An automated microfluidic gene-editing platform for deciphering cancer genes†
Abstract
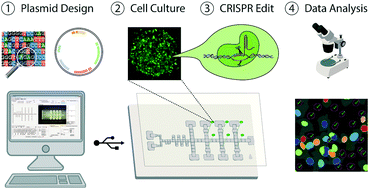
Gene-editing techniques such as RNA-guided endonuclease systems are becoming increasingly popular for phenotypic screening. Such screens are normally conducted in arrayed or pooled formats. There has been considerable interest in recent years to find new technological methods for conducting these gene-editing assays. We report here the first digital microfluidic method that can automate arrayed gene-editing in mammalian cells. Specifically, this method was useful in culturing lung cancer cells for up to six days, as well as implementing automated gene transfection and knockout procedures. In addition, a standardized imaging pipeline to analyse fluorescently labelled cells was also designed and implemented during these procedures. A gene editing assay for interrogating the MAPK/ERK pathway was performed to show the utility of our platform and to determine the effects of knocking out the RAF1 gene in lung cancer cells. In addition to gene knockout, we also treated the cells with an inhibitor, Sorafenib Tosylate, to determine the effects of enzymatic inhibition. The combination of enzymatic inhibition and guide targeting on device resulted in lower drug concentrations for achieving half-inhibitory effects (IC50) compared to cells treated only with the inhibitor, confirming that lung cancer cells are being successfully edited on the device. We propose that this system will be useful for other types of gene-editing assays and applications related to personalized medicine.


 Please wait while we load your content...
Please wait while we load your content...