Live sperm trap microarray for high throughput imaging and analysis†
Abstract
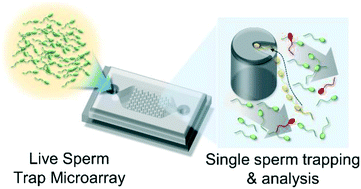
There is a growing appreciation and understanding of cell-to-cell variability in biological samples. However, research and clinical practice in male fertility has relied on population, or sample-based characteristics. Single-cell resolution is particularly important given the winner-takes-all nature of both natural and in vitro fertilization: it is the properties of a single cell, not the population, that are passed to the next generation. While there are a range of methods for single cell analysis, arraying a larger number of live sperm has not been possible due to the strong locomotion of the cells. Here we present a 103-trap microarray that traps, aligns and arrays individual live sperm. The method enables high-resolution imaging of the aligned cell head, the application of dye-based DNA and mitochondrial analyses, and the quantification of motility characteristics, such as tail beat. In testing, a 2400-post array trapped ∼400 sperm for individual analyses of tail beating frequency and amplitude, DNA integrity via acridine orange staining, and mitochondrial activity via staining. While literature results are mixed regarding a possible correlation between motility and DNA integrity of sperm at sample-level, results here find no statistical correlation between tail beat characteristics and DNA integrity at the cell-level. The trap array uniquely enables the high-throughput study of individual live sperm in semen samples – assessing the inherently single-cell selection process of fertilization, with single-cell resolution.


 Please wait while we load your content...
Please wait while we load your content...