Semi-supervised machine learning workflow for analysis of nanowire morphologies from transmission electron microscopy images†
Abstract
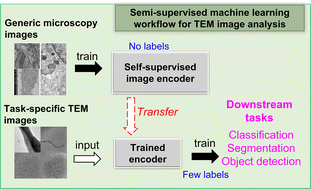
In the field of materials science, microscopy is the first and often only accessible method for structural characterization. There is a growing interest in the development of machine learning methods that can automate the analysis and interpretation of microscopy images. Typically training of machine learning models requires large numbers of images with associated structural labels, however, manual labeling of images requires domain knowledge and is prone to human error and subjectivity. To overcome these limitations, we present a semi-supervised transfer learning approach that uses a small number of labeled microscopy images for training and performs as effectively as methods trained on significantly larger image datasets. Specifically, we train an image encoder with unlabeled images using self-supervised learning methods and use that encoder for transfer learning of different downstream image tasks (classification and segmentation) with a minimal number of labeled images for training. We test the transfer learning ability of two self-supervised learning methods: SimCLR and Barlow-Twins on transmission electron microscopy (TEM) images. We demonstrate in detail how this machine learning workflow applied to TEM images of protein nanowires enables automated classification of nanowire morphologies (e.g., single nanowires, nanowire bundles, phase separated) as well as segmentation tasks that can serve as groundwork for quantification of nanowire domain sizes and shape analysis. We also extend the application of the machine learning workflow to classification of nanoparticle morphologies and identification of different type of viruses from TEM images.


 Please wait while we load your content...
Please wait while we load your content...