A new class of platinum(ii) complexes with the phosphine ligand pta which show potent anticancer activity†
Abstract
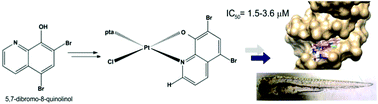
The anticancer potential of sixteen platinum(II) complexes with general formulae [PtCl(hq)(S-dmso)] (1a–8a) and [PtCl(hq)(pta)] (1b–8b) (where hq is 5-chloro-7-iodo-8-quinolinol (clioquinol; cqH) (1a, 1b), 8-hydroxy-5-nitroquinoline (nitroxoline; nxH) (2a, 2b), 5,7-dichloro-8-quinolinol (3a, 3b), 5,7-diiodo-8-quinolinol (4a, 4b), 5,7-dibromo-8-quinolinol (5a, 5b), 5,7-dichloro-8-hydroxy-2-methyl-quinoline (6a, 6b), 8-hydroxyquinoline (7a, 7b) and 8-quinolinethiol (8a, 8b); dmso is dimethyl sulfoxide and pta is 1,3,5-triaza-7-phosphaadamantane) was determined through in vitro cytotoxicity assay in human fibroblasts (MRC5) and two carcinoma cell lines (A375 and A549) and embryotoxicity assay in a zebrafish model. Interactions with double stranded DNA through in vitro assay and a molecular docking study were examined. All complexes, except 6a, exhibited a high cytotoxic effect on MRC5 cells at a concentration of 10 μg mL−1 while 1b, 5a, 6a and 3b showed selective toxicity towards carcinoma cell lines. In general, pta-based complexes (series b) were more toxic according to the results of a MTT screen and the LC50 values obtained in zebrafish (Danio rerio) assay; they also induced higher oxidative stress in this model. Successful cellular uptake of complexes was shown by the ICP-MS methodology. The binding propensity of the complex with DNA obtained in in silico studies can be correlated with those from the experimental investigation. Compounds with the highest binding potential, according to the interaction energy value, were 1b, 3b, 6b and 5b. From observations of the DNA interaction ability and of the in silico assessment, no apparent DNA fragmentation was observed either on DNA extracted from the treated cancer cell line or from the zebrafish embryos.


 Please wait while we load your content...
Please wait while we load your content...