Exploitation of stem-loop DNA as a dual-input gene sensing platform: extension to subtyping of influenza A viruses†
Abstract
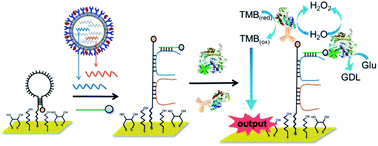
We herein report a strategy coupling DNA structural switches with an enzymatic cascade reaction for the specific and sensitive identification of dual-target nucleic acids toward the subtyping of influenza viruses. During the recognition process, a hairpin DNA was hybridized with dual target sequences at its stem and a partial section of the loop region to form duplexes. The concurrent presence of dual targets was reported as an explicit output interpreted by the enzymatic cascade reaction. The conformational changes of the hairpin–targets duplex and the reaction kinetics upon dual hybridization were explored using a molecular beacon in conjunction with a FRET approach. The unzipping event was found to be predominated by a strand displacement which occurred in the stem of hairpins, and its rate constants were calculated as 3.1 × 103 M−1 s−1 (gHA alone) and 5.8 × 103 M−1 s−1 (gHA & gNA). This strategy was further demonstrated on a multi-well electrochemical chip for the subtyping of influenza viruses (e.g., A/duck/Taiwan/DV30-2/2005 (H5N2)) through the simultaneous identification of genes encoding hemagglutinin (HA) and neuraminidase (NA). Moreover, this genosensor featured an effective capability to analyze unpurified amplicons generated from the genuine viral genome, as it could unambiguously discriminate between H5N2 and non-H5N2 viruses. To the best of our knowledge, this study is the first to demonstrate the leveraging of an individual hairpin DNA across concurrent analysis of dual target genes. Pursuant to the elegant characterizations with high applicability, extensible flexibility, and powerful utility, this platform holds great promise for use as a sophisticated molecular diagnostics tool against variable influenza viruses.

 Please wait while we load your content...
Please wait while we load your content...