Exploring the substrate specificity and catalytic mechanism of imidazolonepropionase (HutI) from Bacillus subtilis†
Abstract
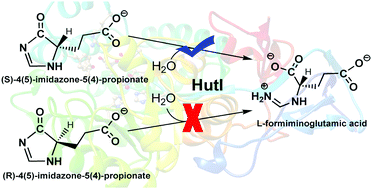
Imidazolonepropionase (HutI, EC 3.5.2.7) catalyzes the hydrolytic cleavage of carbon–nitrogen bond in 4-imidazolone-5-propionic acid (IPA) to yield L-formiminoglutamic acid, which is the third step in the universal histidine degradation pathway. In this article, using a combined quantum mechanics and molecular mechanics (QM/MM) approach, the specificity and catalytic mechanism of HutI from Bacillus subtilis have been explored by considering the four isomers of (S)- and (R)-enantiomers of IPA (SIPA-1, SIPA-2, RIPA-1 and RIPA-2). Our calculations reveal that the activation of hydrolytic water (a zinc-bound water) is performed by residue E252 via a “bridging” water molecule, which occurs before binding of the substrate. After the substrate binding, this activation channel is blocked by the substrate, and the other two residues (D324 and H272) cannot act as the general base to activate the hydrolytic water. For the two (S)-enantiomers of IPA, HutI can specifically convert one isomer of (S)-enantiomer (SIPA-1) to L-formiminoglutamic acid with an energy barrier of 16.6 kcal mol−1. The conversion of another (S)-enantiomer (SIPA-2) corresponds to the energy barrier of 21.9 kcal mol−1. However, for the two isomers of (R)-enantiomer, RIPA-1 corresponds to a higher energy barrier (21.8 kcal mol−1), and RIPA-2 is associated with a weak binding in the active site compared to SIPA-2. Thus, on the basis of our calculations, SIPA-1 is suggested to be the most favorable substrate for HutI, whereas the hydrolytic cleavage of SIPA-2 may require a preliminary isomerization to SIPA-1.

 Please wait while we load your content...
Please wait while we load your content...