Curcumin inhibits proliferation–migration of NSCLC by steering crosstalk between a Wnt signaling pathway and an adherens junction via EGR-1†
Abstract
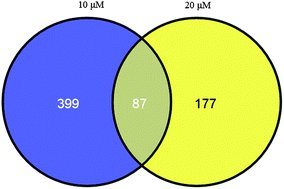
A microarray analysis of differential genes by curcumin treatment was performed and the crucial pathways associated with non-small cell lung cancer (NSCLC) were investigated. Total RNAs from 0, 10 or 20 μM curcumin treated NSCLC 95D cells were used to prepare microarray chips. The differentially expressed genes (DEGs) were identified using the RankProducts package and their function was predicted by DAVID and gene set enrichment analysis. The pathway crosstalk was analyzed by mapping the gene expression profiles into protein–protein interaction databases. Validation of the microarray results was performed by cell viability, cell migration and western blot analyses. A total of 486 (10 μM) and 264 (20 μM) DEGs were screened between the 95D cells in the presence and absence of curcumin. Function enrichment analysis indicated the DEGs were mainly involved in the steroid biosynthetic process and regulation of autophagy. Pathway crosstalk analysis suggested there was a significant interaction between NSCLC and adherens junctions (or Wnt signaling pathways, which are important for cancer cell proliferation and invasion) in both 10 μM and 20 μM curcumin treated 95D cells. Furthermore, early growth response (EGR-1) was demonstrated to regulate the crosstalk between adherens junctions and Wnt signaling pathways, indicating that EGR-1 may also regulate cell proliferation and migration. This hypothesis was validated by in vitro experiments: EGR-1 was decreased after curcumin treatment. Curcumin exhibited a significant anti-proliferation and anti-migration activity in NSCLC 95D cells, possibly by steering the crosstalk between the Wnt signaling pathway and adherens junction via EGR-1.

 Please wait while we load your content...
Please wait while we load your content...