Post hoc support vector machine learning for impedimetric biosensors based on weak protein–ligand interactions†
Abstract
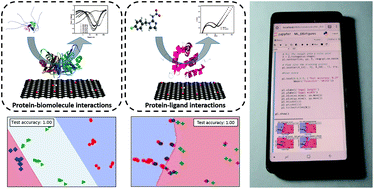
Impedimetric biosensors for measuring small molecules based on weak/transient interactions between bioreceptors and target analytes are a challenge for detection electronics, particularly in field studies or in the analysis of complex matrices. Protein–ligand binding sensors have enormous potential for biosensing, but achieving accuracy in complex solutions is a major challenge. There is a need for simple post hoc analytical tools that are not computationally expensive, yet provide near real time feedback on data derived from impedance spectra. Here, we show the use of a simple, open source support vector machine learning algorithm for analyzing impedimetric data in lieu of using equivalent circuit analysis. We demonstrate two different protein-based biosensors to show that the tool can be used for various applications. We conclude with a mobile phone-based demonstration focused on the measurement of acetone, an important biomarker related to the onset of diabetic ketoacidosis. In all conditions tested, the open source classifier was capable of performing as well as, or better, than the equivalent circuit analysis for characterizing weak/transient interactions between a model ligand (acetone) and a small chemosensory protein derived from the tsetse fly. In addition, the tool has a low computational requirement, facilitating use for mobile acquisition systems such as mobile phones. The protocol is deployed through Jupyter notebook (an open source computing environment available for mobile phone, tablet or computer use) and the code was written in Python. For each of the applications, we provide step-by-step instructions in English, Spanish, Mandarin and Portuguese to facilitate widespread use. All codes were based on scikit-learn, an open source software machine learning library in the Python language, and were processed in Jupyter notebook, an open-source web application for Python. The tool can easily be integrated with the mobile biosensor equipment for rapid detection, facilitating use by a broad range of impedimetric biosensor users. This post hoc analysis tool can serve as a launchpad for the convergence of nanobiosensors in planetary health monitoring applications based on mobile phone hardware.

 Please wait while we load your content...
Please wait while we load your content...