Virtual screening for high affinity guests for synthetic supramolecular receptors†
Abstract
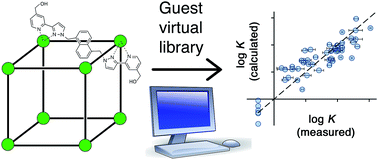
The protein/ligand docking software GOLD, which was originally developed for drug discovery, has been used in a virtual screen to identify small molecules that bind with extremely high affinities (K ≈ 107 M−1) in the cavity of a cubic coordination cage in water. A scoring function was developed using known guests as a training set and modified by introducing an additional term to take account of loss of guest flexibility on binding. This scoring function was then used in GOLD to successfully identify 15 new guests and accurately predict the binding constants. This approach provides a powerful predictive tool for virtual screening of large compound libraries to identify new guests for synthetic hosts, thereby greatly simplifying and accelerating the process of identifying guests by removing the reliance on experimental trial-and-error.

 Please wait while we load your content...
Please wait while we load your content...