The robust NMR toolbox for metabolomics†
Abstract
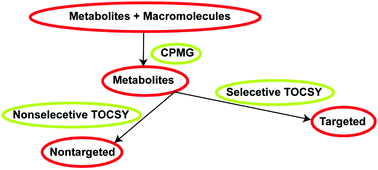
Here, we implemented and validated a suite of selective and non-selective CPMG-filtered 1D and 2D TOCSY/HSQC experiments for metabolomics research. They facilitated the unambiguous identification of metabolites embedded in broad lipid and protein signals. The 2D spectra improved non-targeted analysis by removing the background broad signals of macromolecules.


 Please wait while we load your content...
Please wait while we load your content...