An integrative analysis of transcriptomic response of ethanol tolerant strains to ethanol in Saccharomyces cerevisiae†
Abstract
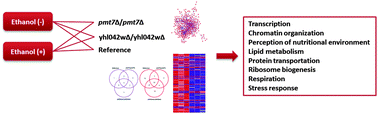
The accumulation of ethanol is one of the main environmental stresses that Saccharomyces cerevisiae cells are exposed to in industrial alcoholic beverage and bioethanol production processes. Despite the known impacts of ethanol, the molecular mechanisms underlying ethanol tolerance are still not fully understood. Novel gene targets leading to ethanol tolerance were previously identified via a network approach and the investigations of the deletions of these genes resulted in the improved ethanol tolerance of pmt7Δ/pmt7Δ and yhl042wΔ/yhl042wΔ strains. In the present study, an integrative system based approach was used to investigate the global transcriptional changes in these two ethanol tolerant strains in response to ethanol and hence to elucidate the mechanisms leading to the observed tolerant phenotypes. In addition to strain specific biological processes, a number of common and already reported biological processes were found to be affected in the reference and both ethanol tolerant strains. However, the integrative analysis of the transcriptome with the transcriptional regulatory network and the ethanol tolerance network revealed that each ethanol tolerant strain had a specific organization of the transcriptomic response. Transcription factors around which most important changes occur were determined and active subnetworks in response to ethanol and functional clusters were identified in all strains.

 Please wait while we load your content...
Please wait while we load your content...