Fragment-based covalent ligand discovery
Abstract
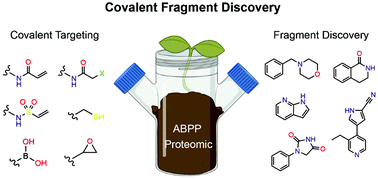
Targeted covalent inhibitors have regained widespread attention in drug discovery and have emerged as powerful tools for basic biomedical research. Fueled by considerable improvements in mass spectrometry sensitivity and sample processing, chemoproteomic strategies have revealed thousands of proteins that can be covalently modified by reactive small molecules. Fragment-based drug discovery, which has traditionally been used in a target-centric fashion, is now being deployed on a proteome-wide scale thereby expanding its utility to both the discovery of novel covalent ligands and their cognate protein targets. This powerful approach is allowing ‘high-throughput’ serendipitous discovery of cryptic pockets leading to the identification of pharmacological modulators of proteins previously viewed as “undruggable”. The reactive fragment toolkit has been enabled by recent advances in the development of new chemistries that target residues other than cysteine including lysine and tyrosine. Here, we review the emerging area of covalent fragment-based ligand discovery, which integrates the benefits of covalent targeting and fragment-based medicinal chemistry. We discuss how the two strategies synergize to facilitate the efficient discovery of new pharmacological modulators of established and new therapeutic target proteins.
- This article is part of the themed collection: 2021 RSC Chemical Biology HOT Article Collection


 Please wait while we load your content...
Please wait while we load your content...