Least absolute shrinkage and selection operator-based prediction of collision cross section values for ion mobility mass spectrometric analysis of lipids†
Abstract
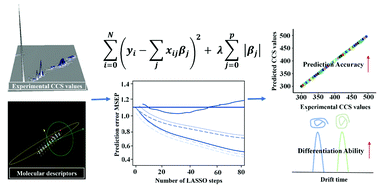
Collision cross section (CCS) values generated from ion mobility mass spectrometry (IM-MS) have commonly been employed to facilitate lipid identification. However, this is hindered by the limited available lipid standards. Recently, CCS values were predicted by means of computational calculations, though the prediction precision was generally not good and the predicted CCS values of the lipid isomers were almost identical. To address this challenge, a least absolute shrinkage and selection operator (LASSO)-based prediction method was developed for the prediction of lipids’ CCS values in this study. In this method, an array of molecular descriptors were screened and optimized to reflect the subtle differences in structures among the different lipid isomers. The use of molecular descriptors together with a wealth of standard CCS values for the lipids (365 in total) significantly improved the accuracy and precision of the LASSO model. Its accuracy was externally validated with median relative errors (MREs) of <1.1% using an independent data set. This approach was demonstrated to allow differentiation of cis/trans and sn-positional isomers. The results also indicated that the LASSO-based prediction method could practically reduce false-positive identifications in IM-MS-based lipidomics.


 Please wait while we load your content...
Please wait while we load your content...