A copper-free and enzyme-free click chemistry-mediated single quantum dot nanosensor for accurate detection of microRNAs in cancer cells and tissues†
Abstract
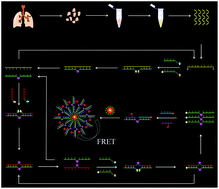
MicroRNAs (miRNAs) play key roles in the post-transcriptional regulation of genes, and their aberrant expression may disturb the normal gene regulation network to induce various diseases, and thus accurate detection of miRNAs is essential to early clinical diagnosis. Herein, we develop for the first time a single-quantum dot (QD)-based Förster resonance energy transfer (FRET) nanosensor to accurately detect miRNAs based on copper-free and enzyme-free cycling click chemistry-mediated tricyclic ligase chain reaction (LCR) amplification. We design four DNA probes namely DNA probes 1–4, with DNA probes 1 and 3 being modified with azide (N3) and DNA probes 2 and 4 being modified with dibenzocyclooctyne (DBCO). When target miRNA is present, DNA probes 1 and 2 can proceed via copper-free and enzyme-free click chemistry to generate the probes 1–2 ligation product. Subsequently, DNA probes 3 and 4 can hybridize with the probes 1–2 ligation product to generate the probes 3–4 ligation product. Both the probes 1–2 ligation product and probes 3–4 ligation product can act as the templates to initiate cycling click chemistry-mediated tricyclic LCR amplification whose products can be easily measured by the single-QD-based FRET nanosensor. This assay does not involve any enzymatic reverse transcription, copper catalyst, and ligase enzyme, and it exhibits excellent selectivity, high sensitivity, and the capability of differentiating even single-base mismatches. Moreover, this nanosensor can accurately quantify miRNA-155 even at the single-cell level, and it can distinguish the miRNA-155 expression in tissues of healthy persons and nonsmall cell lung cancer (NSCLC) patients.
- This article is part of the themed collections: microRNA and its role in gene regulation: Celebrating the 2024 Physiology or Medicine Nobel Prize and 2021 Chemical Science HOT Article Collection


 Please wait while we load your content...
Please wait while we load your content...