Establishment of a universal and rational gene detection strategy through three-way junction-based remote transduction†
Abstract
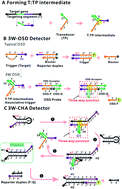
The polymerase chain reaction and many isothermal amplifications are able to achieve super gene amplification. Unfortunately, most commonly-used transduction methods, such as dye staining and Taqman-like probing, still suffer from shortcomings including false signals or difficult probe design, or are incompatible with multi-analysis. Here a universal and rational gene detection strategy has been established by translating isothermal amplicons to enzyme-free strand displacement circuits via three-way junction-based remote transduction. An assistant transduction probe was imported to form a partial hybrid with the target single-stranded nucleic acid. After systematic optimization the hybrid could serve as an associative trigger to activate a downstream circuit detector via a strand displacement reaction across the three-way junction. By doing so, the detection selectivity can be double-guaranteed through both amplicon–transducer recognition and the amplicon–circuit reaction. A well-optimized circuit can be immediately applied to a new target detection through simply displacing only 10–12 nt on only one component, according to the target. More importantly, this property for the first time enables multi-analysis and logic-analysis in a single reaction, sharing a single fluorescence reporter. In an applicable model, trace amounts of Cronobacter and Enterobacteria genes have been clearly distinguished from samples with no bacteria or one bacterium, with ultra-high sensitivity and selectivity.


 Please wait while we load your content...
Please wait while we load your content...