultimately requires facile manipulation of sulfation patterns associated with GAGs. However, satisfying this requirement in full remains challenging, given that synthesis of GAGs is technically arduous while convenient GAG mimetics often produce sulfation patterns that are uncharacteristic of GAGs. To overcome this, we develop saccharide-free polyproline-based GAG mimetics (PGMs) that can be facilely assembled via amide coupling chemistry. Molecular dynamics simulations show that PGMs recapitulate key GAG structural features (i.e. ∼9 Å-sized repeating units, periodicity and helicity) and as with GAGs, can be tuned to introduce systematic variations in sulfate clustering and spacing. Functionally, a variety of PGMs control various GAG activities (concerning P-selectin, neurotrophic factors and heparinase) and exhibit GAG-like characteristics such as progressive modulation, comparable effectiveness with heparins, need for different sequences to suit different activities and the presence of a “minimal bioactive length”. Furthermore, PGMs produce consistent effects in vivo and successfully provide therapeutic benefits over cancer metastasis. Taken together with their high level of biosafety, PGMs answer the long-standing need for an effective and practicable strategy to manipulate GAG-appropriate sulfation patterns and exploit GAG activity in medicine and biotechnology.


 Please wait while we load your content...
Please wait while we load your content...