Reconstruction and validation of entire virus model with complete genome from mixed resolution cryo-EM density
Abstract
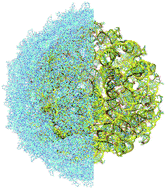
It is very difficult to reconstruct computationally a large biomolecular complex in its biological entirety from experimental data. The resulting atomistic model should not contain gaps structurally and it should yield stable dynamics. We, for the first time, reconstruct from the published incomplete cryo-EM density a complete MS2 virus at atomistic resolution, that is, the capsid with the genome, and validate the result by all-atom molecular dynamics with explicit water. The available experimental data includes a high resolution protein capsid and an inhomogeneously resolved genome map. For the genomic RNA, apart from 16 hairpins with atomistic resolution, the strands near the capsid’s inner surface were resolved up to the nucleic backbone level, and the innermost density was completely unresolved. As a result, only 242 nucleotides (out of 3569) were positioned, while only a fragmented backbone was outlined for the rest of the genome, making a detailed model reconstruction necessary. For model reconstruction, in addition to the available atomistic structure information, we extensively used the predicted secondary structure of the genome (base pairing). The technique was based on semi-automatic building of relatively large strands of RNA with subsequent manual positioning over the traced backbone. The entire virus structure (capsid + genome) was validated by a molecular dynamics run in physiological solution with ions at standard conditions confirming the stability of the model.
- This article is part of the themed collection: Challenges in biological cryo electron microscopy


 Please wait while we load your content...
Please wait while we load your content...