Oxidative dearomatisation: the key step of sorbicillinoid biosynthesis†
Abstract
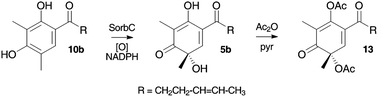
An FAD-dependent monooxygenase encoding gene (SorbC) was cloned from Penicillium chrysogenum E01-10/3 and expressed as a soluble protein in Escherichia coli. The enzyme efficiently performed the oxidative dearomatisation of sorbicillin and dihydrosorbicillin to give sorbicillinol and dihydrosorbicillinol respectively. Bioinformatic examination of the gene cluster surrounding SorbC indicated the presence of two polyketide synthase (PKS) encoding genes designated sorbA and sorbB. The gene sorbA-encodes a highly reducing iterative PKS while SorbB encodes a non-reducing iterative PKS which features a reductive release domain usually involved in the production of polyketide aldehydes. Using these observations and previously reported results from isotopic feeding experiments a new and simpler biosynthetic route to the sorbicillin class of secondary metabolites is proposed which is consistent with all reported experimental results.

 Please wait while we load your content...
Please wait while we load your content...