A simple fluorescence biosensing strategy for ultrasensitive detection of the BCR–ABL1 fusion gene based on a DNA machine and multiple primer-like rolling circle amplification†
Abstract
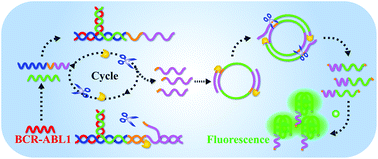
A one-step, rapid fluorescence biosensing method has been developed for ultrasensitive detection of the BCR–ABL1 fusion gene based on a polymerase/nicking endonuclease DNA machine and multiple primer-like rolling circle amplification (RCA). In the strategy, the BCR–ABL1 fusion gene can be specifically identified by using a dual probe to form a three-way junction structure (3-WJ). Then the 3-WJ based DNA machine is driven by polymerase and nicking endonuclease to generate a large number of triggers, initiating a downstream RCA reaction. The introduction of two nicking endonuclease recognition sites into a circular DNA template makes RCA occur in a multiple primer-like manner, achieving exponential growth of the signal. Benefiting from the cascade amplification, the developed method generates a wide linear response from 10 fM to 1 nM with a low detection limit of 5.52 fM. In addition, the one-step operation allows the assay to be completed within 60 min and acceptable recovery is obtained in complex samples. These merits endow the biosensing strategy with certain potential for the clinical diagnosis and scientific research of the BCR–ABL1 fusion gene.


 Please wait while we load your content...
Please wait while we load your content...