Workflow for fast lipid tissue screening using LESA-FT-ICR-MS†
Abstract
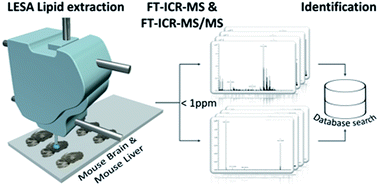
Lipid screening of biological substrates is an important step during biomarker detection and identification. In this work, a fast workflow is described capable of rapid screening for lipid components from biological tissues at ambient pressure based on liquid microjunction extraction in tandem with nano-electrospray ionization (nESI) with ultra-high resolution mass spectrometry, i.e., liquid extraction surface analysis (LESA) coupled to Fourier-transform ion cyclotron resonance (tandem) mass spectrometry (LESA-FT-ICR-MS/MS). Lipid profiles are presented for thin tissue sections of mouse brain (MB) and liver (ML) samples, analyzed in both positive and negative mode by data-dependent acquisition (DDA) tandem FT-ICR-MS/MS. Candidate assignments were based on fragmentation patterns using mostly SimLipid software and accurate mass using mostly the LipidMaps database (average sub-ppm mass error). A typical, single point surface analysis (<1 mm spatial sampling resolution) lasted less than 15 minutes and resulted in the assignment of (unique and mulitple) lipid identifications of ∼190 (MB) and ∼590 (ML) m/z values. Despite the biological complexity, this led to unique identifications of distinct lipid molecules (sub-ppm mass error) from 38 different lipid classes, corresponding to 10–30% of the lipid m/z identifications.


 Please wait while we load your content...
Please wait while we load your content...
