Capturing coacervate formation and protein partition by molecular dynamics simulation†
Abstract
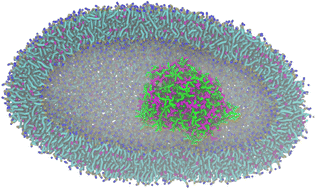
Biomolecules localize and function in microenvironments where their local concentration, spatial organization, and biochemical reactivity are regulated. To compartmentalize and control the local properties of the native microenvironment, cellular mimics and artificial bioreactors have been developed to study the properties of membraneless organelles or mimic the bio-environment for life origin. Here, we carried out molecular dynamics simulation with the Martini 3.0 model to reproduce the experimental salt concentration and pH dependency of different complex coacervates. We showed that coacervates inside vesicles are able to change their shape. In addition, we used these coacervate systems to explore the partitioning of the ubiquitous cytoskeletal protein actin and found that actin spontaneously partitions to all the coacervate peripheries. Therefore, we believe that our study can provide a better understanding of the versatile coacervate platform, where biomolecules partition and gather to fulfill their biological duties.


 Please wait while we load your content...
Please wait while we load your content...